Introduction to Active Learning Glide
Active Learning Glide combines traditional Glide SP docking with state-of-the-art deep learning to efficiently screen ultra-large compound libraries.
Building Machine Learning (ML) Models
Selecting Compounds for Active Learning
Background

Active Learning Glide combines Glide's high-performance ligand-receptor docking with Schrödinger’s state-of-the-art deep learning to efficiently screen ultra-large compound libraries.
Active Learning Glide is built on the premise that given docking scores for a sufficient number of ligands, an Machine Learning (ML) model can “learn” to recognize features in the ligand structures that can be predictive of the docking score. This trained model can be used to predict the docking scores of ligands outside the training set without needing to dock them first. This method exploits the computational efficiency of ML evaluation (ML evaluation in the Schrodinger Suite is roughly 1000x faster than Glide docking) to allow screening of ultra-large libraries (+ 1BLN ligands) in a reasonable amount of time.
Building Machine Learning (ML) Models
ML models in Active Learning Glide are built using the DeepAutoQSAR package inside Schrodinger Suite. DeepAutoQSAR optimizes the parameters of many types of ML models (hyperparameters) and constructs an ensemble of three best-performing models. Every model is trained to a different pool of data “fold”. The program will continue hyperparameter search until a sufficient number of models have been reached or a predetermined stop time has been reached.
Types of models: DeepAutoQSAR considers traditional ML models and neural network models from deep-learning. The following classical models are considered: Random Forest, XGBoost. The following neural network models are considered: Graph Convolutional (GCNN), Fully-Connected Convolutional.
Selecting Compounds for Active Learning
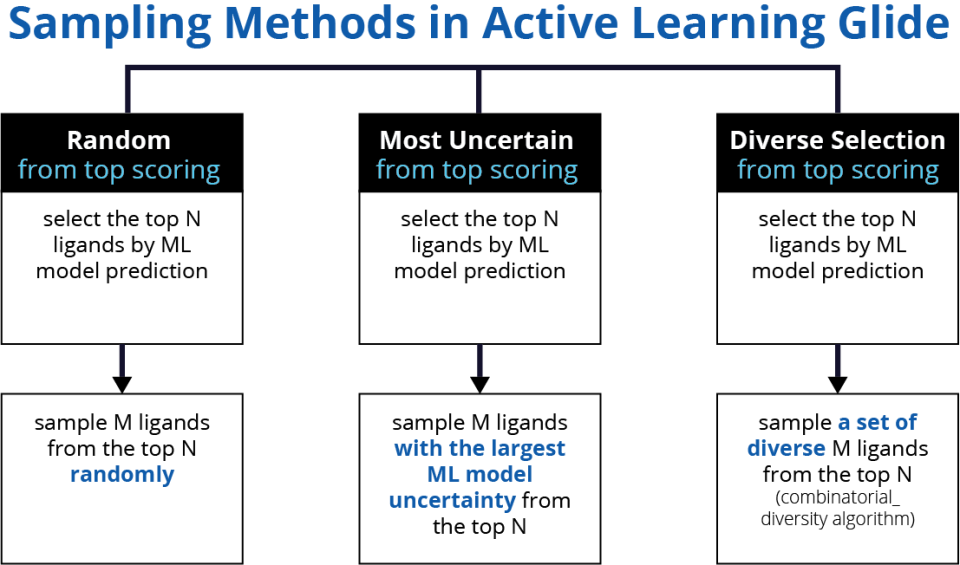
In the first iteration, a random selection of ligands is made. These ligands are docked with Glide and form the initial training set for ML training. For subsequent iterations, we bias selection toward ligands that score well using the ML-model score over the entire library. This is done to improve model performance where it matters the most - at the very top of the ranked list.

Pilot Mode vs. Glide
Active Learning Glide trains ML models to the docking scores of ligands and not to any experimental binding affinity. When evaluating the Active Learning Glide workflow as a substitute for large brute force docking, we want to see how well the workflow captures the very best ligands in the library according to Glide. Using pilot mode in AL-Glide we can answer this question using a slice of the screening library called the pilot library. The pilot library is run through a single iteration of the Active Learning Glide workflow. In parallel, the pilot library is docked fully using Glide SP.
Pilot Report example
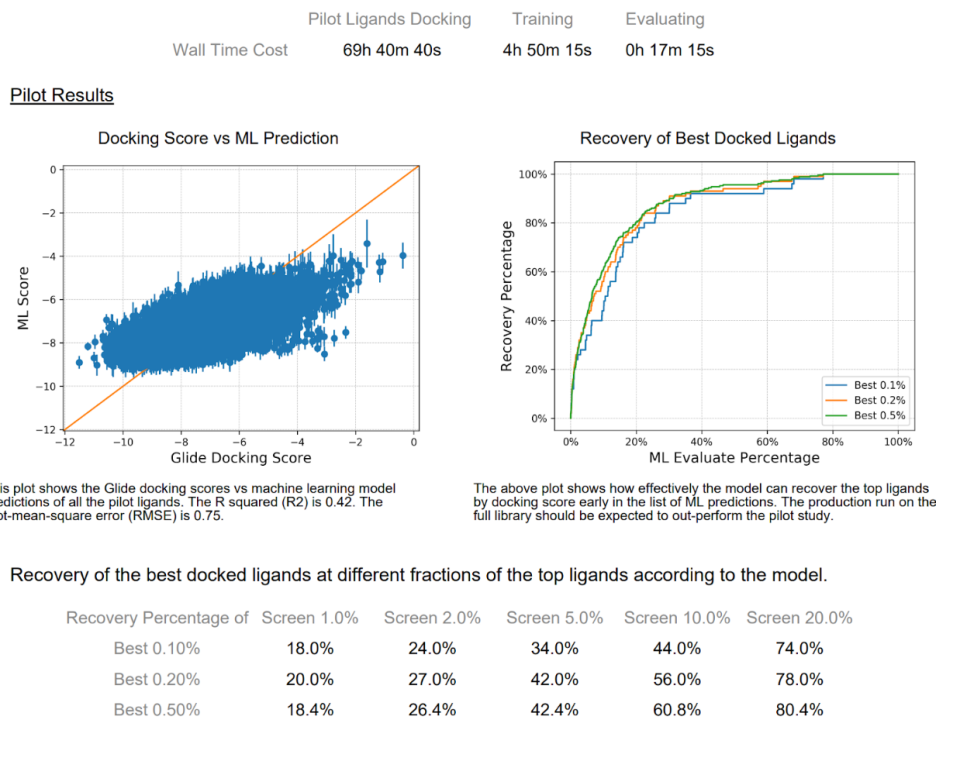
Report Interpretation
A PDF report is generated after a pilot run of Active Learning Glide. There are two plots that are shown. The first (left hand side) is a scatter plot of the ML score (docking score prediction) vs the true Glide docking score. There will typically not be a strong quantitative relationship between these two variables. An R^2 value > 0.3 is a good sign that the model is able to learn the Glide score. The most important analysis is represented in the plot on the right side of the report. Here we look at enrichment of the best compounds in the pilot library by the ML model. In virtual screening, we typically care about recovery at the very top of the list. A model with modest correlation can show excellent early enrichment of top scoring compounds. We see this in the example report above. Ultimately, the enrichment problem is really a ranking problem and not a regression problem. Therefore, we find the enrichment plot to be the single most important analysis of the pilot results. The enrichment of the productive runs is expected to be better than the pilot run, as there is a larger training set typically being used in the former.
Additionally, there is an HTML report file available for all active learning workflows. This report summarizes the workflow execution status, results, and analysis, and can be downloaded during execution. The report includes a workflow chart with color-coded stages, details for each stage; including mini flowcharts and relevant parameters, and analysis such as histograms of selected ligands, PCA plots, model performance metrics, and enriched substructures.