Epik Calculation Modes
Epik can perform four types of calculations:
-
1. pKa query (mirrors the same behavior as previous versions of Epik and can be accessed via both the command line interface (CLI) and the GUI)
-
2. State Prediction (mirrors the same behavior as previous versions of Epik and can be accessed via both the command line interface (CLI) and the GUI)
-
3. Speciation Report (Command Line only) (new to Epik and only available through the CLI at this time)
-
4. Experimental Match (Command Line only) (new to Epik and only available through the CLI at this time)
1. pKa query
This calculation is a straightforward pKa value prediction on the input molecule(s); no additional states are generated. In previous versions of Epik, pKa query was the default calculation type, but in Epik it must be called with the --query keyword via the command line or in the GUI by deselecting the “Generate states” box.
Command Line
$SCHRODINGER/epikx input.mae output.mae --query
GUI
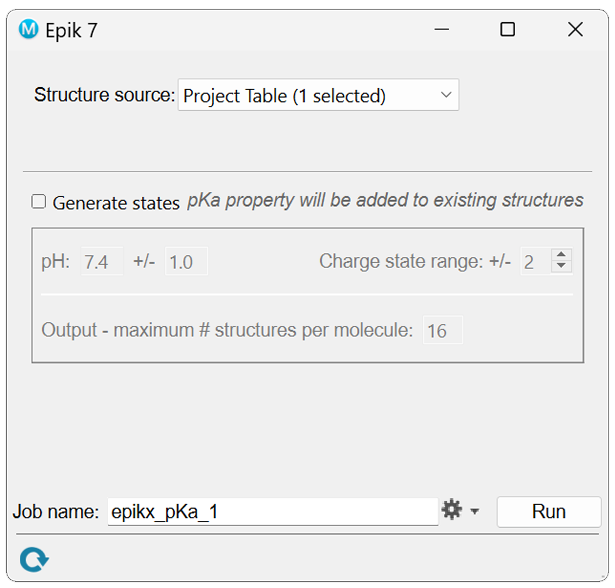
In the resulting output file the micro-pKa values and corresponding uncertainties for the ionizing atoms can be visualized in Maestro by applying the pKa-H2O” atom property label.
2. State Prediction
This calculation takes input molecules, enumerates their states, predicts the pKa values for each state, calculates the state populations at the specified pH, and then returns the most populated states ordered by population above some population threshold. This is the default mode in both the CLI and GUI.
Command Line
The user can set the pH using the keyword -ph [PH] (default = 7.4). The population threshold can be altered either directly with -p [P] (default = 0.01) or indirectly with the pH threshold -pht [PHT] (default = ±2). The example command below performs a state prediction at pH 8.5 ± 1.
$SCHRODINGER/epikx input.mae output.mae -ph 8.5 -ph1
GUI
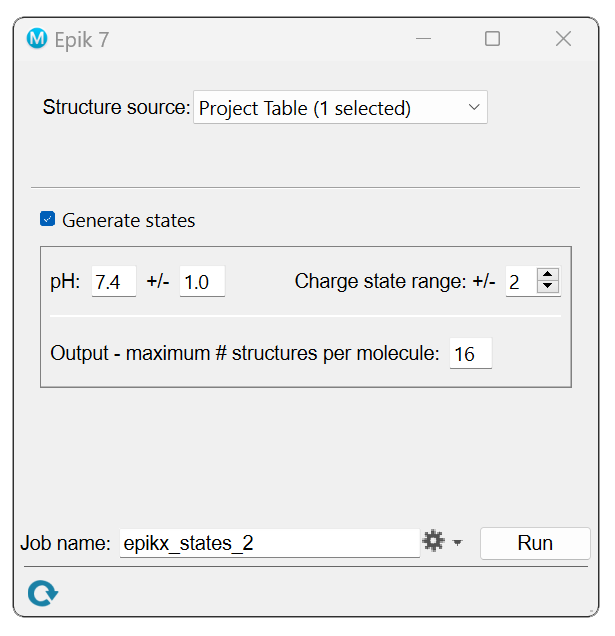
The resulting output file will contain at least as many structures as are in the input file, but usually more due to multiple protonation states for each input molecule. The resulting output states descend in order by population. As in the query mode, the micro-pKa values and their uncertainties are available through the atom-level properties. For each state, the fractional population is available through the structure-level Maestro property Population, and the state penalty in kcal/mol is available through the structure-level property State Penalty. The macro- pKa values are available through the structure-level property macro-pKa.
The macro- pKa value between q and q+1 is considered basic if the most acidic site (i.e., hydrogen with the lowest micro-pKa value) in the most populated microstate of the q+1 macrostate is either a) cationic or b) a neutral nitrogen and the number of cationic nitrogens decreases between the q+1 and q states. All other types of transitions are acidic. The top four of each category of macro- pKa values are ranked and assigned the Maestro property acidic macro-pKa {1-4} for the lowest acidic macro- pKa values and basic macro-pKa {1-4} for the highest basic macro- pKa values. For example, the most acidic macro- pKa value is assigned the property acidic macro-pKa 1. The ionizing site that corresponds to each categorized macro- pKa contains the structure-level property acidic atom {1-4} and basic atom {1-4}.
3. Speciation Report (Command Line only)
Epik can also produce a detailed speciation report for a single structure entry. This mode is not available via the GUI, and is accomplished by passing the -- report or - r argument in the CLI.
$SCHRODINGER/epikx input.mae output.mae --report
This will produce a report directory that contains a number of files, the most important of which is the HTML file ending in “..._populations.html”. Opening this file in an Internet browser of your choice displays the following page, which includes the most populated species, their pH-dependent populations (in black text), pH-independent populations (in green text), the macro-pKa values between charge levels, a speciation diagram showing their pH-dependent populations across the pH range, and the individual species’ SMILES strings.
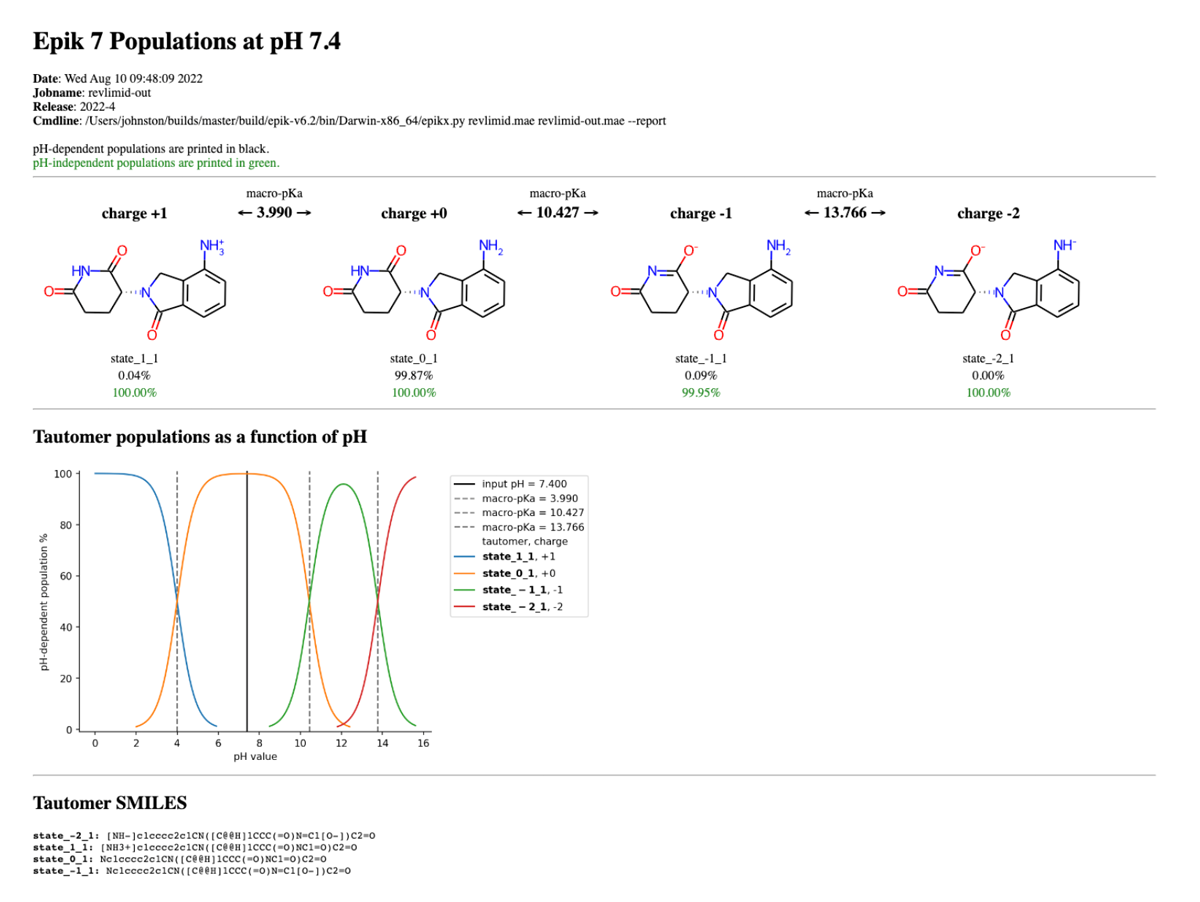
The user may generate the report at a different pH by passing the -ph [PH] argument, which will adjust the pH-dependent populations accordingly in the top half of the report. The user may similarly combine with other options, e.g., -pht [PHT], -p [P], etc., to augment the results of the speciation report.
4. Experimental Match (Command Line only)
This calculation takes an input molecule and, given an experimental pKa value in a structure property, predicts the macro-pKa values and returns the best protonated state that matches the experimental pKa value that is present as a structure-level property on each molecule in the input structure. For multiple experimental pKa values, the experimental pKa property must correspond to a single value, and each experimental value associated with a single structure. To use this mode, provide the property label that contains the experimental pKa value with --match_exp [EXP_PKA_PROPERTY]:
$SCHRODINGER/epikx input.mae output.mae --match_exp <EXP_PKA_PROPERTY>
The output file will contain as many structures as are in the input with the most populated protonated state from the ionization whose macro-pKa value most closely matches the experimental value. The properties of the previous modes are still available through the usual routes. The index of the most acidic atom in the protonation state is available through the structure property label “best acid”.
The user may generate the report at a different pH by passing the -ph [PH] argument, which will adjust the pH-dependent populations accordingly in the top half of the report. The user may similarly combine with other options, e.g., -pht [PHT], -p [P], etc., to augment the results of the speciation report.