Using a Custom Core in the FEP+ Panel
Ligands with asymmetric substituted rings may have multiple binding poses that cannot be efficiently sampled by the default FEP+ protocol. Using custom cores to put these asymmetric rings in the alchemical region can enhance the sampling of the flipping of these rings. Another place where a custom core is needed is when the ligands consist of the same two fragments joined by different linkers ("linker enumeration"); here the linker is identified, and the "core" is the two fragments.
Custom Core
A custom core can be defined when you generate a perturbation map. When you click Generate Map, choose Core from the Define custom substructure option menu in the Map Options Dialog Box, then enter the SMARTS pattern of the core. It is a good idea to first select a node in the current map so that one of the ligands is displayed in the Workspace; then you can select the atoms of the core on that ligand to generate the SMARTS pattern for the custom core. (You can also specify multiple SMARTS patterns - see below.)
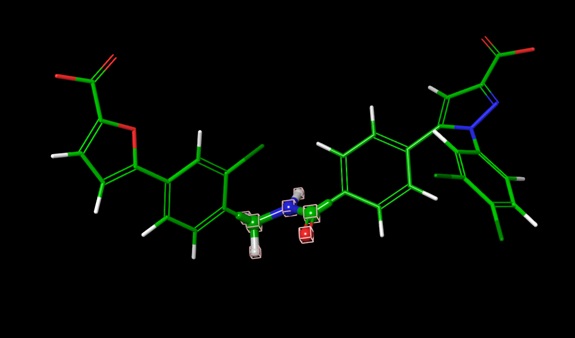
As an example, for the following three Thrombin ligands 6a, 3a, and 1d, each with an asymmetrically substituted phenyl ring, the default FEP+ protocol includes the phenyl ring in the core region, and only the substitutions on the ring are in the alchemical region. Since the phenyl ring is in the core region, and has full interaction with the protein environment in all replicas, the flipping of the ring might be blocked, resulting in insufficient sampling.
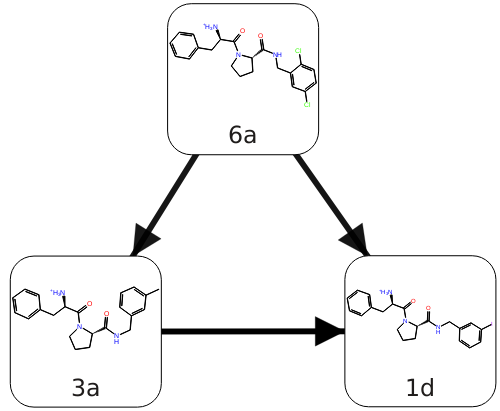
Figure 1:Three thrombin ligands with asymmetric rings.
By using the custom core option when generating the map and setting the core of the perturbation to be everything except the asymmetric ring (Fig. 2), the asymmetric ring will be in the alchemical region for these perturbations. The asymmetric ring in the dummy state does not have interactions with the protein environment, thus can easily flip the orientations, and these flipped orientations will propagate into the physical state through replica exchange. By this means, using the custom core option can enhance the sampling of the asymmetric ring.
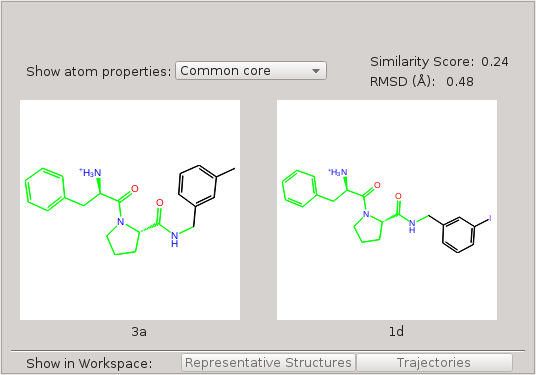
Figure 2:Custom core corresponding to everything except the asymmetric ring.
The application of a custom core to a pair of structures depends on whether they match the core SMARTS pattern.
-
Both molecules match the core SMARTS pattern
If both molecules have a substructure that matches the core SMARTS pattern, this substructure will be the core of the perturbation. For the example above, both molecules 3a and 1d have a substructure that matches the SMARTS pattern, and the core of the perturbation is this substructure. (Fig. 2)
-
Only one molecule matches the core SMARTS pattern
Now, suppose we add another two molecules, 3a_mod and 1d_mod, into the above map. 3a_mod and 1d_mod differ from the corresponding 3a and 1d by an extra methyl group on the carbon that joins the asymmetric ring to the rest of the molecule. For the perturbation between 1d_mod and 1d, only 1d has a substructure matching the above SMARTS pattern while 1d_mod does not, due to the additional methyl group. In this case, the core of the perturbation is the maximum common substructure (MCS) between 1d_mod and the substructure in 1d that matches the SMARTS pattern. In other words, the core of the perturbation is the intersection of the maximum common substructure between the two molecules and the core SMARTS pattern.
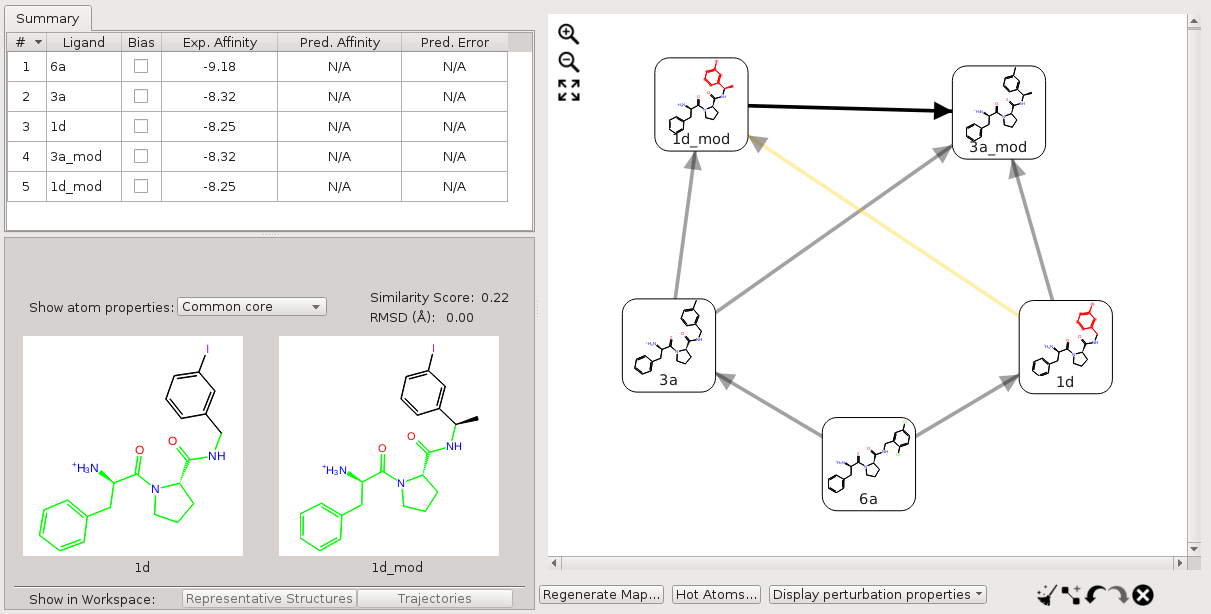
Figure 3:Core of the perturbations with only one molecule matching the custom core SMARTS pattern.
-
Neither molecule matches the core SMARTS pattern
For perturbations, between 1d_mod and 3a_mod, neither molecule has a substructure matching the above custom core SMARTS pattern. In this case, the custom core is ignored, and the default mapping without a custom core is used (Fig. 4).
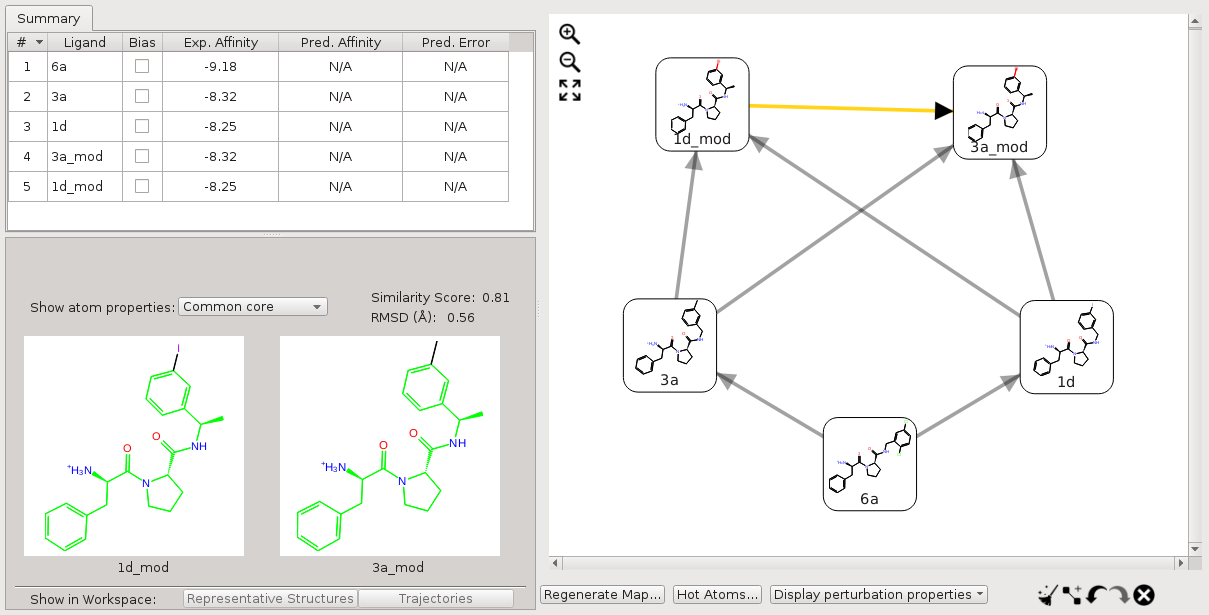
Figure 4:Perturbations where neither molecule has a substructure matching the core SMARTS pattern.
For a set of molecules where only a subset has a substructure that matches a core SMARTS pattern while others do not, the default map may contain edges where neither molecule matches the SMARTS pattern. If this happens, it is best to connect the molecules that do not match the SMARTS pattern with other molecules that match the SMARTS pattern, and delete the edges where neither molecule matches the SMARTS pattern. For example, for the five molecules discussed above, we can add edges 3a→3a_mod, and 1d→1d_mod, and delete the edge between 3a_mod and 1d_mod. In this way, the custom core is effective for all the perturbations.
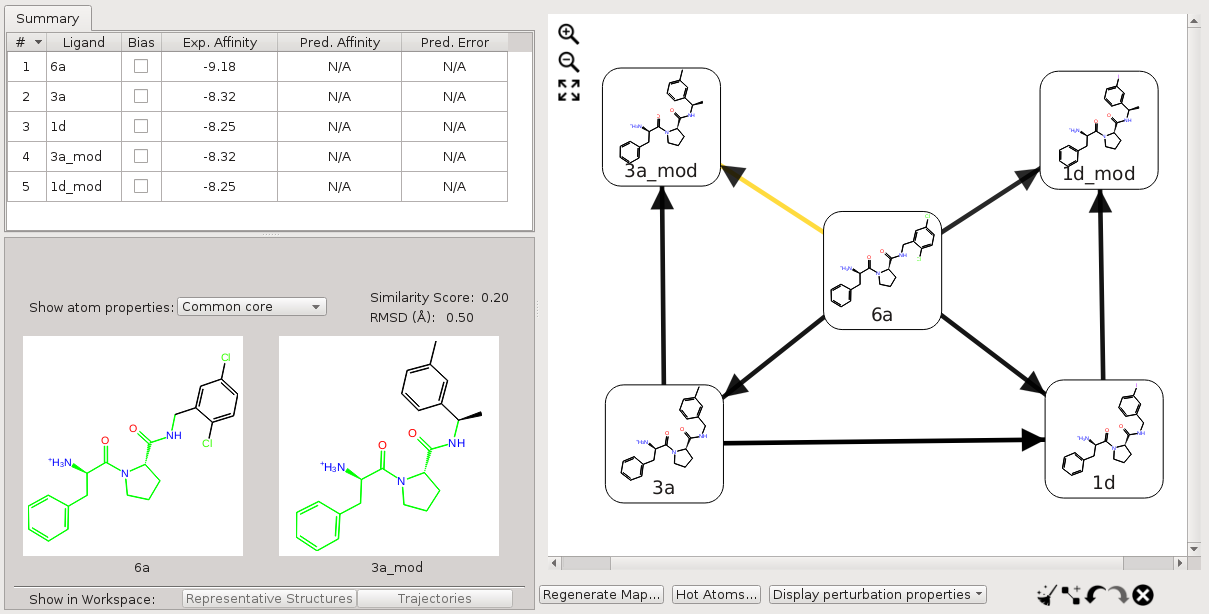
Figure 5:Perturbation map for a set of molecules where only a subset has a substructure matching a core SMARTS pattern.
An alternative is to provide multiple SMARTS patterns, so that all (or more) structures match a core SMARTS pattern. You can enter multiple SMARTS patterns in the text box, separated by spaces. The SMARTS patterns are tried in order of size (number of atoms) on each ligand and the first one that matches for a given ligand is used for the core for that ligand. The matching rules are the same as above when one ligand or neither ligand matches a core SMARTS pattern, or both ligands match the same core SMARTS pattern. In the case where each ligand matches a different SMARTS pattern, the core is the intersection of the two patterns, i.e. it is the maximum common substructure of the two patterns.
For example, consider the following three molecules.
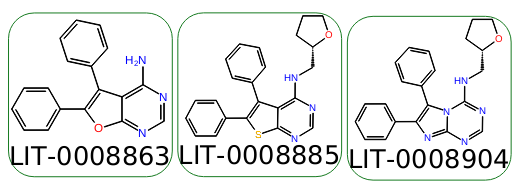
We want the phenyl rings to be in the hot region, but by default, the phenyl rings are included in the core. The core we want to use is the fused ring system, but none of them are the same. If we use a single SMARTS pattern, "n1ccn(c12)cnc(n2)[H]", which matches the third ligand (LID-0008904), the two edges that include this ligand have mappings only within the desired core region:
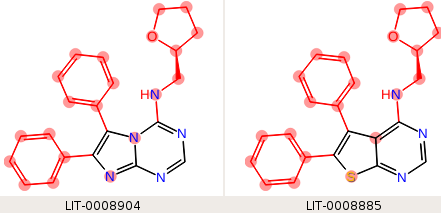
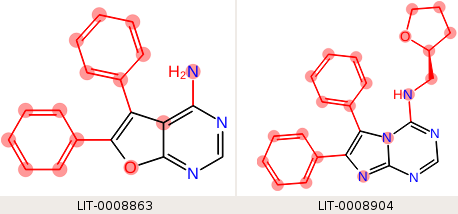
but neither of the ligands in the third edge (the first two above) matches the core SMARTS pattern, so the maximum common substructure is used, and the hot region does not include the phenyl rings.
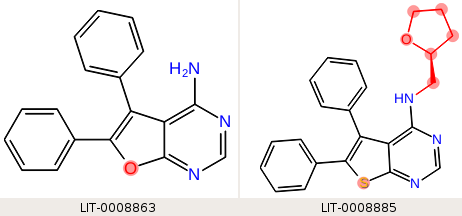
If we provide three SMARTS patterns, one for each of the fused ring systems, "c1csc(c12)nc([H])nc2 n1ccn(c12)cnc(n2)[H] c1coc(c12)nc([H])nc2", then each ligand matches one core SMARTS pattern, and the third leg of the perturbation has only the fused ring system in the core:
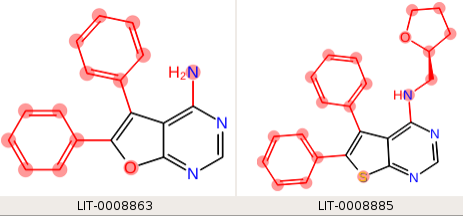
Linker Enumeration
For linker enumeration, the linker can be defined when you generate a perturbation map. When you click Generate Map, choose Linker from the Define custom substructure option menu in the Map Options Dialog Box, then enter the SMARTS pattern of the linker.
You can select a node in the current map to display one of the ligands in the Workspace, and pick the linker atoms in the Workspace to generate the SMARTS pattern. The two fragments that constitute the "core" are identified as the atoms not in the SMARTS pattern. The fragments must be the same in each ligand in the perturbation map.