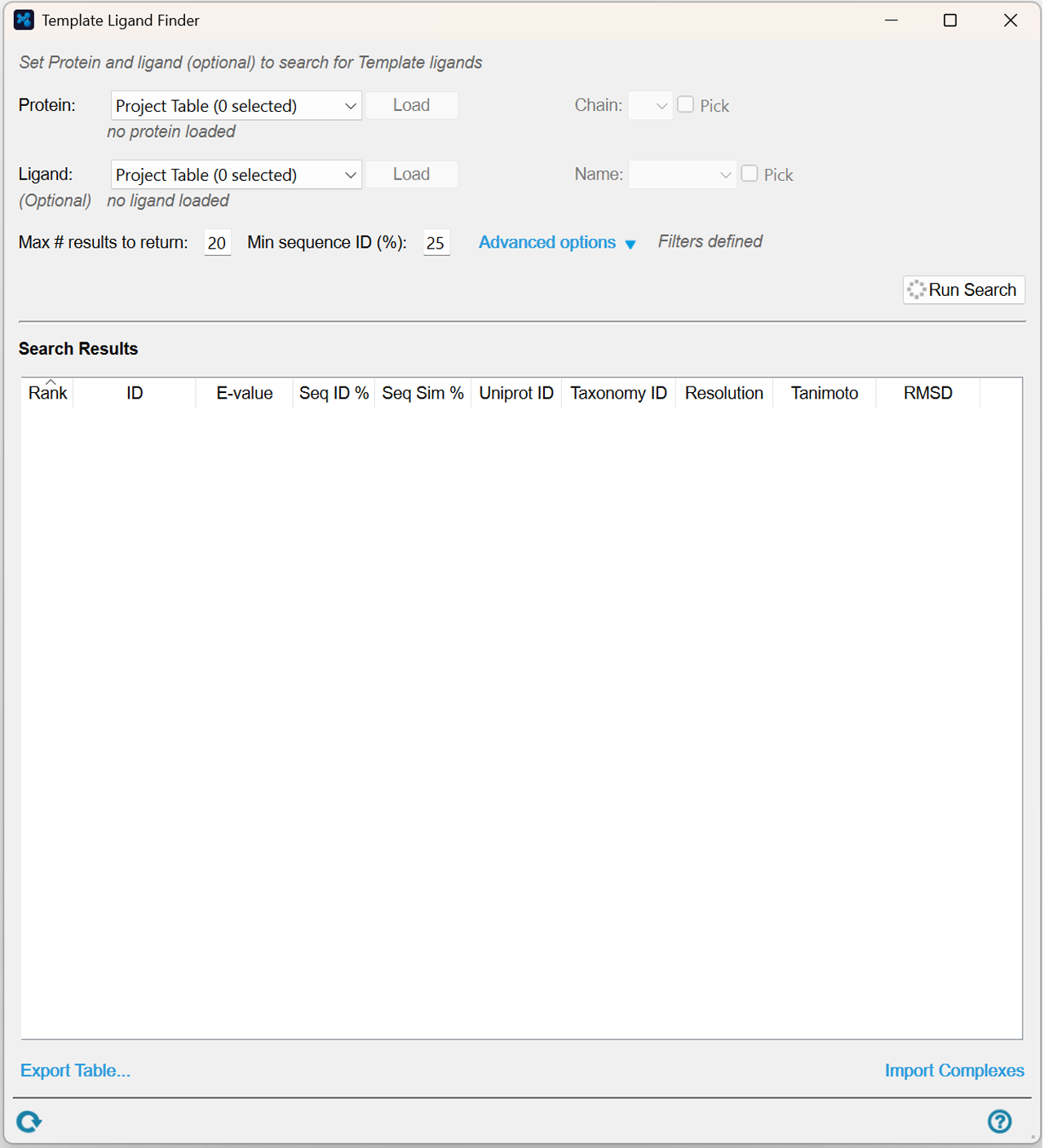
Template Ligand Finder (Beta)
Template Ligand Finder panel is for ligand-binding mode prediction and cryptic binding site identification. Rapidly identify and visualize homologous proteins based on both protein and ligand similarity to uncover potential template ligands as references for IFD-MD.
To open this panel, click the Tasks button and browse to Receptor-Based Virtual Screening → Template Ligand Finder. Alternatively, click the Tasks button and browse to Induced Fit Docking → Template Ligand Finder.
- Features
- Additional Resources
Template Ligand Finder Features
Input Settings
Set protein and the optional ligand to search for template ligands.
- Protein and chain selection menus
-
Select the source (Project Table, from File, or from an entry in the Workspace) for the desired protein from the dropdown menu and click the Load button. Select the chain letter from the adjacent dropdown menu (this is required if the protein is from the Workspace). Click the Pick button to make the selection for the chain in the Maestro workspace.
- Ligand and name selection menus
-
Select the source (Project Table, from File, or from an entry in the Workspace) for the optional ligand from the dropdown menu and click the Load button. Also select the ligand name from the adjacent dropdown menu. Click the Pick button to make the selection for the ligand in the Maestro workspace.
Search Filters
- Max # results to return option
-
Enter a value in the text box to set the maximum number of results to return in the Search Results.
- Min sequence ID (%) option
-
Enter a value in the text box to set the threshold for querying BLAST hits. Hits with a sequence identity value below the threshold will be ignored. The default value is 25.
Clicking the blue text opens a menu with several settings for Structure Options, BLAST Search Options, and ID Filters.
- Structure options
- Use the text boxes, arrow keys and dropdown menu to adjust the Minimum Tanimoto score, Maximum resolution (in Å), and Release date range.