Conformational Searches With Embrace
Embrace can perform conformational searches in addition to minimizations. Energy difference mode is the only mode supported for conformational searches with Embrace. Searches are conducted on the receptor, each ligand, and each ligand-receptor complex. The energies used in the energy difference equation
for the receptor and the ligand are the values from the lowest energy conformations found for those systems. However, multiple complex conformations may be retained and a separate Ecomplex is used in the energy difference equation for each one (MCOP arg6 specifies the number of conformations to keep).
The Embrace Conformational Search Panel is used to set up and submit Embrace conformational search jobs. To open this panel, choose click the Tasks button and browse to MacroModel → Embrace Conformational Search. The controls for the Embrace settings are located in the Embrace tab, which is described in the Embrace Minimization Panel topic. The conformational search parameters are set up in the CSearch tab. This tab contains a subset of the controls found in the CSearch tab for regular conformational searches and is described in the Embrace Conformational Search Panel topic.
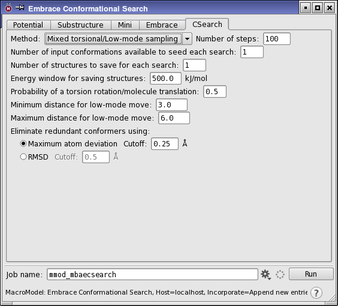
Embrace conformation searches automatically employ the AUTO Automatic Setup mechanism. AUTO selects the MCMM and comparison atom parameters for each individual ligand-receptor complex. Computations prepared from Maestro have the AUTO opcode added to the job’s command file (jobname.com) automatically. In addition, AUTO is substructure-aware. Parameters are only indicated for the freely moving receptor and ligand atoms, but not for fixed or frozen regions of the receptor. Thus, only a proper receptor substructure needs to be indicated to prepare the appropriate MCMM parameters. Examples can be found in Specifying a Substructure for Embrace. In addition, only the substructure facility can be used to indicate fixed or frozen atoms in an Embrace calculation, and ligand atoms cannot be fixed or frozen. Other than by adjusting the substructure, it is not possible to adjust the AUTO parameters for individual complexes in an Embrace automated calculation.
Conformational searches on protein-ligand complexes are computationally intensive, and the use of substructures is strongly recommended to reduce the CPU time and memory required. Even with fairly small substructures and very short searches, such searches require hours or days for each ligand processed.