Prime MM-GBSA
The Prime MM-GBSA Panel can be used to calculate ligand binding energies and ligand strain energies for a set of ligands and a single receptor, using the MM-GBSA technology available with Prime. You can also run the job from the command line, which offers more flexibility than the panel (see prime_mmgbsa—Calculate Ligand Binding Energies). The energies and other properties are added to the Maestro file, and are described in Prime MM-GBSA Output.
The ligands and the receptor must be properly prepared beforehand, for example, by using LigPrep and the Protein Preparation Workflow. Waters and ions must be removed from the protein, as an implicit solvent model is used. The ligands must be pre-positioned with respect to the receptor, and the receptor must be prepared as for a Prime refinement calculation.
If the receptor is a membrane-bound protein, addition of an implicit membrane must be done as a preparation task. See Setting Up an Implicit Membrane for more information.
To open the Prime MM-GBSA Panel, click the Tasks button and browse to Lead Optimization → MM-GBSA.
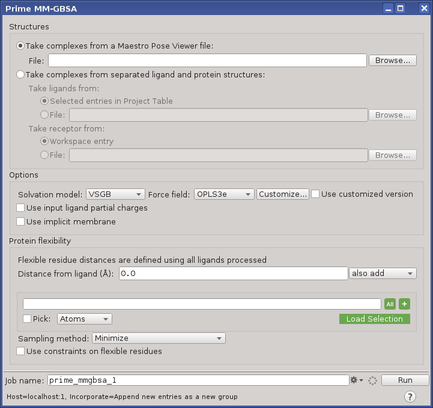
Figure 1:The Prime MM-GBSA panel.
See the Prime MM-GBSA Panel for more information.