Receptor Grid Generation
Glide searches for favorable interactions between one or more ligand molecules and a receptor molecule, usually a protein. The shape and properties of the receptor are represented on a grid by several different sets of fields that provide progressively more accurate scoring of the ligand poses. For receptors that adopt more than one conformation on binding, such as a change in side-chain conformation, backbone location or loop conformation, you should consider docking to multiple protein conformations, to ensure that possible actives are not missed. To do this you would need to prepare grids for each conformation. Glide can, however, handle different hydroxyl conformations of Ser, Thr, or Tyr residues and thiol conformations of Cys residues with a single grid generation.
If you have a receptor in which there is substantial movement upon docking, you can also use the Induced Fit Docking protocol to account for receptor flexibility. This protocol uses Glide and Prime, and is much more computationally demanding than Glide docking alone. It is therefore mainly useful for docking a small number of ligands. See the Induced Fit Docking User Manual — Contents for more information. To simulate a small amount of adjustment of the receptor atoms on docking, the van der Waals radii of the nonpolar atoms can be scaled, so that there is a little more room for the ligand to dock without clashing with receptor atoms.
Receptor grid generation requires a “prepared” structure: an all-atom structure with appropriate bond orders and formal charges. Information on structure preparation is given in Protein and Ligand Preparation for Glide.
The receptor grid can be set up and generated from the Receptor Grid Generation panel.
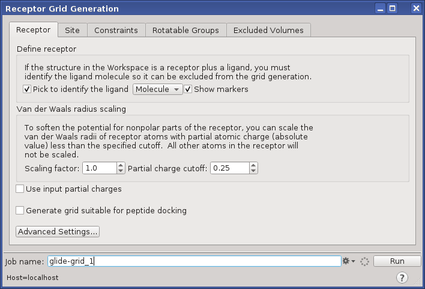
The options in each tab of this panel allow you to define the receptor structure by excluding any cocrystallized ligand that may be present, determine the position and size of the active site as it will be represented by receptor grids, set up Glide constraints, set up flexible hydroxyl or thiol groups, and define excluded volumes, where ligand atoms are heavily penalized. Ligand docking jobs cannot be performed until the receptor grids have been generated.
To open the Receptor Grid Generation panel: click the Tasks button and browse to Receptor-Based Virtual Screening → Receptor Grid Generation.
For information on setting receptor parameters, see the Receptor Grid Generation Panel topic.
When you have set up the parameters, you can run the job by clicking Run. If you want to change job settings, reset the panel settings, or write the input to files for running from the command line, use the Settings button menu. For more information, see Job Toolbar.