Adding Atoms to the Selection
There are two ways of adding atoms to the selection: by switching to add mode, and by expansion.
Using Add Mode
To add atoms to the selection by picking, you can switch into Add mode in several ways:
- Hold down the Shift key while you pick or drag.
- Click the Add to selection button on the Selection toolbar,
 .
. - Press the + key.
- Choose Select → Pick to Select → Add.
The picking mode switches to Add mode until you exit the mode. When you switch to Add mode, a plus sign appears next to the pointer to indicate that you are in this mode, and the Add to selection button is highlighted. An add button is added next to Quick Select on the toolbar. If you pick already selected atoms, they remain selected, so you do not need to be careful about picking only unselected atoms.
To exit Add mode, do one of the following:
- Release the Shift key, if you are holding it down.
- Click the Add to selection button
 .
. - Click the add button next to Quick Select on the toolbar. (The add button is hidden.)
- Press the Esc key or the + key.
- Choose Select → Pick to Select → Normal.
- Click the main Select by picking button. (Note that this also changes the draw mode for selection.)

or 
As well as picking atoms in the Workspace to add to the selection, you can pick any object in the Quick Select section of the toolbar to add to the selection, including those in the Select Objects panel, which you open by clicking the Choose item button,
or Select → More Objects. Both standard sets and custom sets can be added to the selection.
The selection scope for these objects can be modified to exclude nonpolar hydrogen atoms from the selection, so only the polar hydrogens are selected. See Selecting Hydrogen Atoms for details.
Expanding the Selection
To expand the selection to the next pick level (selection scope), double click an atom in the selection. For example, if the pick level is Atoms, double-clicking expands the selection to the next level, which is Residues or Repeating Units.
The Expand Selection tool allows you to expand the selection by various relationships between the selection and other atoms.
To open this tool, do one of the following:
-
click
 on the toolbar
on the toolbar -
right-click on a selected atom and choose Expand Selection
-
choose Select → Expand.
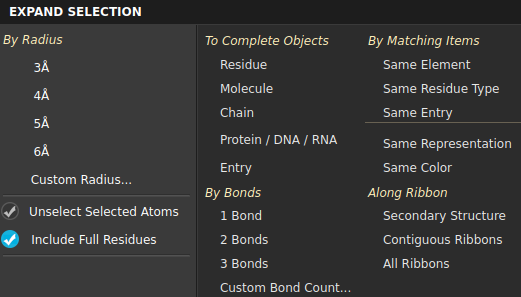
The primary option is to expand the selection by radius but several other means of expanding the selection are available. These options depend on the Maestro profile: BioLuminate has more protein-related items, whereas the protein-related items are not present in the Materials Science profile.
-
By Radius—add all atoms that are within the specified distance of any atom in the selection. If you select the option Include Full Residues, these atoms are expanded to include all atoms in the residues that they are in. This is equivalent to picking with the selection scope set to Residues rather than Atoms. Several preset radius values are available, but you can set your own by choosing Custom Radius, and entering the desired value in the dialog box that opens.
For a structure with periodic boundaries, if the radius crosses the cell boundary the image atoms in the neighboring cell are used to determine the selection, in addition to the atoms inside the cell. The selection includes all atoms found to be within the radius and their images in all cells that are displayed. For example, suppose you have a cell has a benzene molecule at both ends. You select an atom in one of them, then expand the selection to a given radius. The direct distance to the benzene molecule at the other end of the cell is larger than the radius, but the distance to its image in the neighboring cell is inside the radius. Both benzene molecules in the cell are selected. If you are also showing the neighboring cell, both benzene molecules in both cells are selected.
The expansion to a given radius can also be done as a replacement, by selecting the option Unselect Selected Atoms. In this mode, the atoms in the expansion are selected, and the original selection is cleared. This is useful, for example, if you want to pick the residues in the active site of a protein that are near the ligand. First you pick the ligand, then you expand the selection to the desired radius, with the options Unselect Selected Atoms and Include Full Residues active. Note that this option does not apply to expansions other than by radius.
-
By Bonds— add any unselected atom that is within the chosen number of bonds from a selected atom. These options expand the selection according to the connectivity. You can use a different number of bonds by clicking Custom Bond Count and entering the desired number.
-
By Matching Items—add all atoms with the same chosen characteristic as the atoms in the selection. For example, if a GLY and an ARG residue are selected, choosing Same Residue Type selects all GLY and ARG residues.
-
Along Ribbon—expand the selection to include all atoms within the same chosen ribbon feature. For example, if you have some atoms selected in a helix and others selected in a loop, choosing Secondary Structure selects all atoms in the same helix as the selected atoms in the helix and all atoms in the same loop as the selected atoms in the loop.
Note that the expanded selection can include atoms that are not visible.
The selection scope for expansion can be modified to exclude nonpolar hydrogen atoms from the selection, so only the polar hydrogens are selected. See Selecting Hydrogen Atoms for details.