Sequence Viewer
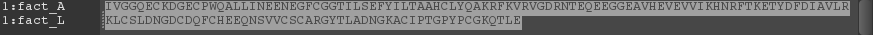
The sequence viewer displays the sequence of the proteins that are included in the Workspace. Each chain is displayed on a separate line. The left column of the sequence viewer lists the entry name for the sequence, with the chain name appended. The right column displays the actual sequence as a list of single-letter codes, and also displays the secondary structure assignment, if one is available. If the protein contains ligands, waters, or ions, these are displayed in separate "sequences", one for each type of molecule. (These molecules are identified by the corresponding ASL expressions.)
To show or hide the sequence viewer, choose Window → Sequence Viewer.
Clicking the blue menu icon on the left side of the Sequence Viewer title bar will open the Sequence Viewer Shortcut Menu. To expand or collapse the Sequence Viewer pane, click the chevron icon on the right side of the Sequence Viewer title bar to toggle. When the pane is expanded, clicking and dragging the splitter above the Sequence Viewer title bar re-sizes the Workspace and the Sequence Viewer pane.
The background colors of the sequences reflect the current coloring scheme for the alpha carbon in the Workspace. Each residue in the sequence also has a tooltip that gives the residue name, number, and insertion code.
If the protein has a secondary structure assignment, the assignment is displayed above the sequence, using the following symbols:
- Helices are displayed as tubes, colored red
- Strands are displayed as arrows, colored cyan
- Loops are displayed as a gray line
You can select residues in the sequence and perform actions on these residues.
- To select a single residue, click the residue.
- To add single residues to the selection or remove single residues from the selection, control-click the residues.
- To select a range of residues, either click the first and shift-click the last, or drag over the residues. If the sequence viewer is wrapped, you can drag over multiple rows to select residues in more than one row; the residues that are selected are those in the sequence in which you started dragging.
- To change the selection of a range of residues to the selection, hold down CTRL and drag over the residues.The unselected residues are selected, and the selected residues are deselected.
The selected residues are highlighted in reverse video (as colored letters on a black background), and are marked in yellow in the Workspace. You can also select the missing residue locations in gaps.
You can zoom in on a single residue by middle-clicking on it. This action does not select the residue, but spot-centers on the alpha carbon and fits the residue to the screen. If you middle-click on a gap, the structure is not spot-centered but the view is zoomed in to the residues on either side of the gap.
The sequence viewer and the Workspace are synchronized: selection of atoms in the Workspace selects the corresponding residues in the sequence viewer, and selection of residues in the sequence viewer selects them in the Workspace. Likewise, changing the color scheme in the Workspace changes the color scheme in the sequence viewer, and vice versa. Structural changes, such as mutating a residue, are also reflected in the sequence viewer.
To zoom in on a residue in the Workspace without selecting it, middle-click the residue in the sequence viewer.
The sequence viewer has a shortcut menu (right-click and hold), which you can use to perform various actions on the sequences or the display. The Sequence Viewer Shortcut Menu is displayed when you right-click in the left pane of the sequence viewer, or when you right-click in the right pane of the sequence viewer when there are no residues selected. If you have residues selected, the Sequence Viewer Selection Shortcut Menu is displayed.