Ligand Docking
Glide ligand docking jobs require a set of previously calculated receptor grids and one or more ligand structures. The ligand structures must satisfy the conditions listed in Ligand Preparation for Glide. Information on setting up grid generation jobs is given in Receptor Grid Generation.
Preparation of the ligands before docking is strongly recommended. LigPrep or MacroModel can be used to prepare ligands—see Protein and Ligand Preparation for Glide for more information.
If a correct Lewis structure cannot be generated for a ligand, it is skipped by the docking job. Glide also automatically skips ligands containing unparametrized elements, such as tin, or atom types not supported by the OPLS force fields, such as explicit lone pair “atoms.”
Detailed descriptions of the Ligand Docking panel in Maestro and each of its tabs, including instructions for applying Glide constraints, using extra-precision Glide docking (Glide XP), and distributed processing, are given in the Ligand Docking Panel DEPRECATED help topic. To open the Ligand Docking panel, click the Tasks button and browse to Receptor-Based Virtual Screening → Ligand Docking (Deprecated).
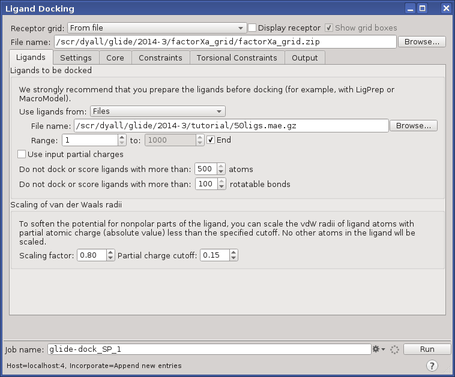
This section contains the following topics: