Nanoreactor
Tutorial Created with Software Release: 2026-1
Topics: Catalysis & Reactivity , Energy Capture & Storage
Methodology: Molecular Quantum Mechanics
Products Used: AutoTS , Jaguar , MS Maestro , MS Reactivity
|
1.71GB |
This tutorial is written for use with a 3-button mouse with a scroll wheel.
Words found in the Glossary of Terms are shown like this: Workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Abstract:
In this tutorial, we will learn to use the Nanoreactor panel to predict thermal decomposition products of a molecule without any prior knowledge.
Tutorial Content
1. Introduction to Nanoreactor
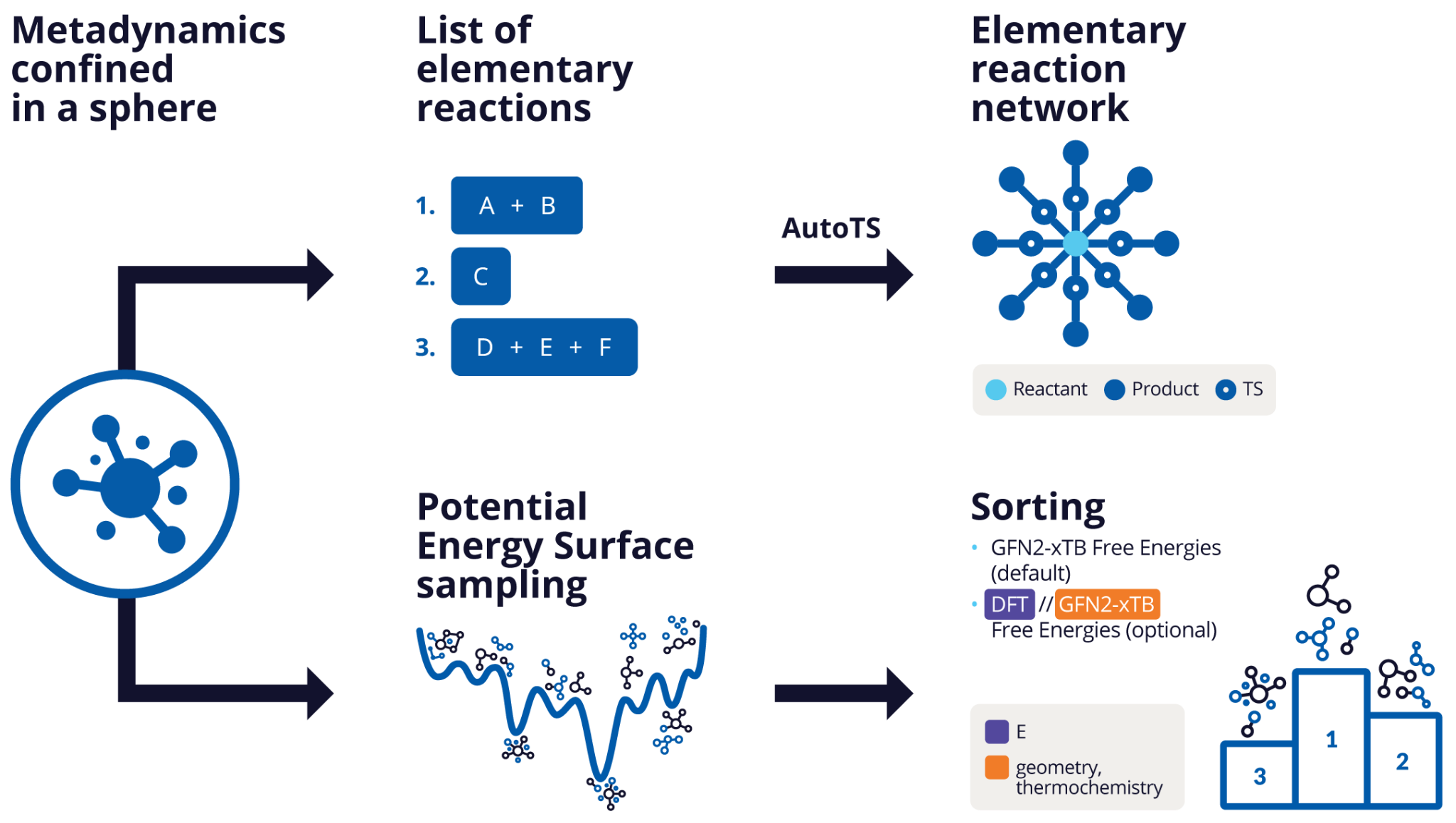
Automated reactions discovery lies at the heart of predictive chemistry, enabling chemists to design chemical processes that are smarter, faster, cleaner, and more efficient. Nanoreactor – Elementary Reaction Network (ERN), a tool by Schrödinger, automates the identification of energetically relevant elementary reactions starting from a known local minimum on the GFN2-xTB potential energy surface, using root-mean-square deviation (RMSD)-based metadynamics. When combined with AutoTS, it enables the efficient generation and refinement of an Elementary Reaction Network (ERN) using xTB, MLFF-corrected or DFT-corrected free energy surface. Complementing this, Potential Energy Surface Sampling-Sorting (PSS) enhances Nanoreactor’s capabilities by systematically exploring and ranking minima states on GFN2-xTB (or DFT, or MLFF) methods. Since chemical reactions tend to follow the downhill path on the free energy surface, this feature focuses on pinpointing the most probable final products.
In this tutorial, we will use the Nanoreactor and Nanoreactor Results panels to sample relevant elementary reactions for 3-hydroperoxypropanal, and the PSS of acetic acid and benzene to investigate its thermal decomposition products.
Nanoreactor is an integration and extension of work done by multiple individuals. Please cite appropriate references in any publication that contains results from the use of this panel. Specifically, papers by Koerstz et. al., Rasmussen et. al., and Grimme should be referenced. These papers are also linked in our For further reading section.
2. Creating and Saving Projects
At the start of the session, change the file path to your chosen Working Directorythe location where files are saved in MS Maestro to make file navigation easier. Each session in MS Maestro begins with a default Scratch Projecta temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project, which is not saved. A MS Maestro project stores all your data and has a .prj extension. A project may contain numerous entries corresponding to imported structures, as well as the output of modeling-related tasks. Once a project is saved, the project is automatically saved each time a change is made.
Structures can be built in MS Maestro or can be imported using File > Import Structures (or drag-and-dropped), and are added to the Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data. The Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion is located to the left of the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed. The Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data can be accessed by Ctrl+T (Cmd+T) or Window > Project Table if you would like to see an expanded view of your project data.
- Double-click the Materials Science icon
- (No icon? See Starting Maestro)
- Go to File > Change Working Directory
- Find your directory, and click Choose
- Pre-generated files are included for running jobs or examining output. Download the zip file here: schrodinger.com/sites/default/files/s3/release/current/Tutorials/zip/nanoreactor.zip
- After downloading the zip file, unzip the contents in your Working Directory for ease of access throughout the tutorial
- Go to File > Save Project As
- Change the File name to nanoreactor_tutorial, click Save
- The project is now named
nanoreactor_tutorial.prj
- The project is now named
We will proceed to import the systems that we will use in the subsequent sections: 3-hydroperoxypropanal, acetic acid, and benzene. If you would prefer to draw or build these components yourself using the 2D Sketcher, feel free to do so following similar steps outlined in the Introduction to Materials Science Maestro tutorial. Otherwise:
- Go to File > Import Structures
- Navigate to where you have downloaded the provided files (presumably in your working directorythe location where files are saved), and choose the
inputs.maegzfile - Click Open
- A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion containing two entries
3. Performing a Nanoreactor Elementary Reaction Network Calculation
In this section, we will explore the first step of thermal decomposition of 3-hydroperoxypropanal (3hyd) using the Nanoreactor panel.
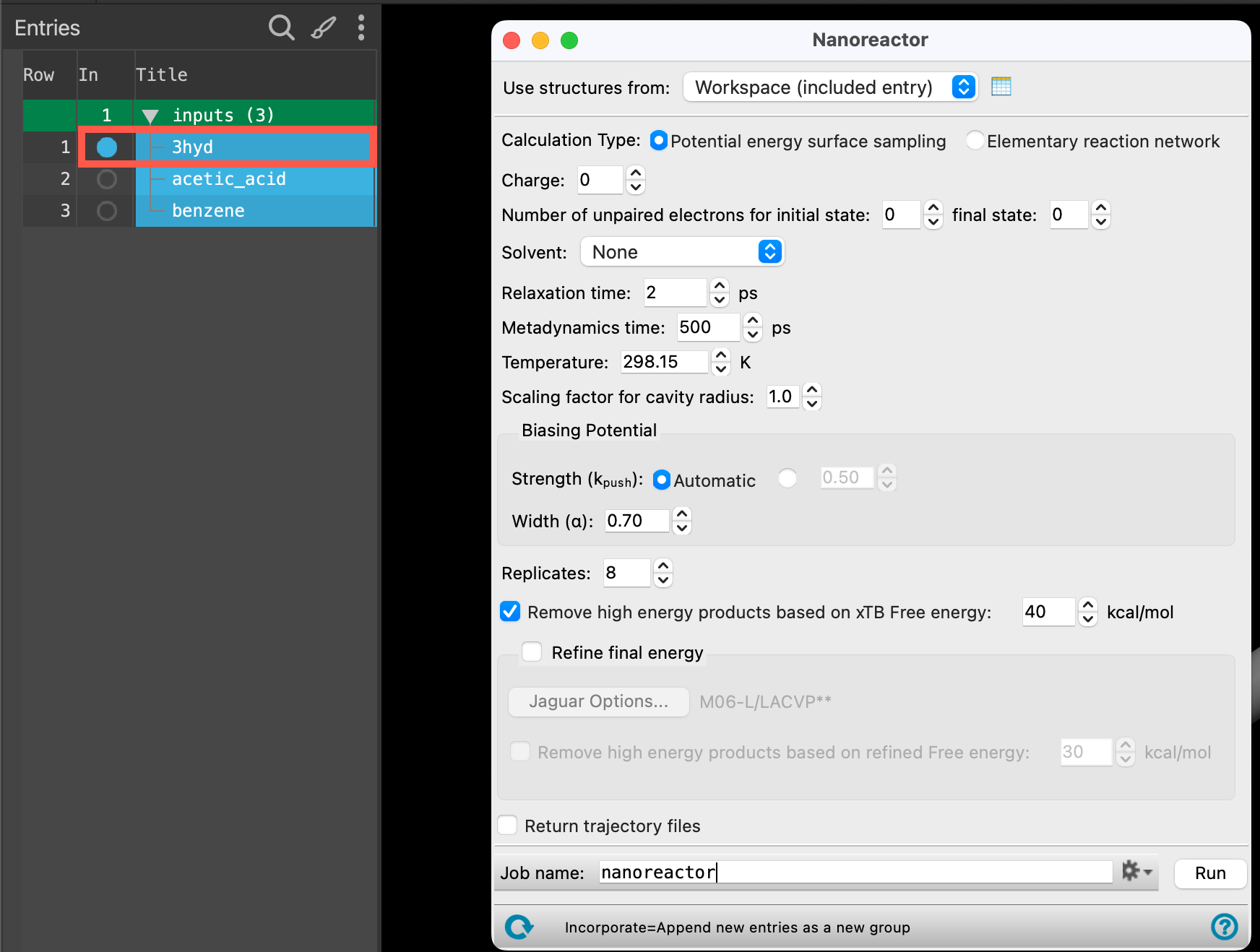
- With the 3hyd entry selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Nanoreactor Calculations
- The Nanoreactor panel opens
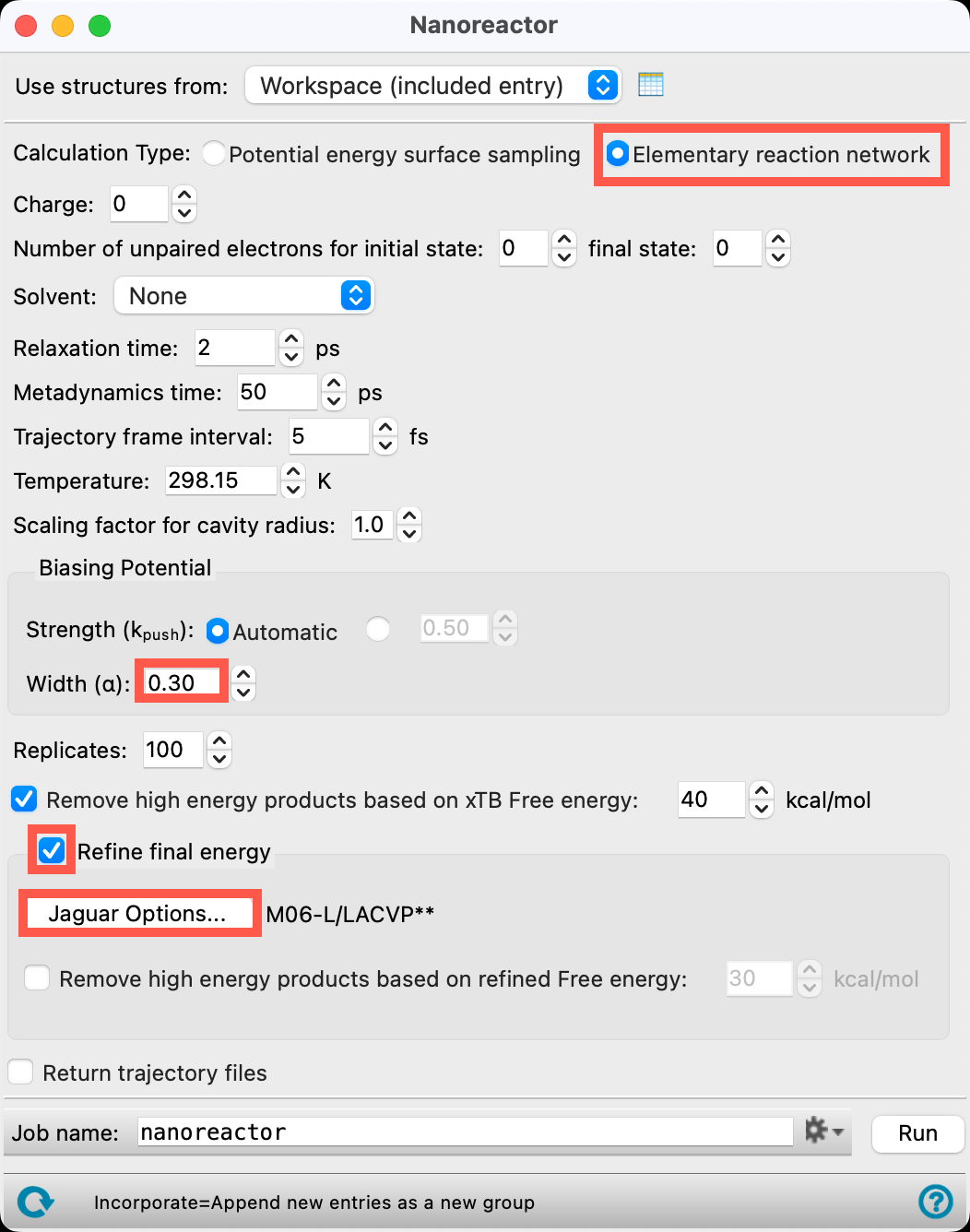
- Set the Calculation Type to Elementary reaction network
- Choosing this calculation type allows for the transition state to be calculated (though we will not be selecting this feature in this particular example)
- Set the Width to 0.30
- Check Refine final energy
- This setting introduces a DFT calculation on nanoreactor products, specifically electronic energy is calculated with DFT and thermal corrections are calculated with GFN2-xTB. Be aware that adding this parameter might significantly increase computation time, especially for large molecules
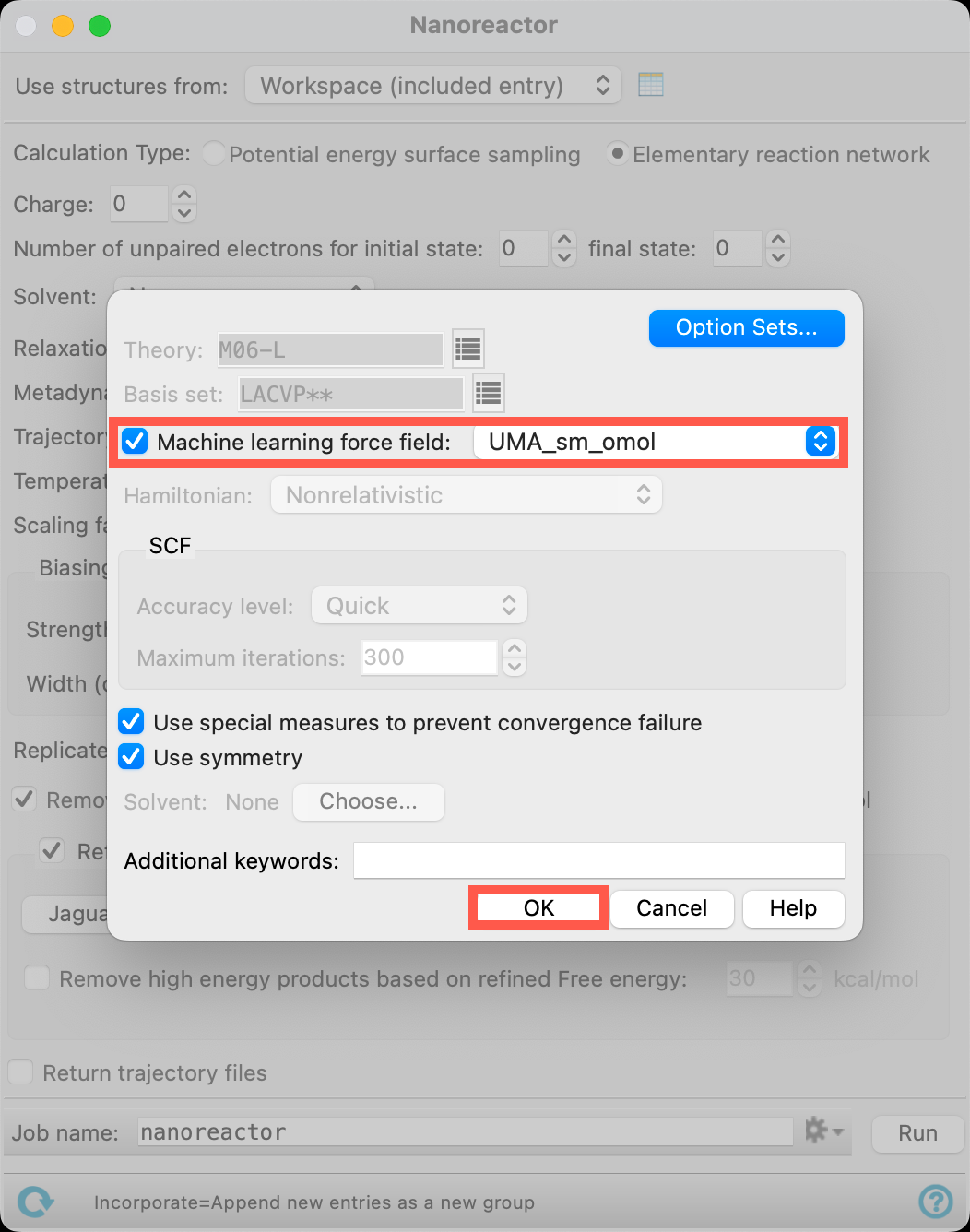
- Open the Jaguar Options panel
- Select Machine learning force field to be UMA_sm_omol
- Click OK to close the panel
Note: If you are interested in using Machine Learning Force Fields (MLFF) but the necessary options are not visible, please contact your Account Manager. Additional information about MLFF, including details on available models, is available in the help documentation and on our website.
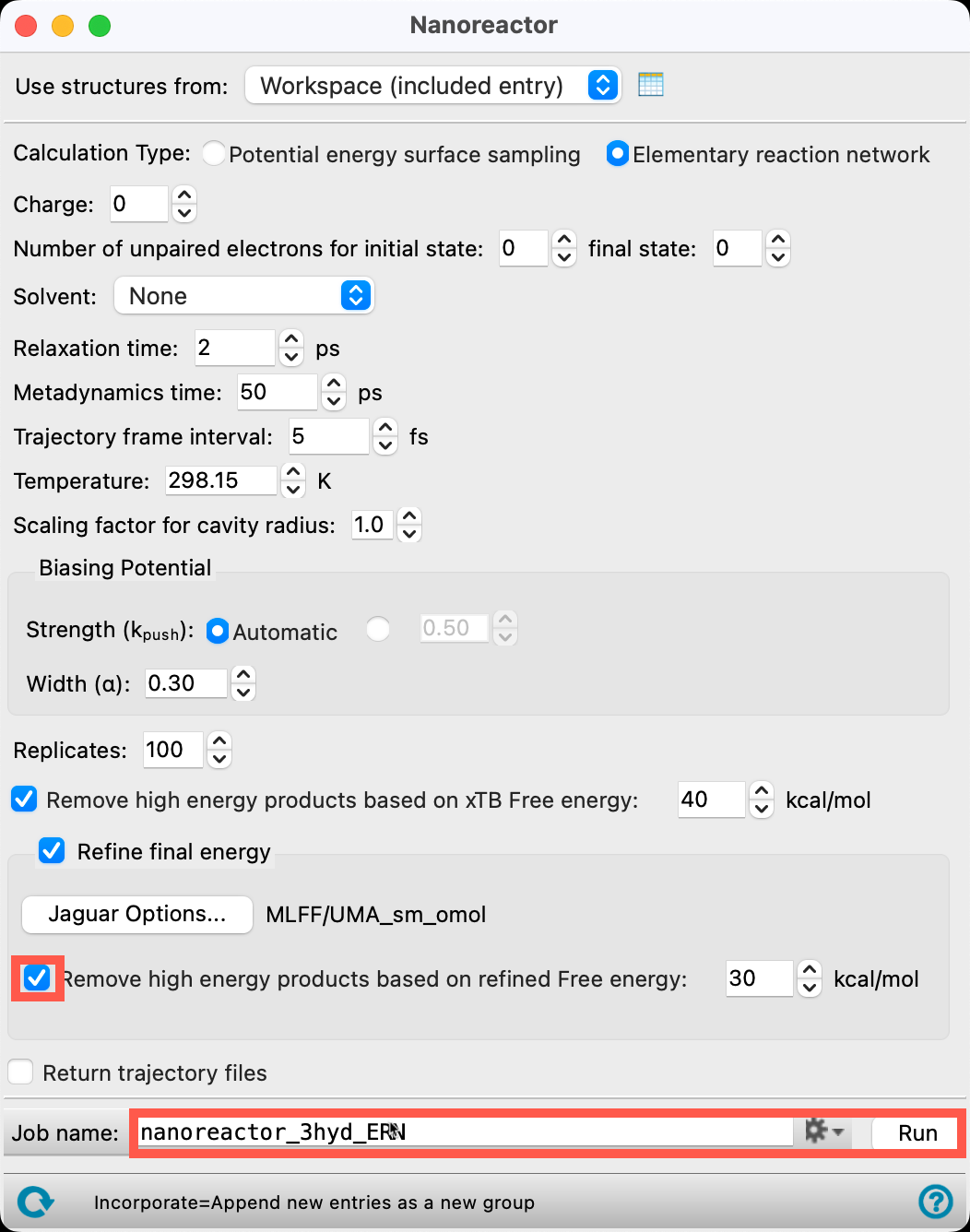
- Check Remove high energy products based on refined Free energy
- Change the Job name to nanoreactor_3hyd_ERN
-
Adjust the job settings (
 ) as needed
) as needed
- This job requires a CPU host. The job can be completed in 15 minutes on 100 CPUs.
- If you would like to run the job yourself, click Run. Otherwise, we will proceed with imported results files in the next section
- Close the Nanoreactor panel
Let’s take a closer look at the features in this panel. More detailed information can be found by clicking the blue question mark ( ) located at the bottom right corner of the panel or by visiting the related documentation page.
) located at the bottom right corner of the panel or by visiting the related documentation page.
- The Charge textbox allows the user to specify the overall charge of the system, while it can be positive, negative, or zero, the net charge does need to be an integer.
- The initial and final number of unpaired electrons allows the user to specify how many unpaired electrons there should be in the initial state and in the final state of products, if any. The GFN2-xTB method does not properly distinguish between ⟨S2⟩ spin eigenstates and only conserves a total spin.
- A solvent modeled by the Poisson-Boltzmann model can be selected from the Solvent option menu, if None is selected the calculation is run in the gas phase.
- The Relaxation time is the total simulation time for the molecular dynamics pre-equilibration.
- The Metadynamics time is the total simulation time for the metadynamics calculation.
- The Temperature describes bath temperature.
- There are three hyperparameters for metadynamics:
- s = cavity radius, generally smaller value (wrt default) = more isomerization is favored smaller value (wrt default) = faster events for bimolecular and higher molecular reactions. If you run bimolecular and higher molecular reactions, we suggest setting multiplication factor of 0.8 with respect to the default cavity radius determined by the program, see replicate_1_md-out.log file by searching “spherical wallpotential with radius”
- kpush = strength of the biasing potential (defines how strongly the system should be “pushed” out of previously visited structures) too small value = we approach unbiased MD limit = no reaction takes place too high value = chemical irrelevant reactions might occur
- α = width of the biasing potential (determines how far from the saved structures the system is still affected by the biasing potential) too small value = means that very large changes in the structure are needed to being affected by the biasing potential too high value = narrow biasing potential meaning that the system can “escape” the biasing potential by smaller changes to the structure (e.g. conformational changes)
- The Replicates textbox can be used to specify the number of trajectories used in each molecular dynamics, metadynamics, and optimization calculation.
- Removing products with a relative energy greater than the given threshold, relative to the lowest xTB Free energy structure, allows the user to remove high-energy products generated by the panel before the structures are refined by a quantum mechanical calculation. It results in removing physically unlikely structures from the simulation. Knowing the energies of the expected decomposition products can help determine what this threshold should be.
- The Refine final energy option allows the user to improve the accuracy of the final free energy calculation using single-point DFT (or MLFF) refinement; the specifics of the calculation can be changed in the Jaguar Options menu.
- The Calculate transition states option allows the user to calculate the transition states with AutoTS using single-point DFT (or MLFF) refinement; the specifics of the calculation can be changed in the AutoTS Options menu.
4. Analyzing Nanoreactor Elementary Reaction Network Results
In this section, we will view and analyze the ERN of the 3-hydroperoxypropanal calculation with the Nanoreactor Results panel.
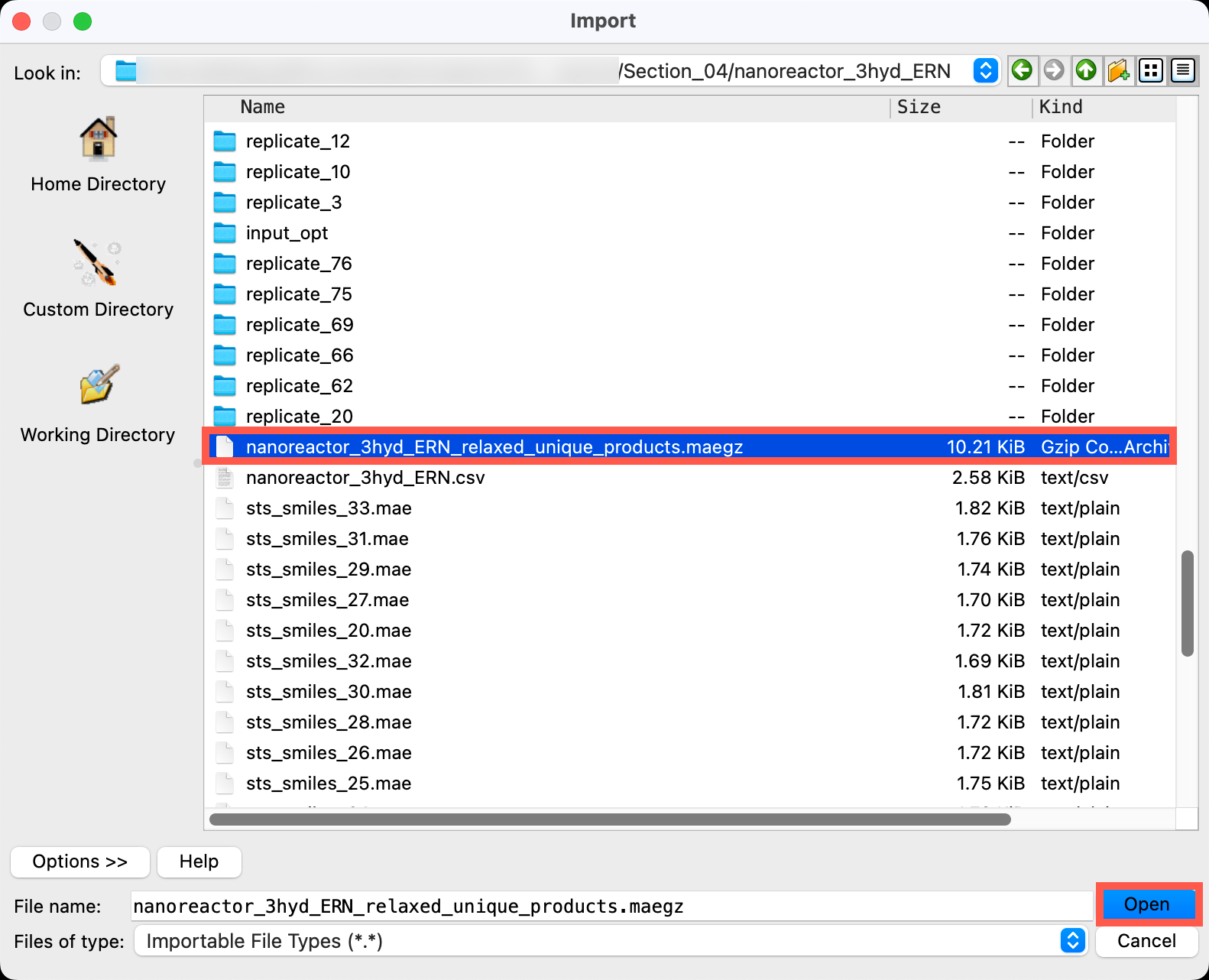
- Go to File > Import Structures
- Navigate to the provided files and choose
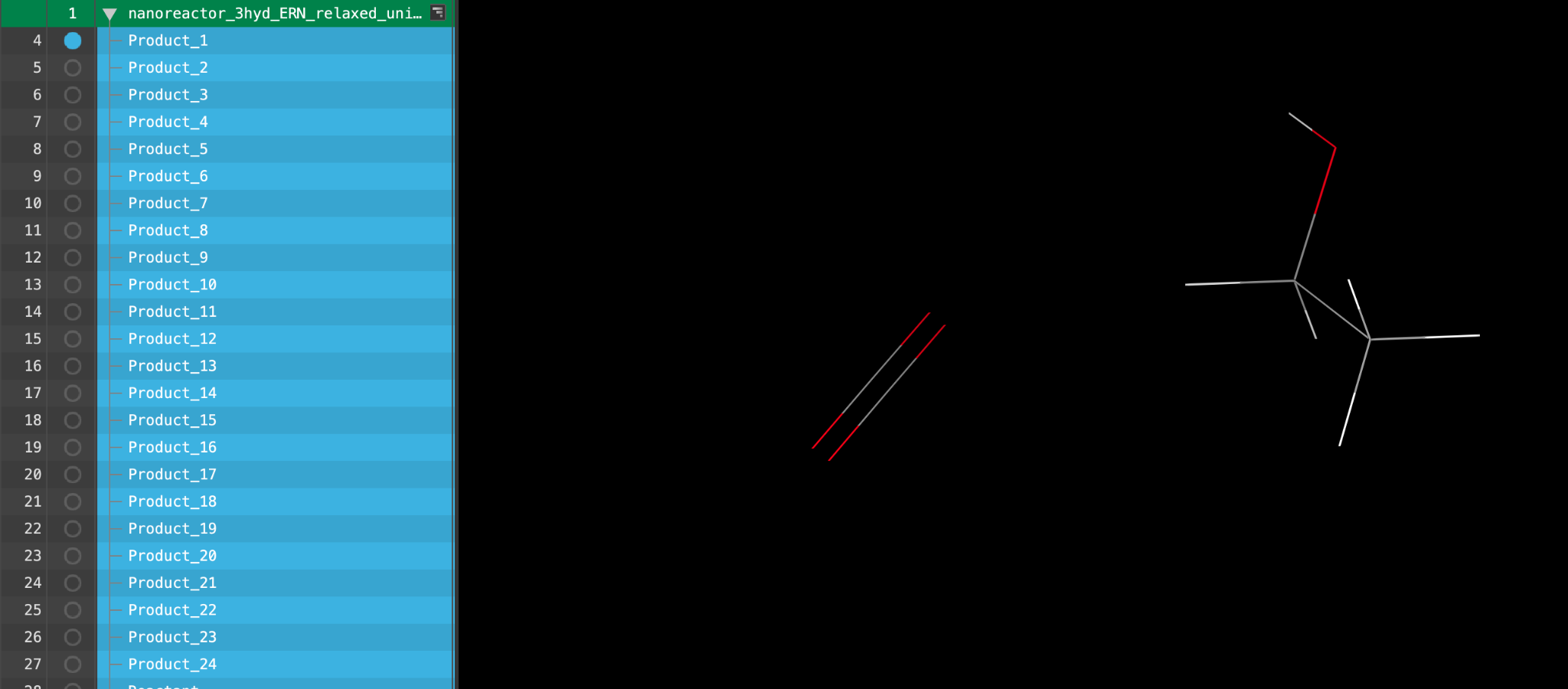
Section_04 > nanoreactor_3hyd_ERN > nanoreactor_3hyd_ERN_relaxed_unique_products.maegz - Click Open
A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion. The group contains 32 entries.
Note that if you performed the calculation yourself, the exact number of output structures may vary due to randomness in the metadynamics simulations. The variation (should) mostly come from the high-energy products.
Feel free to visualize any of the entries in the workspace.
Quantitative analysis can be performed in the results panel:
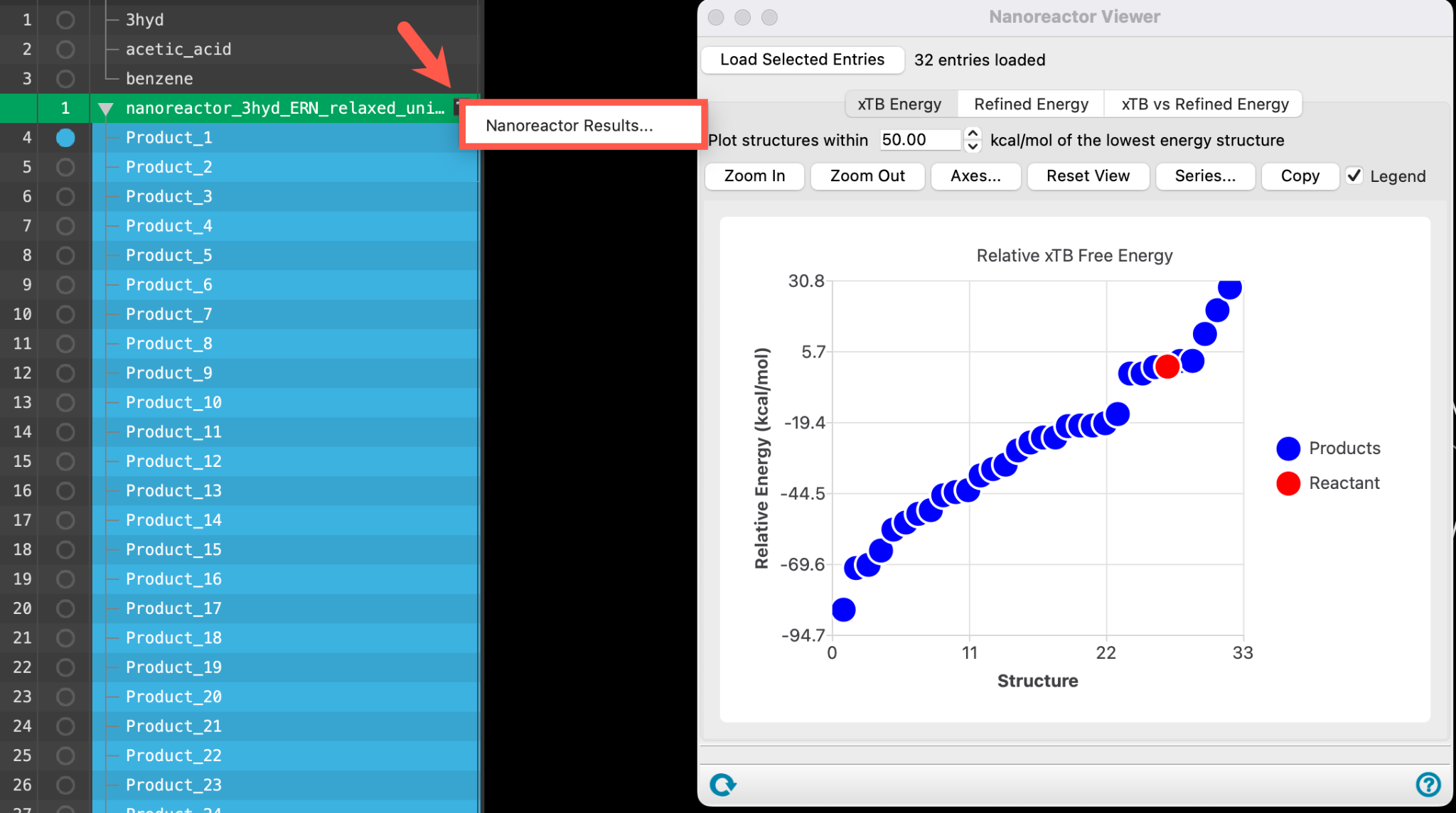
- Use the WAM (workflow action menu) button (
 ) to access the Nanoreactor Results panel
) to access the Nanoreactor Results panel
- Alternatively, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Nanoreactor Results and use the Load selected entries button to load the entire entry group into the panel
- The Nanoreactor Results panel opens
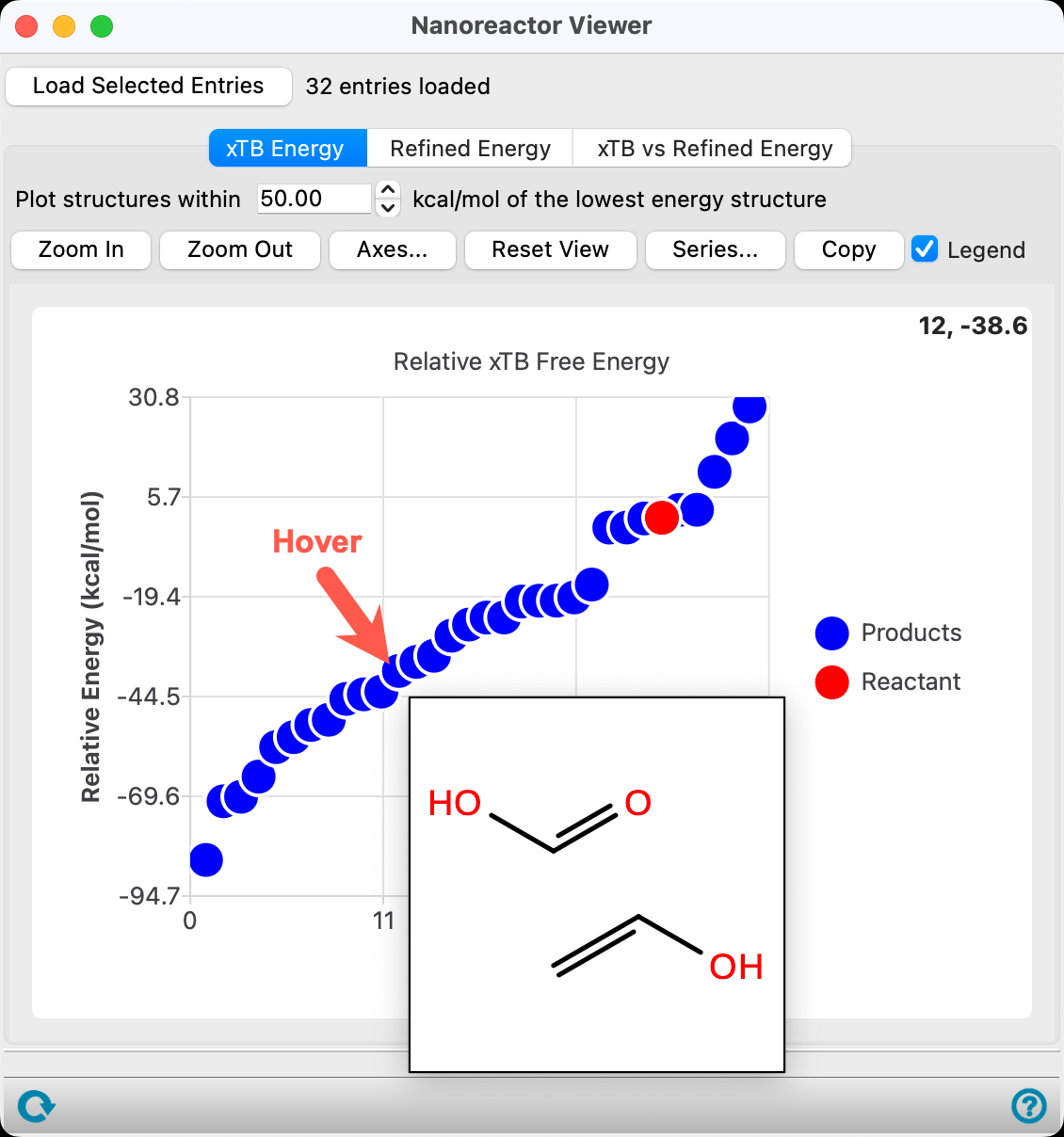
The energy is calculated with respect to the most thermodynamically stable species taken as 0.
- For confirmation, hover the mouse over the lowest energy blue data point to visualize a 2D sketch of the products
- Close the Nanoreactor Viewer panel
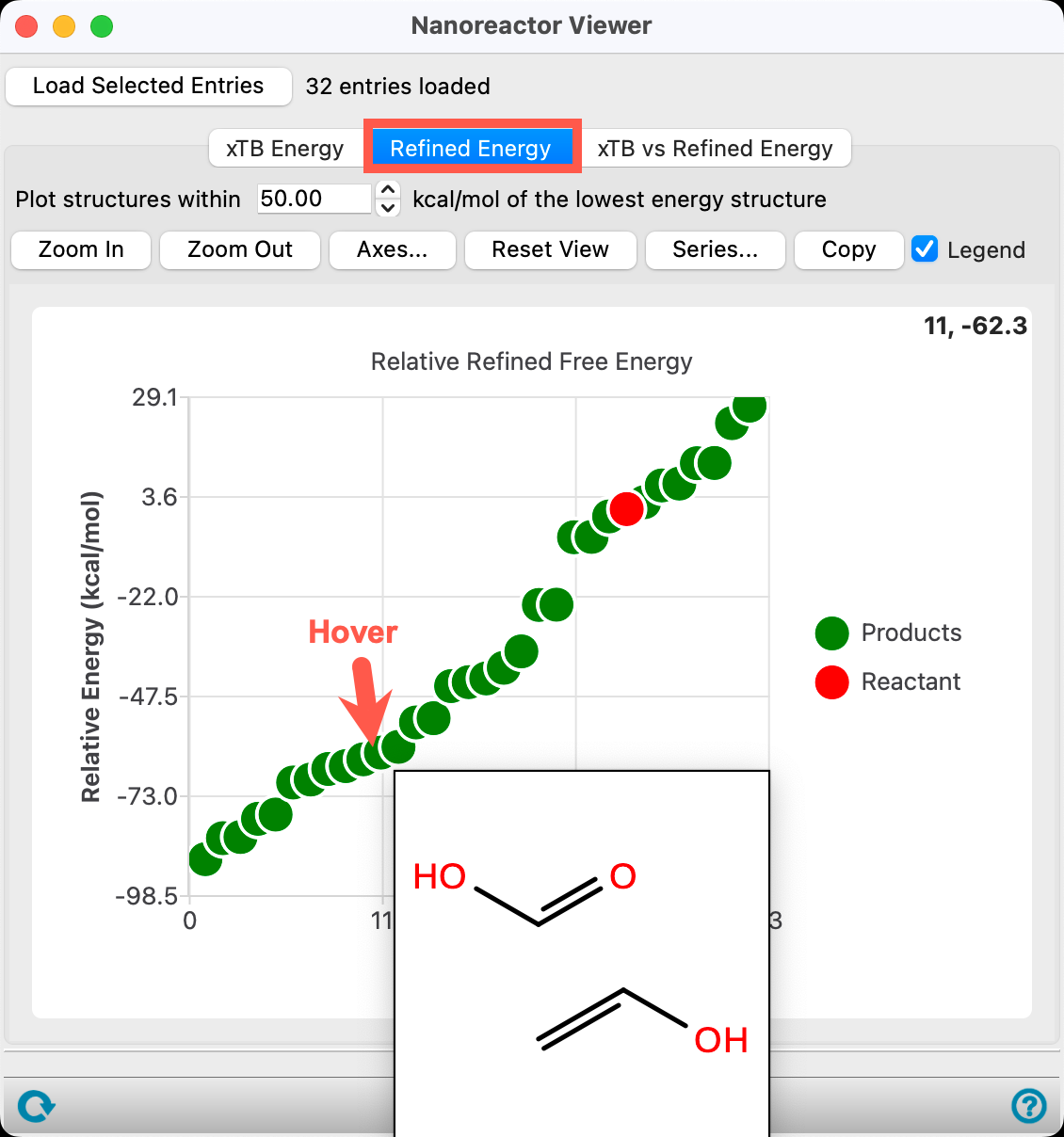
- Go to the Refined Energy tab
- Since MLFF methods were used in the refinement process, the DFT//xTB tab displays the MLFF//xTB energy
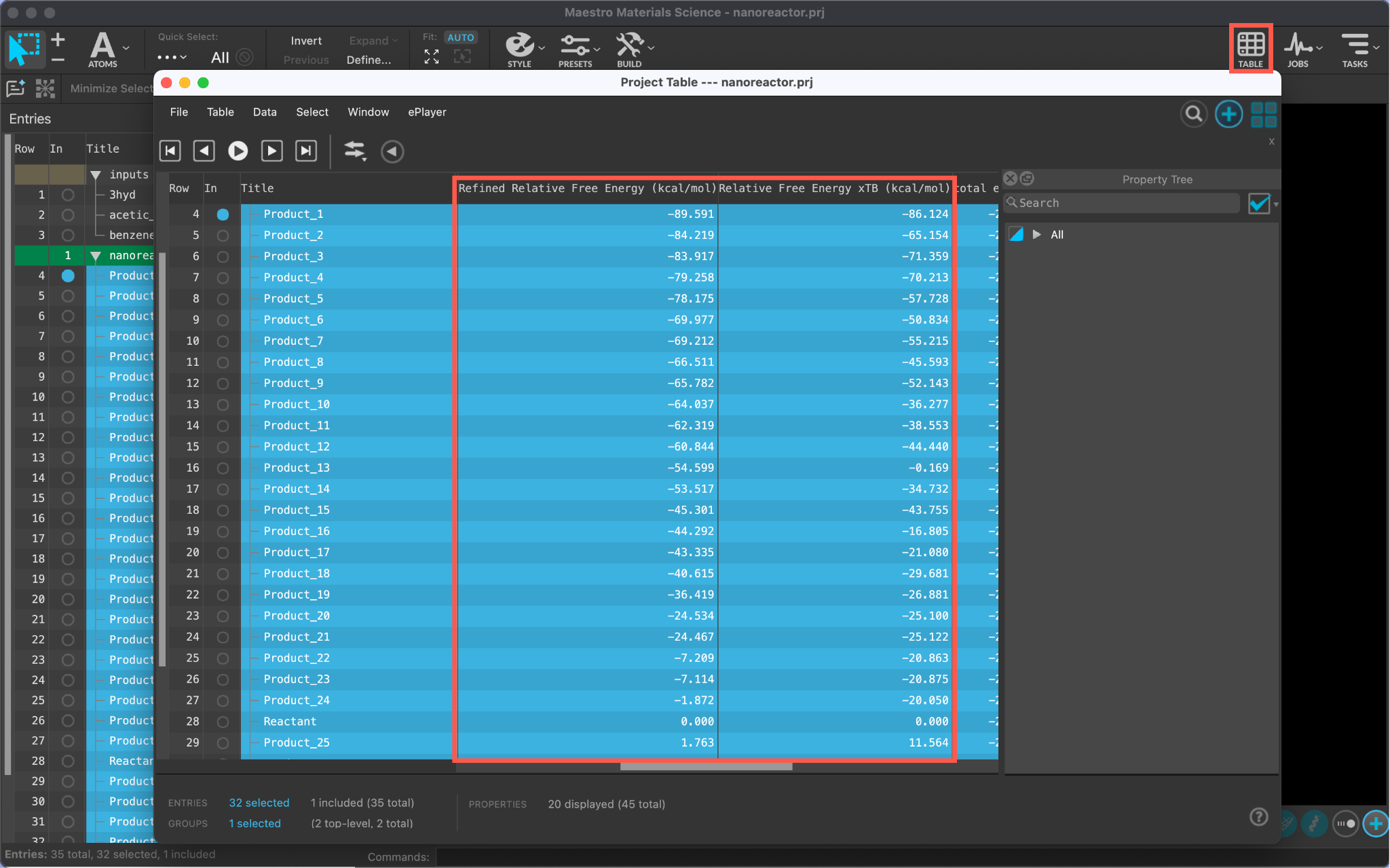
The relative free energy values can be viewed in the project table.
- Open the TABLE
- Scroll over to view the Refined Relative Free Energy and Relative Free Energy xTB values
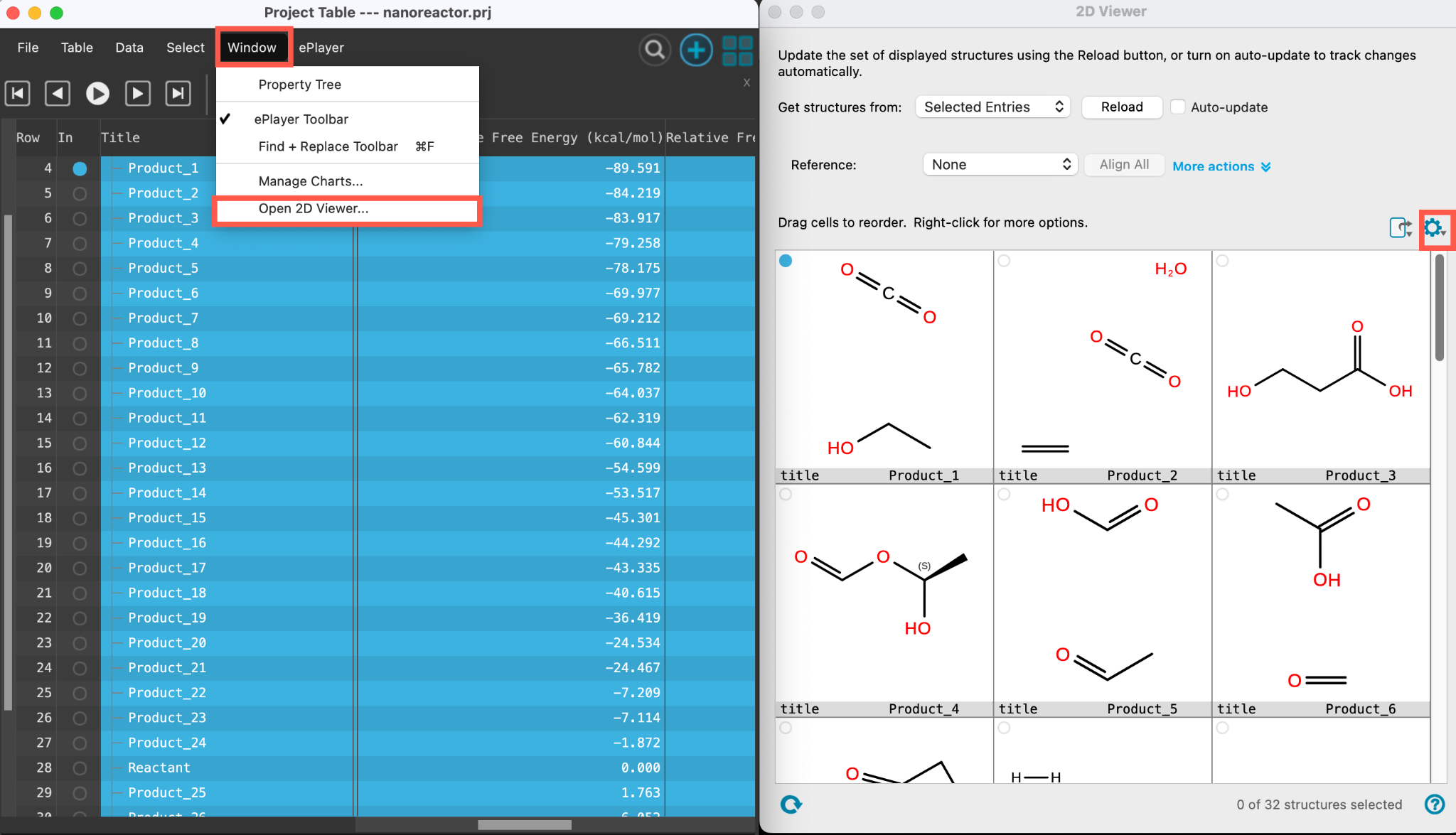
- Go to Window > Open 2D Viewer
- The 2D Viewer panel opens
- To view more structures simultaneously, adjust the number of columns in the settings menu, shown by the cog
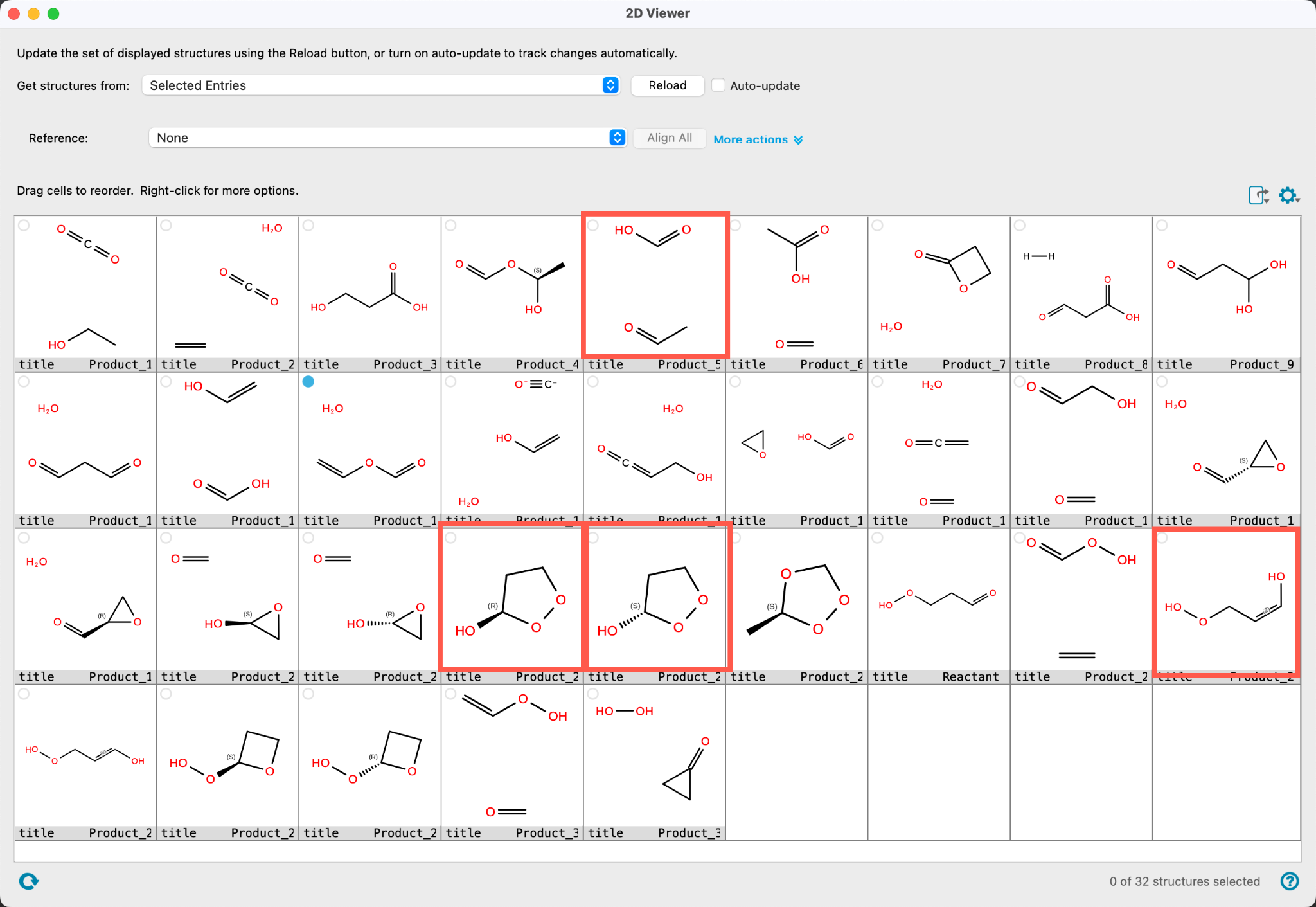
The 2D Viewer panel shows all 32 product structures. Explore the products and close the panel once finished.
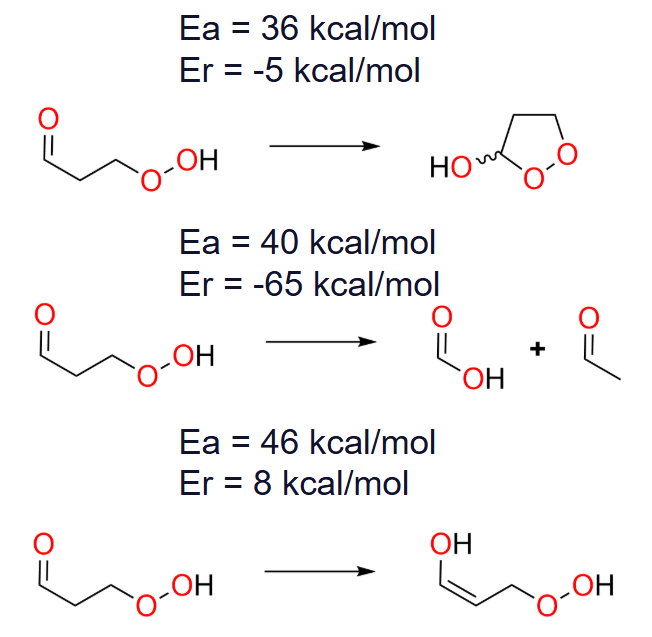
Based on an exhaustive search algorithm, 3-hydroperoxypropanal exhibits 75 unimolecular elementary reactions, assuming no more than two bonds being broken or formed, see References for more details specifically the work by Koerstz et. al. The four identified products by Nanoreactor-ERN include the three lowest-barrier elementary reactions from the list of 75 products found in the exhaustive search by Grambow et. al. This nicely demonstrates that the semiempirical method is both accurate and fast enough to automatically pinpoint promising candidates for DFT refinement for the low barrier reactions of 3-hydroperoxypropanal.
Shown below are the three lowest-barrier unimolecular elementary reactions for 3-hydroperoxypropanal, see the References for more details specifically the work by Koerstz et. al:
Note that these Nanoreactor results do not provide kinetic information per se, but the means to obtain kinetic information quickly and in an unbiased manner. Due to limitations in the algorithm, not all products generated in an ERN run necessarily correspond to true elementary steps. This must be confirmed through transition state analysis and subsequent intrinsic reaction coordinate (IRC) calculations with AutoTS.
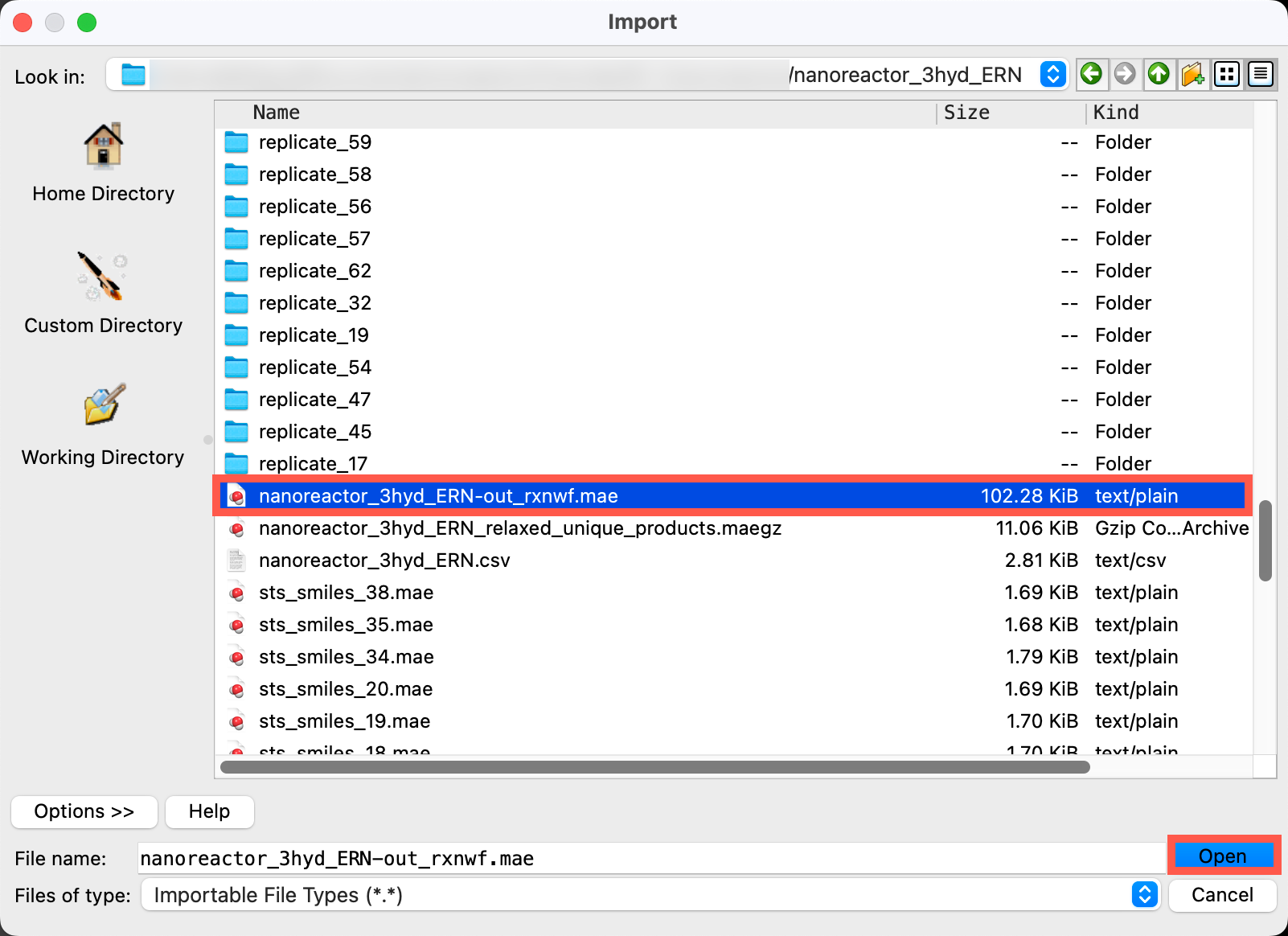
- Go to File > Import Structures
- Navigate to the provided files and choose
Section_04 > nanoreactor_3hyd_ERN > nanoreactor_3hyd_ERN-out_rxnwf.mae - Click Open
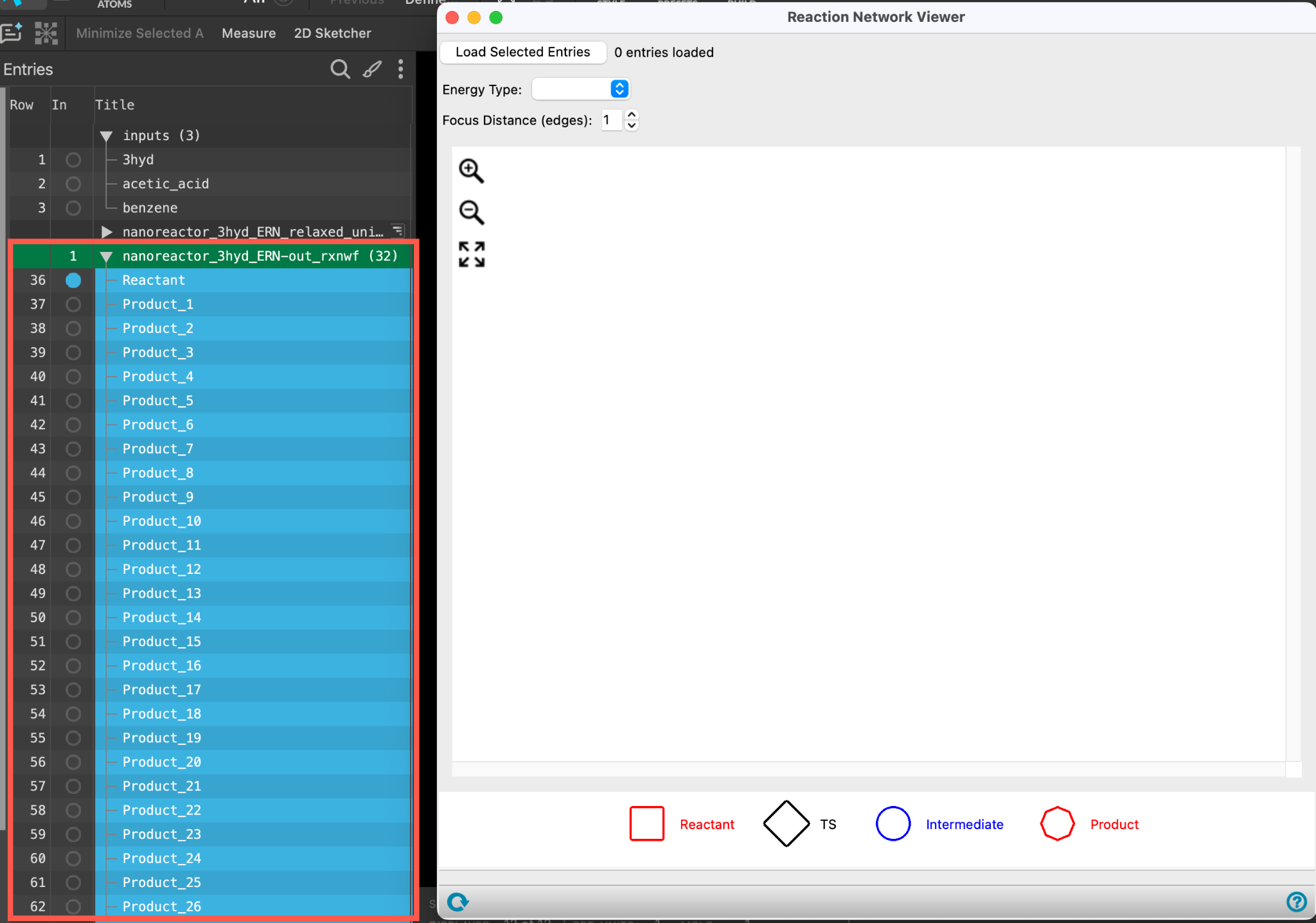
- With the nanoreactor_3hyd_ERN-out_rxnwf entry selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Reaction Network Viewer
- The Reaction Network Viewer panel opens
- Click Load Selected Entries
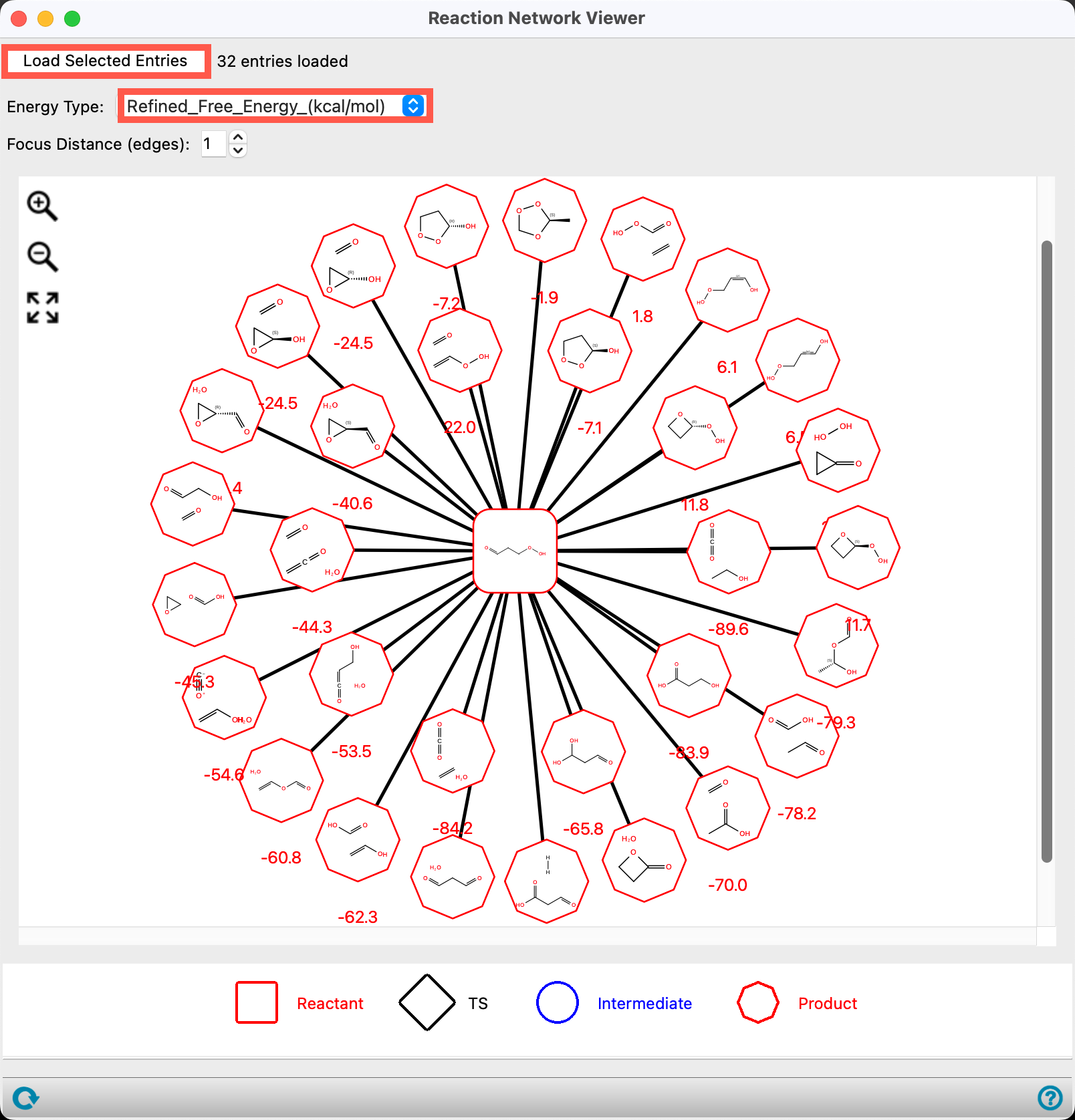
- Set the Energy Type to Refined_Free_Energy_(kcal/mol)
The reaction network visualization shows 3-hydroperoxypropanal as the central reactant, with its various products displayed in the surrounding octagons.
5. Performing a Nanoreactor Potential Energy Surface Sampling Calculation
This section will examine the PSS of acetic acid using the Nanoreactor panel. Following this, the process for performing a PSS for a more complex system, benzene, will be demonstrated in Section 7.
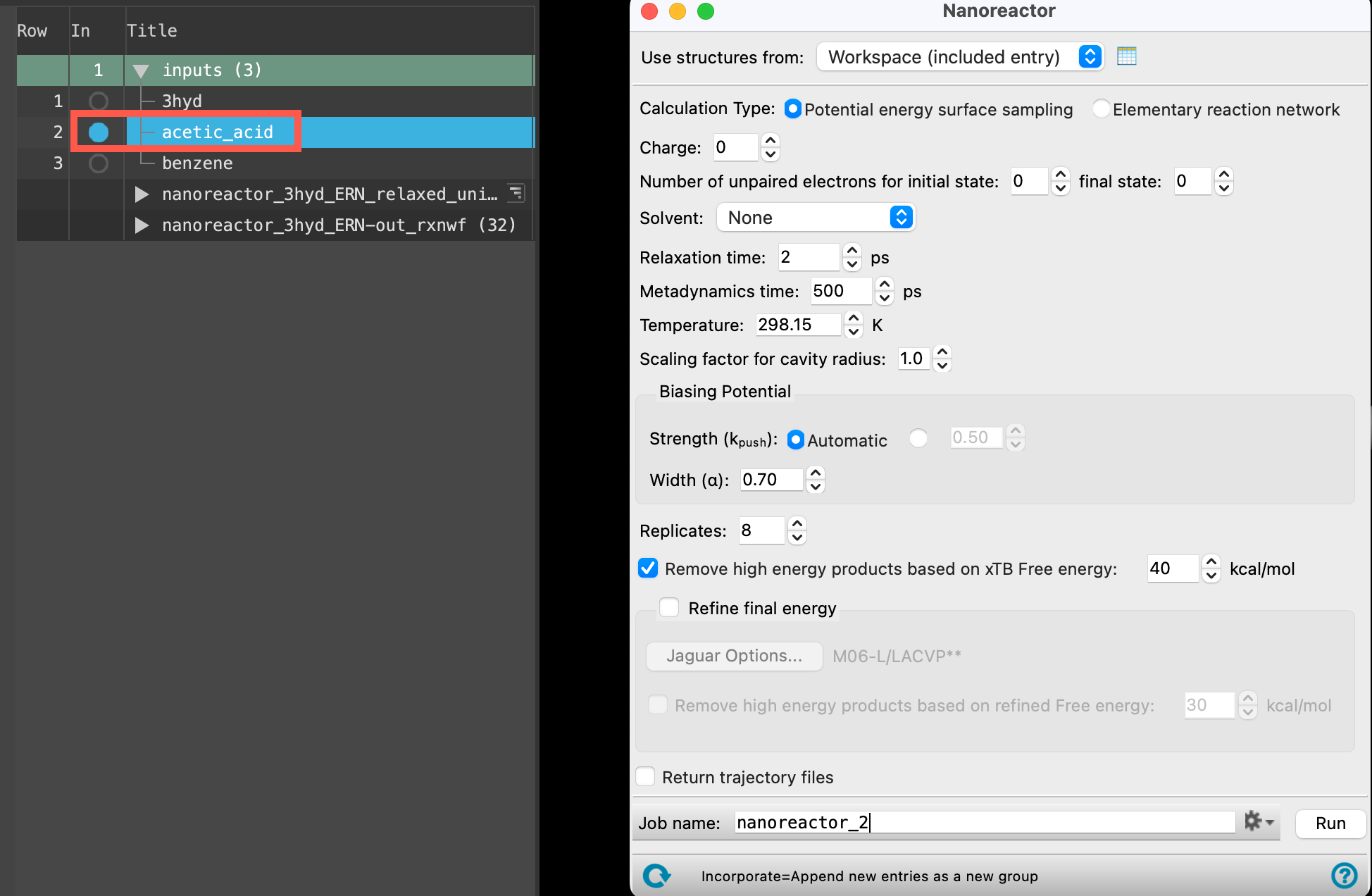
- With the acetic_acid entry selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Nanoreactor Calculations
- The Nanoreactor panel opens
- Reset
 the panel
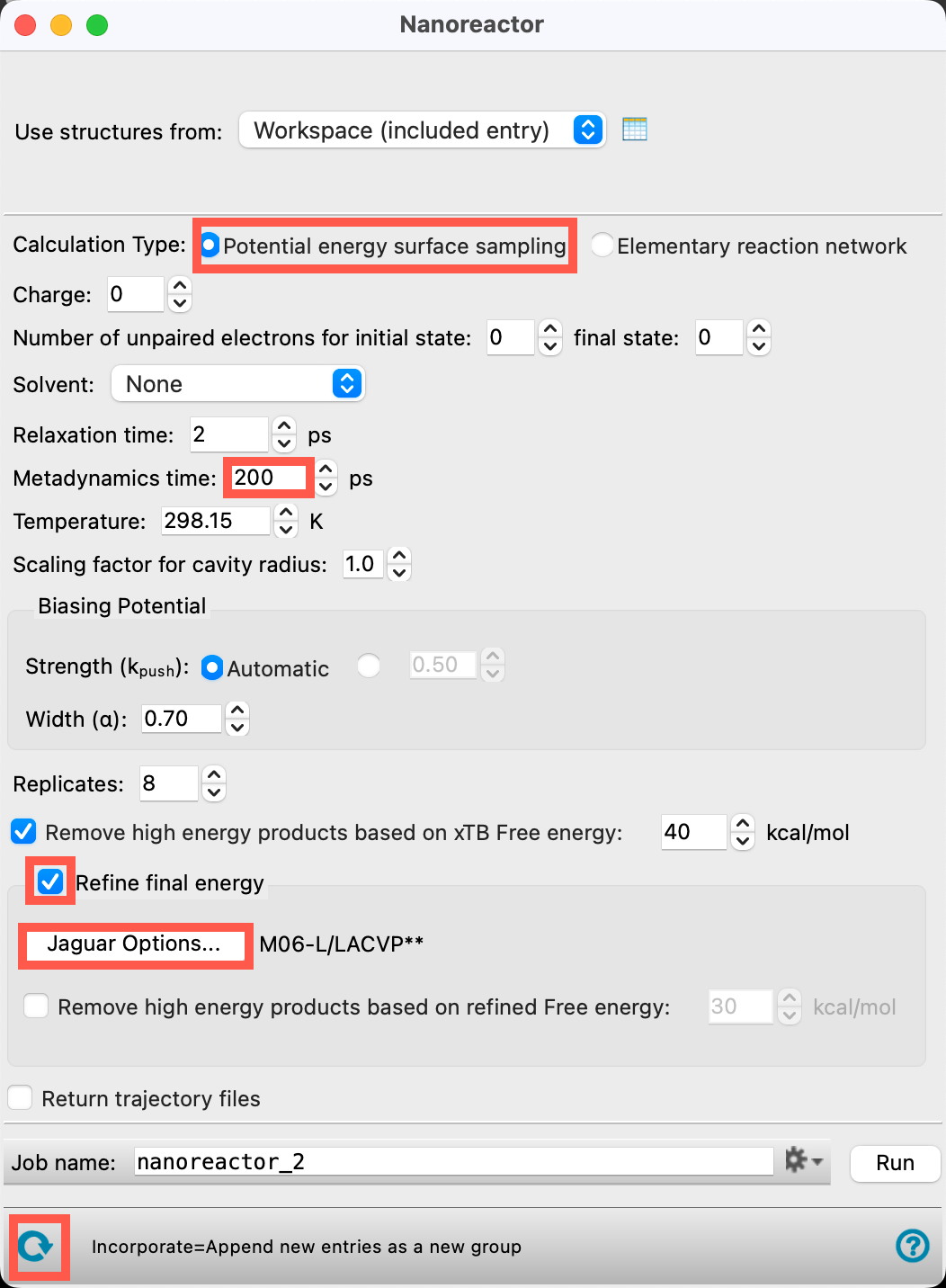
the panel - Ensure that Potential energy surface sampling is selected for the Calculation Type
- Change the Metadynamics time to 200 ps
- This system is small so increasing the relaxation time of the molecular dynamics simulations will not increase the total time the calculation takes by an exorbitant amount
- Check Refine final energy
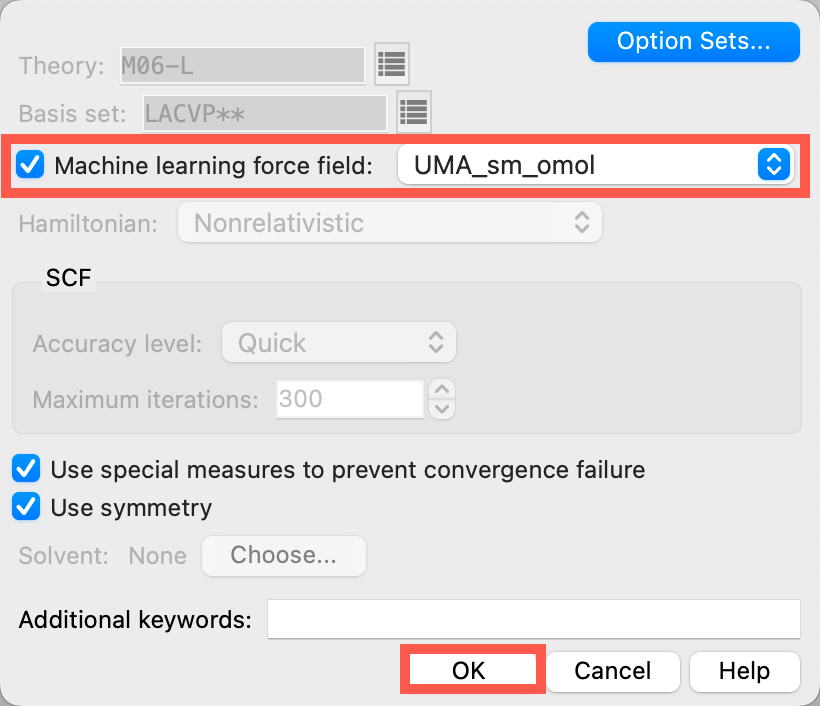
- Open the Jaguar Options panel
- Select Machine learning force field to be UMA_sm_omol
- Click OK to close the panel
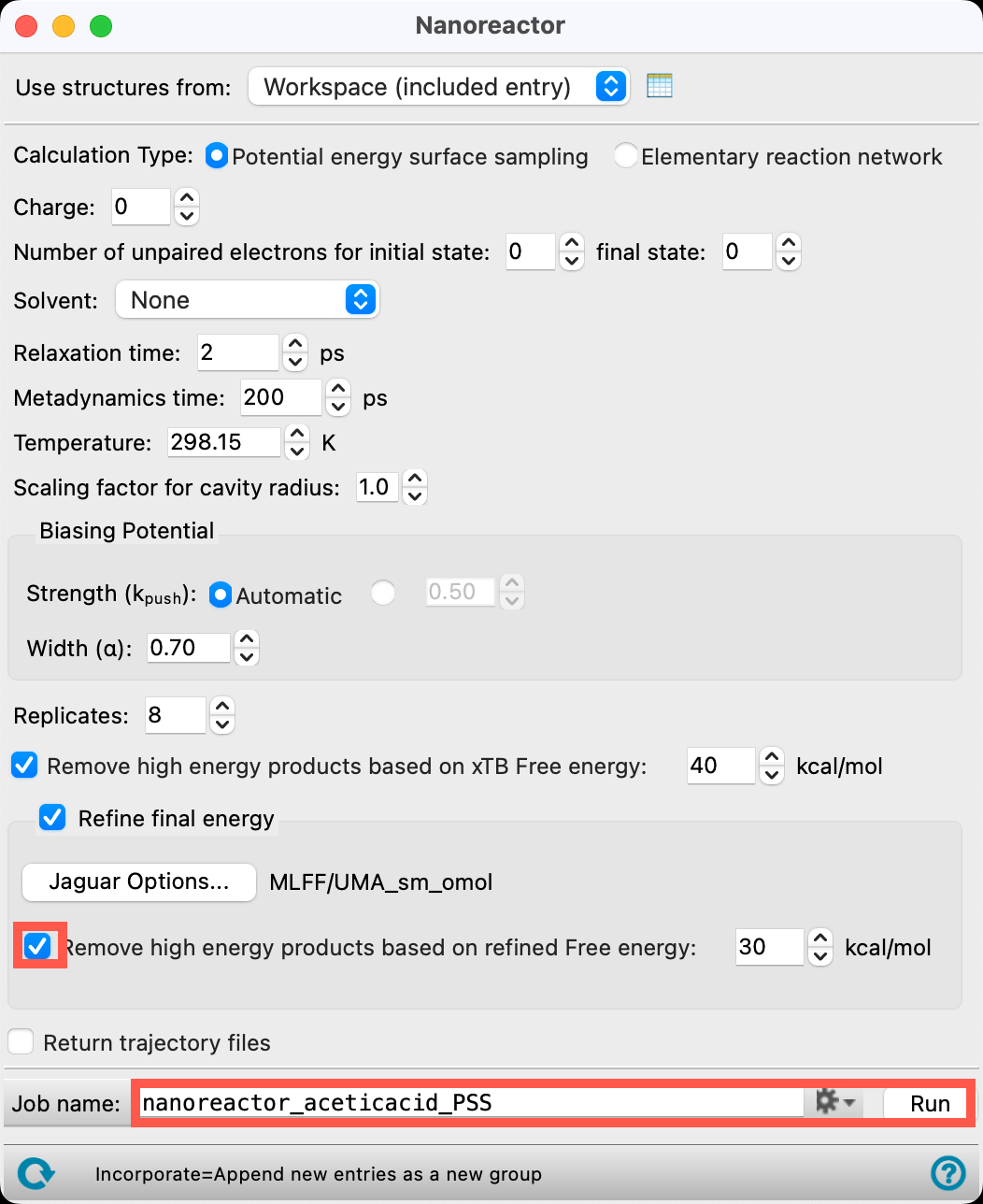
- Check Remove high energy products based on refined Free energy
- Change the Job name to nanoreactor_aceticacid_PSS
-
Adjust the job settings (
 ) as needed
) as needed
- This job requires a CPU host. The job can be completed in about 2 hours on 8 CPUs
- If you would like to run the job yourself, click Run. Otherwise, we will proceed with imported results files in the next section
- Close the Nanoreactor panel
6. Analyzing Nanoreactor Potential Energy Surface Sampling Results
In this section, we will view and analyze the results from the decomposition of acetic acid calculation with the Nanoreactor Results panel.
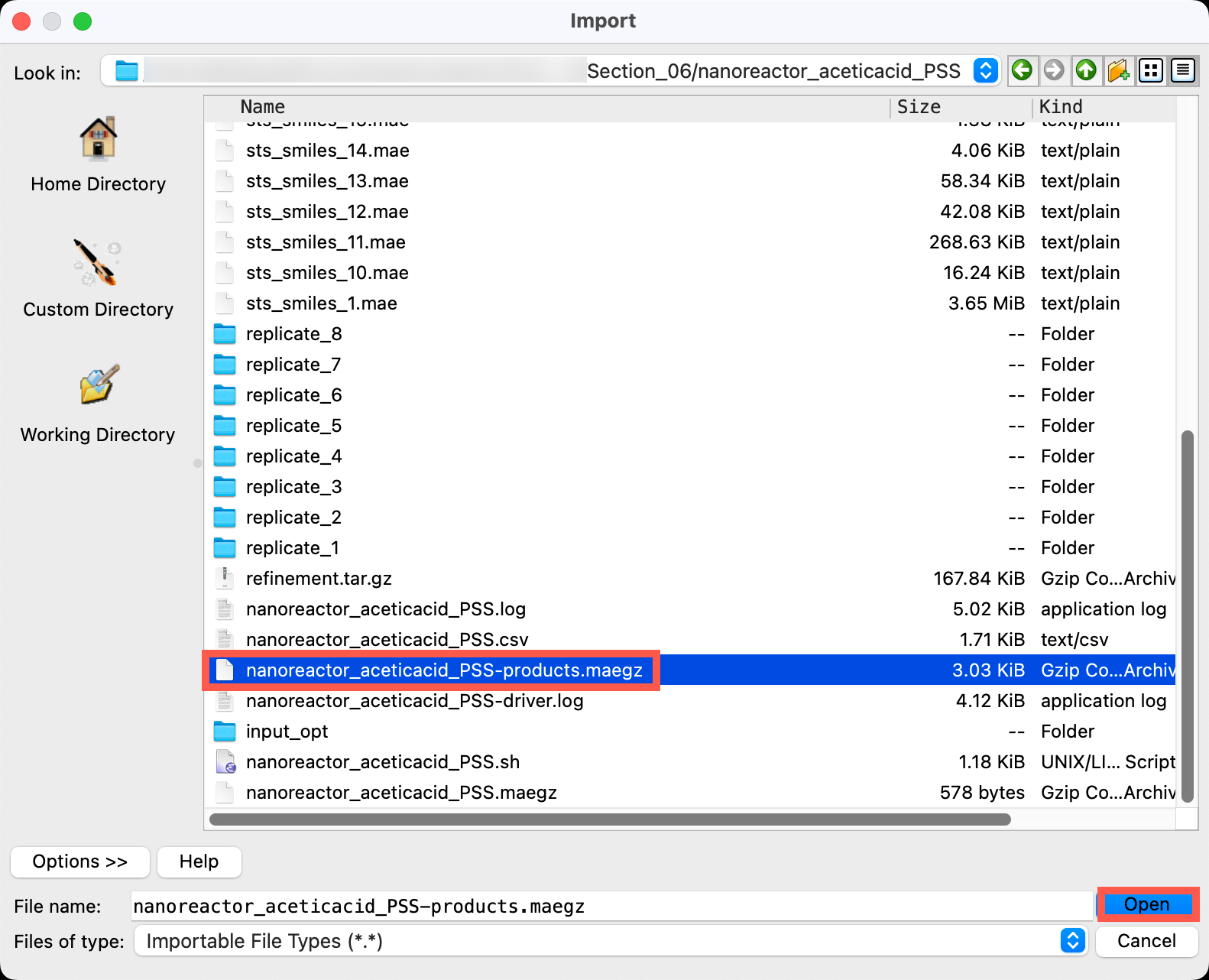
- Go to File > Import Structures
- Navigate to the provided files and choose
Section_06 > nanoreactor_aceticacid_PSS > nanoreactor_aceticacid_PSS-products.maegz - Click Open
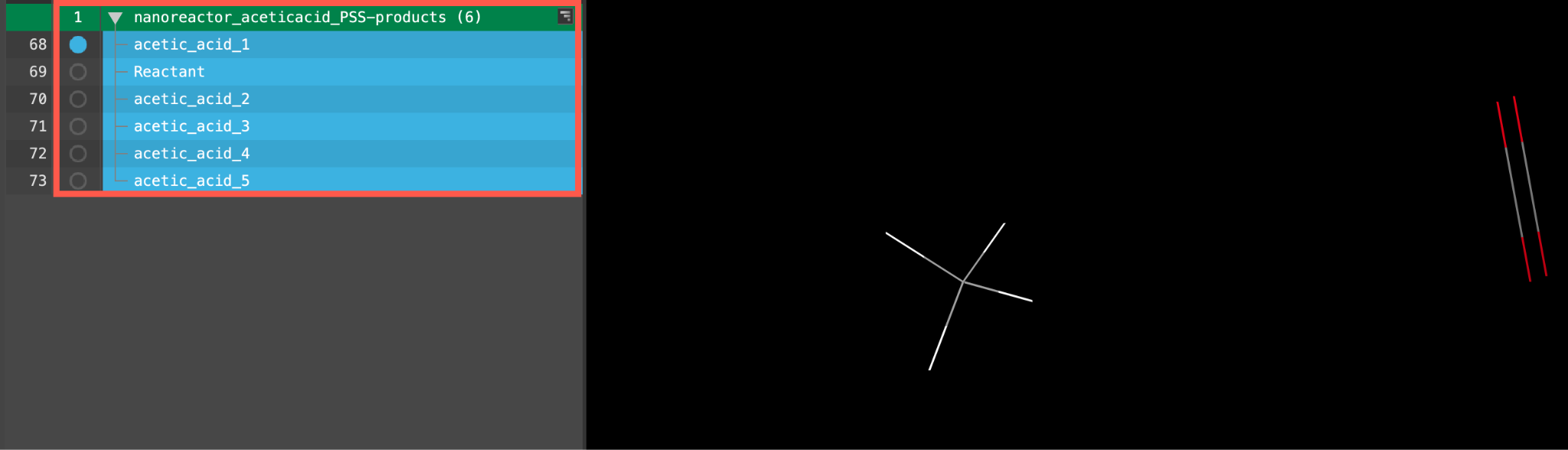
A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion. The group contains 6 entries.
Note that if you performed the calculation yourself, the exact number of output structures may vary due to randomness in the metadynamics simulations.
Feel free to visualize any of the entries in the workspace.
Quantitative analysis can be performed in the results panel:
- Use the WAM (workflow action menu) button (
 ) to access the Nanoreactor Results panel
) to access the Nanoreactor Results panel
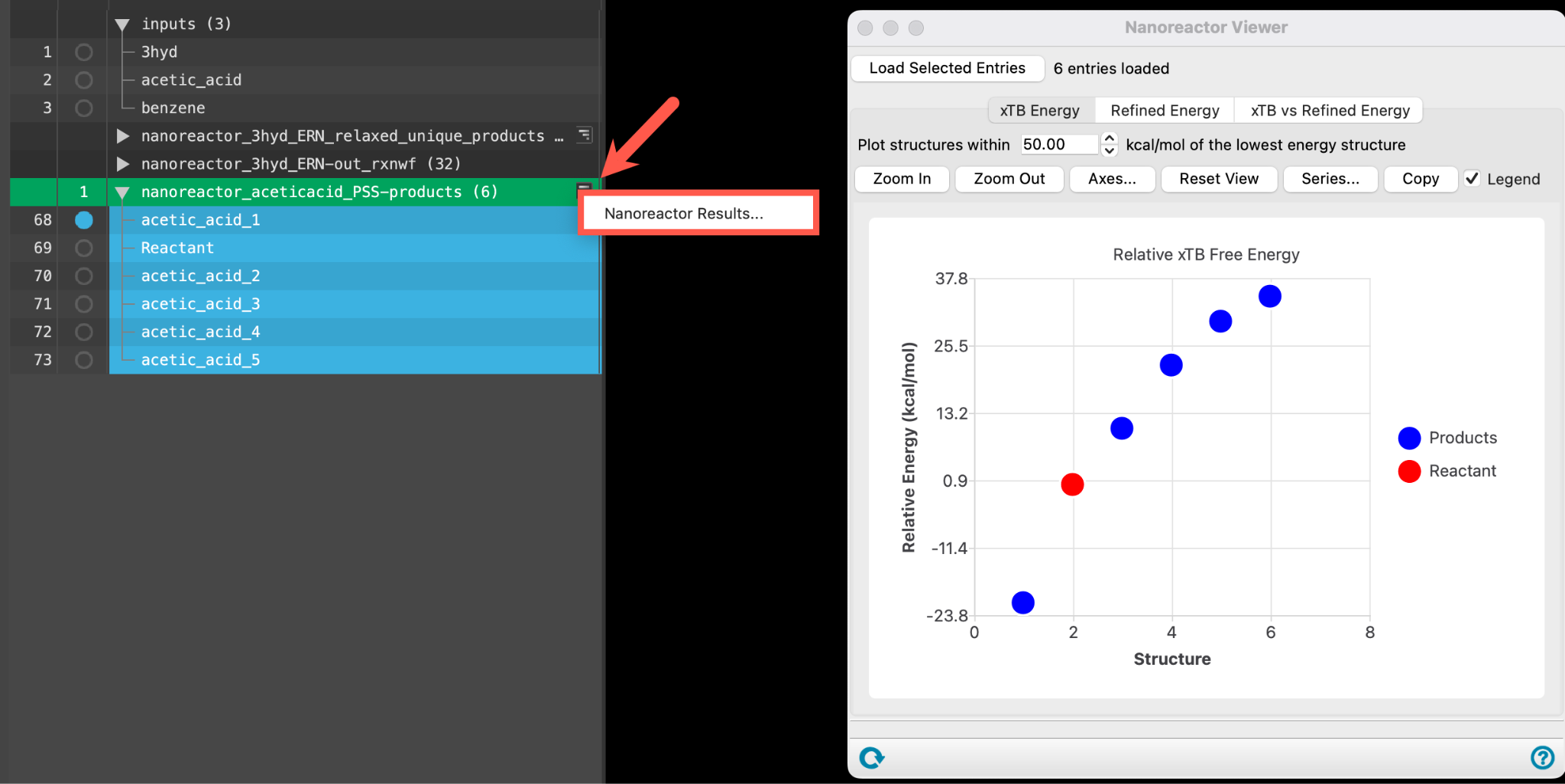
- Alternatively, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Nanoreactor Results and use the Load selected entries button to load the entire entry group into the panel
- The Nanoreactor Results panel opens
The first tab (xTB Energy) shows the relative GFN2-xTB energies for the product structures.
Note: The structures on the plot correspond to within 50 kcal/mol of the lowest energy structure.
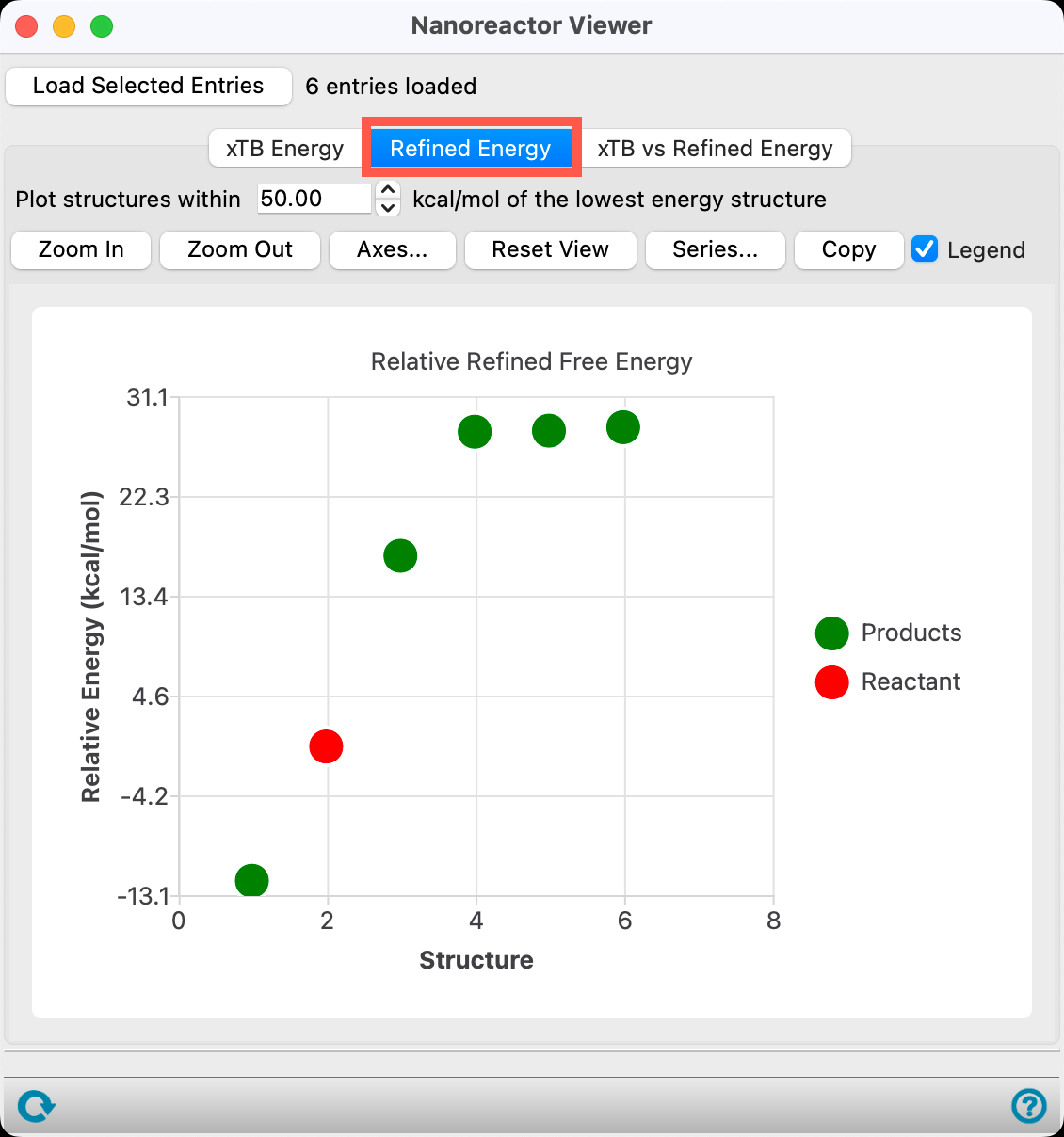
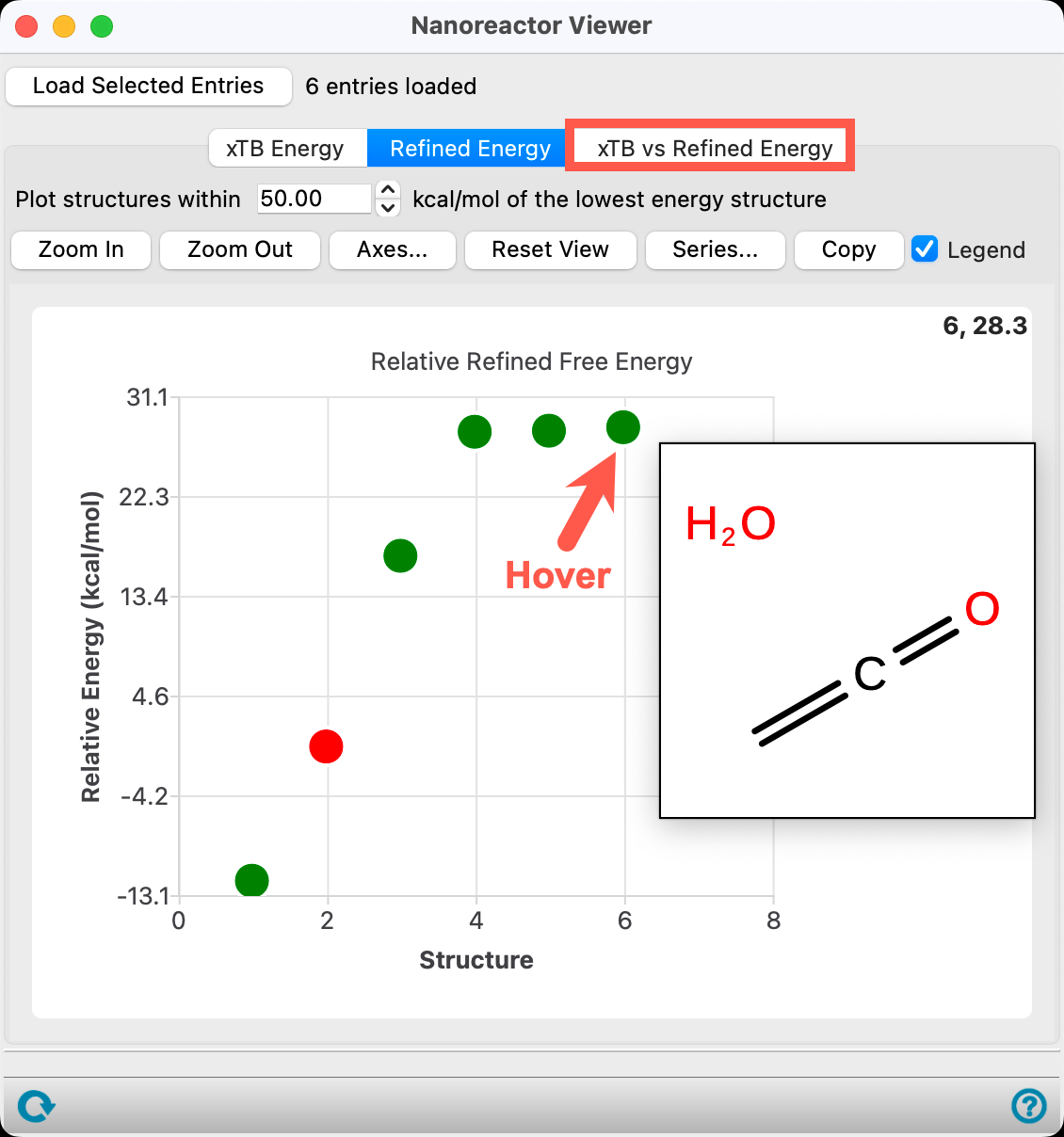
- Go to the Refined Energy tab
Because we checked Refine final energy in the initial calculation, relative energies are available.
The relative energy tab sorts the products based on DFT//GFN2-xTB thermodynamics. Specifically, electronic energy is calculated with DFT and thermal corrections are calculated with GFN2-xTB.
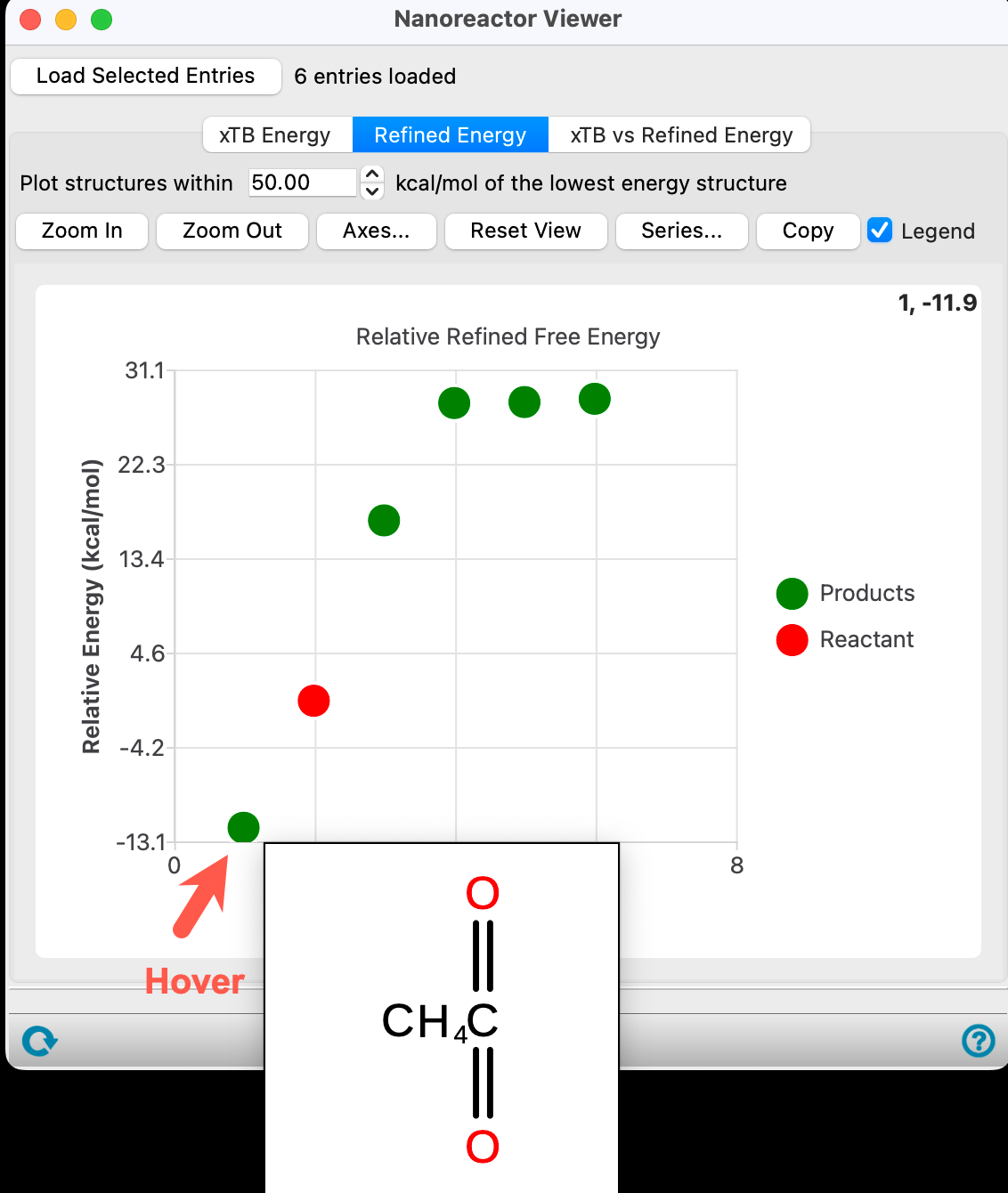
The energy is calculated with respect to the most thermodynamically stable species taken as 0, in this case CO2 and CH4.
- For confirmation, hover the mouse over the lowest energy blue data point to visualize a 2D sketch of the products
Note: In addition to hovering in order to see a structure, clicking on the data point will update the workspacethe 3D display area in the center of the main window, where molecular structures are displayed to include the structure.
The second product is trans-CH3CO2H (the input state). Proceed to visualize the various structures. Keep going and visualize all other products. In the provided output, the 5th structure represents ketene (CH2=C=O) and water.
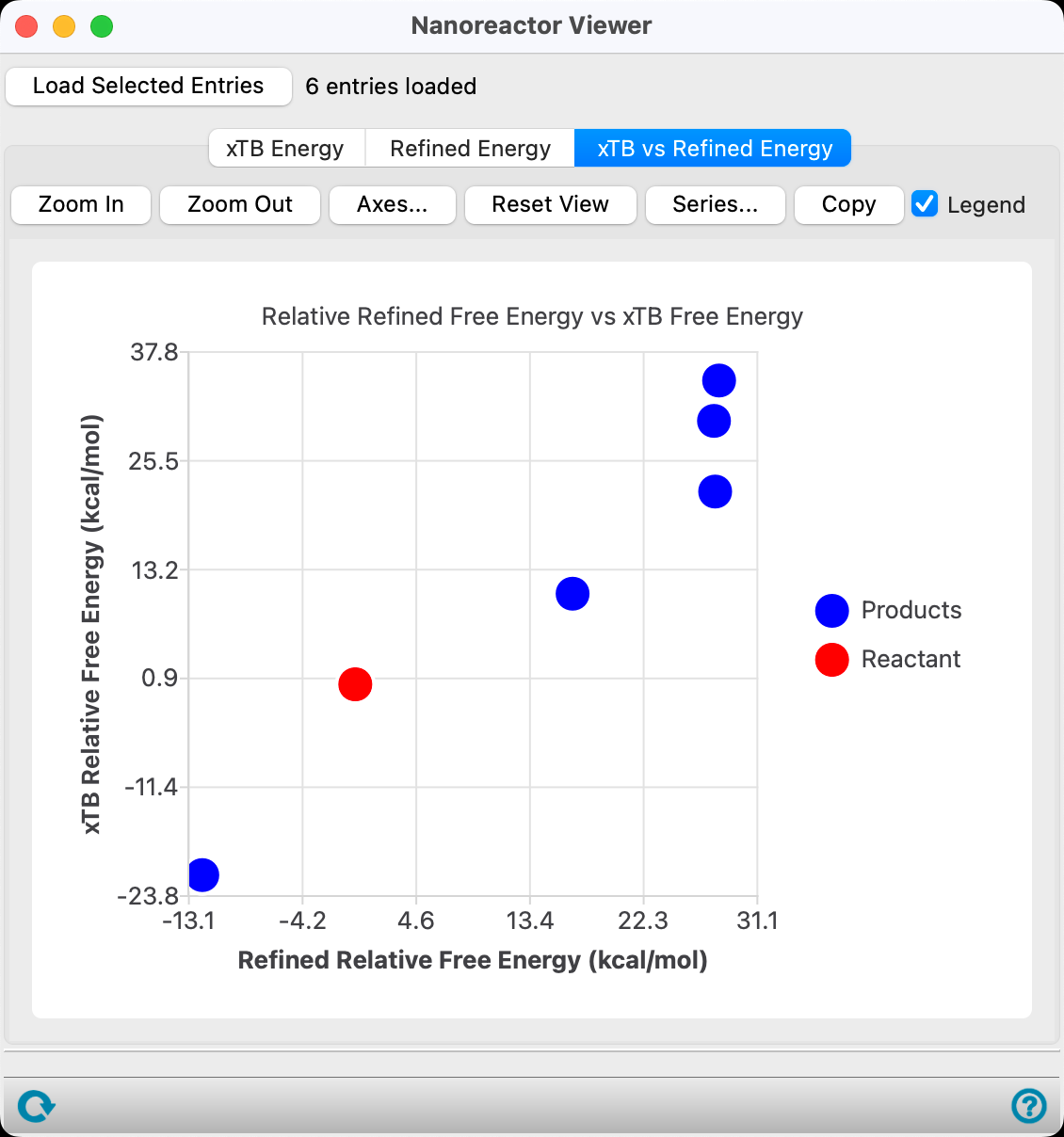
- Go to the xTB vs DFT//xTB Energy tab.
The low-energy CO2 / CH4 product as well as the CH2=C=O / H2O products are well-known experimentally detected products of acetic acid thermal decomposition in the gas-phase. See the References section (specifically, DOI:10.1002/kin.550160504).
There is also great qualitative agreement between DFT//GFN2-xTB and DFT thermodynamics. DFT thermodynamics were computed for acetic acid decomposition in the literature. See the References section (specifically, DOI:10.2174/1874123101812010014).
Moreover, Nanoreactor points out that in addition to these products, there could be several more products formed in this reaction (20+ based on the output entry group). Some of these products are more thermodynamically favorable than water and the ketene.
Finally, we can compare the GFN2-xTB based free energy with DFT//GFN2-xTB based free energy . Of course, the GFN2-xTB data is significantly less computationally expensive.
7. Performing and Analyzing a Nanoreactor Potential Energy Surface Sampling Calculation for Benzene
This section will demonstrate performing and analyzing a PSS calculation on benzene.
- With the benzene entry selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Nanoreactor Calculations
- The Nanoreactor panel opens
- Ensure that Potential energy surface sampling is selected for the Calculation Type
- Change the Metadynamics time to 100 ps
- Uncheck Remove high energy products based on xTB Free energy
- Check Refine final energy
- Open the Jaguar Options panel
- Select Machine learning force field to be UMA_sm_omol
- Click OK to close the panel
- Change the Job name to nanoreactor_benzene_pss
-
Adjust the job settings (
 ) as needed
) as needed
- This job requires a CPU host. The job can be completed in about 2 hours on 16 CPUs.
- If you would like to run the job yourself, click Run. Otherwise, we will proceed with imported results files in the next section
- Close the Nanoreactor panel
- Go to File > Import Structures
- Navigate to the provided files and choose
Section_07 > nanoreactor_benzene_pss > nanoreactor_benzene_pss-products.maegz - Click Open
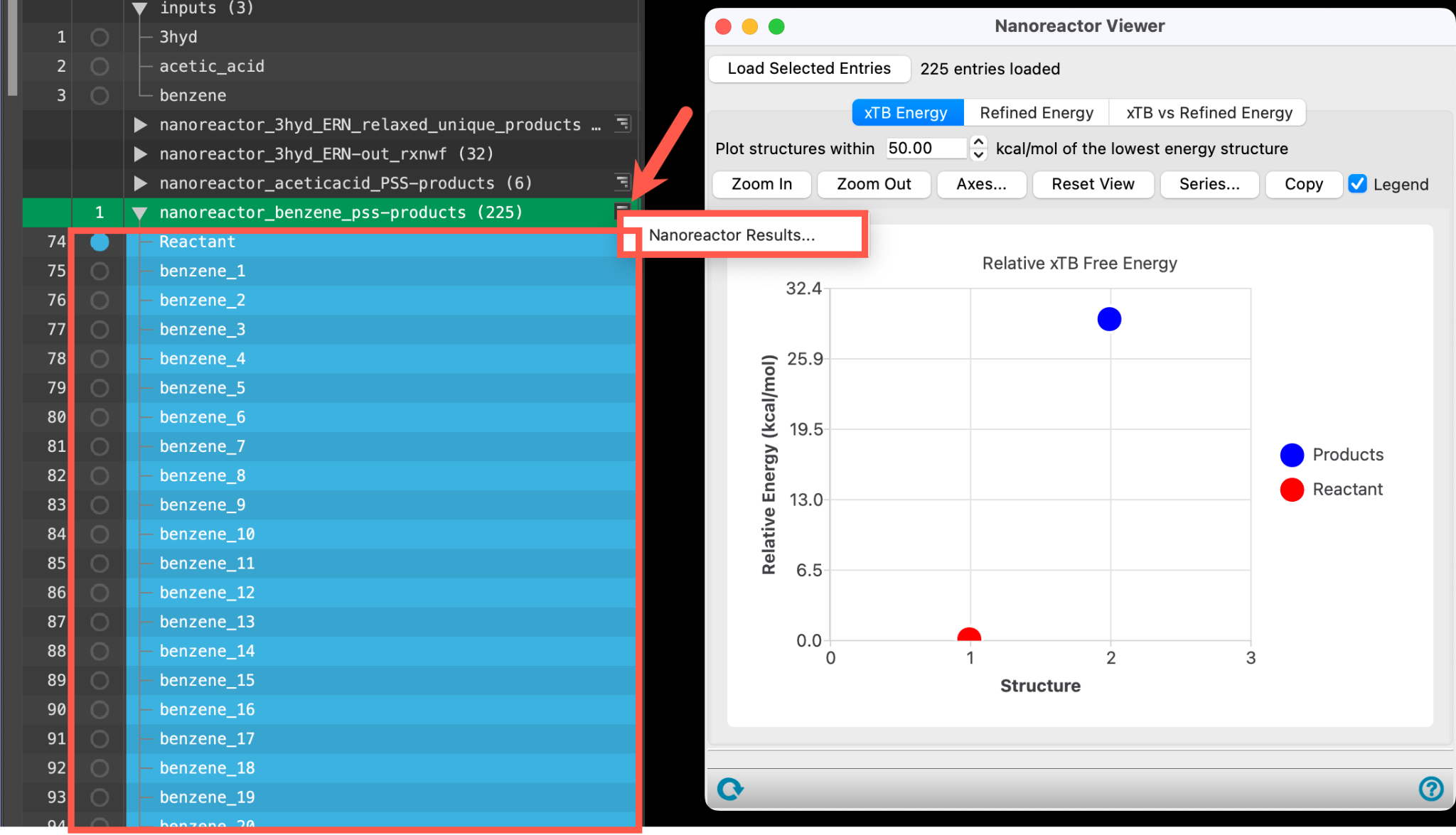
- A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion. The group contains 225 entries.
Note that if you performed the calculation yourself, the exact number of output structures may vary due to randomness in the metadynamics simulations.
Feel free to visualize any of the entries in the workspace.
- Use the WAM (workflow action menu) button (
 ) to access the Nanoreactor Results panel
) to access the Nanoreactor Results panel
- Alternatively, go to Tasks > Materials > Quantum Mechanics > Reaction Network > Nanoreactor Results and use the Load selected entries button to load the entire entry group into the panel
- The Nanoreactor Results panel opens
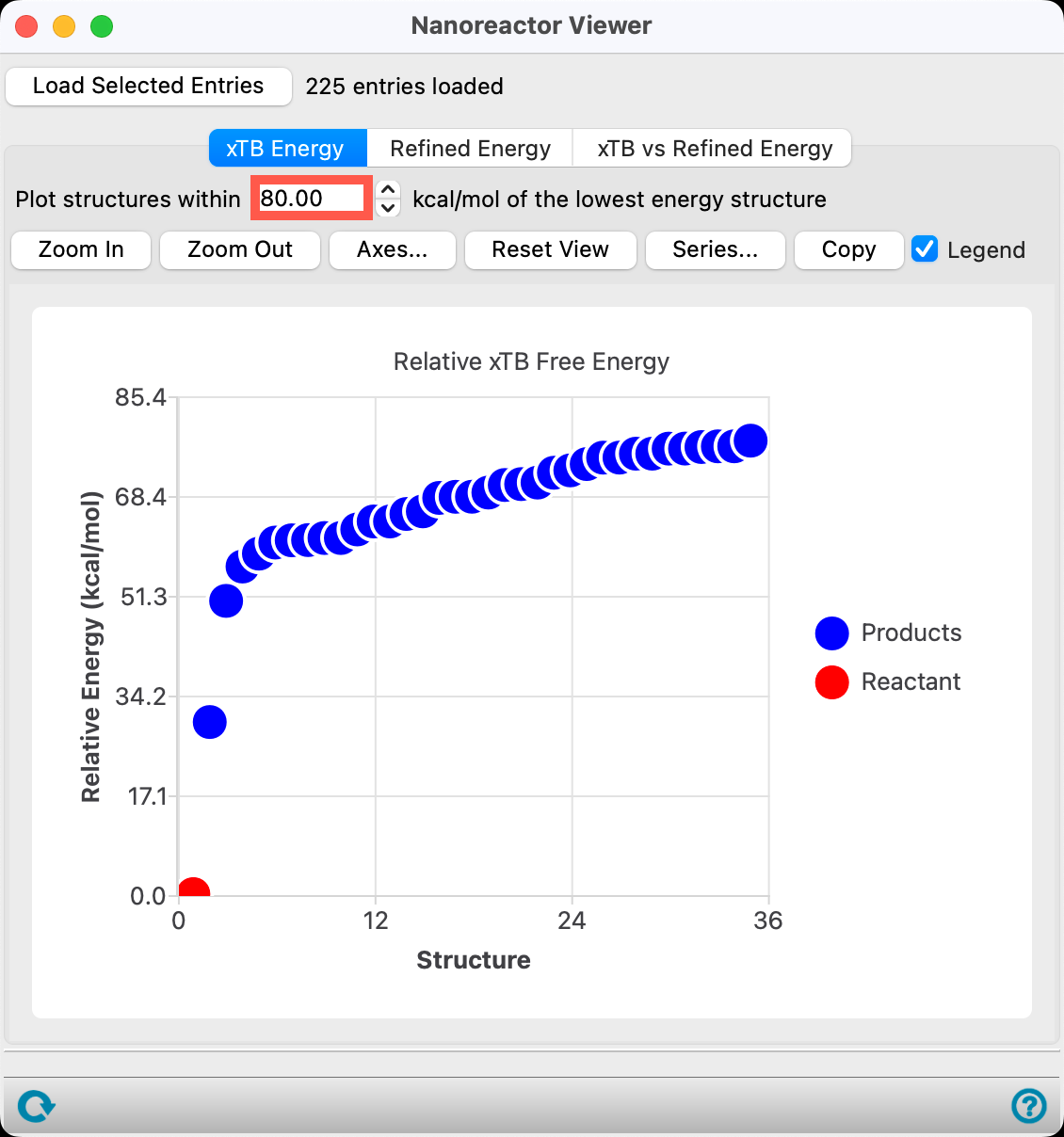
- Set the Plot structure within 80.00 kcal/mol of the lowest energy structure
- The number used for this value is flexible and can be adjusted by the user, as needed
- A threshold of 80 kcal/mol was chosen to illustrate a wide range of potential products
The majority of the products are benzene isomers; however, the results also show products from two distinct fragmentation pathways. See the References section (specifically, DOI: 10.1002/asia.202000535).
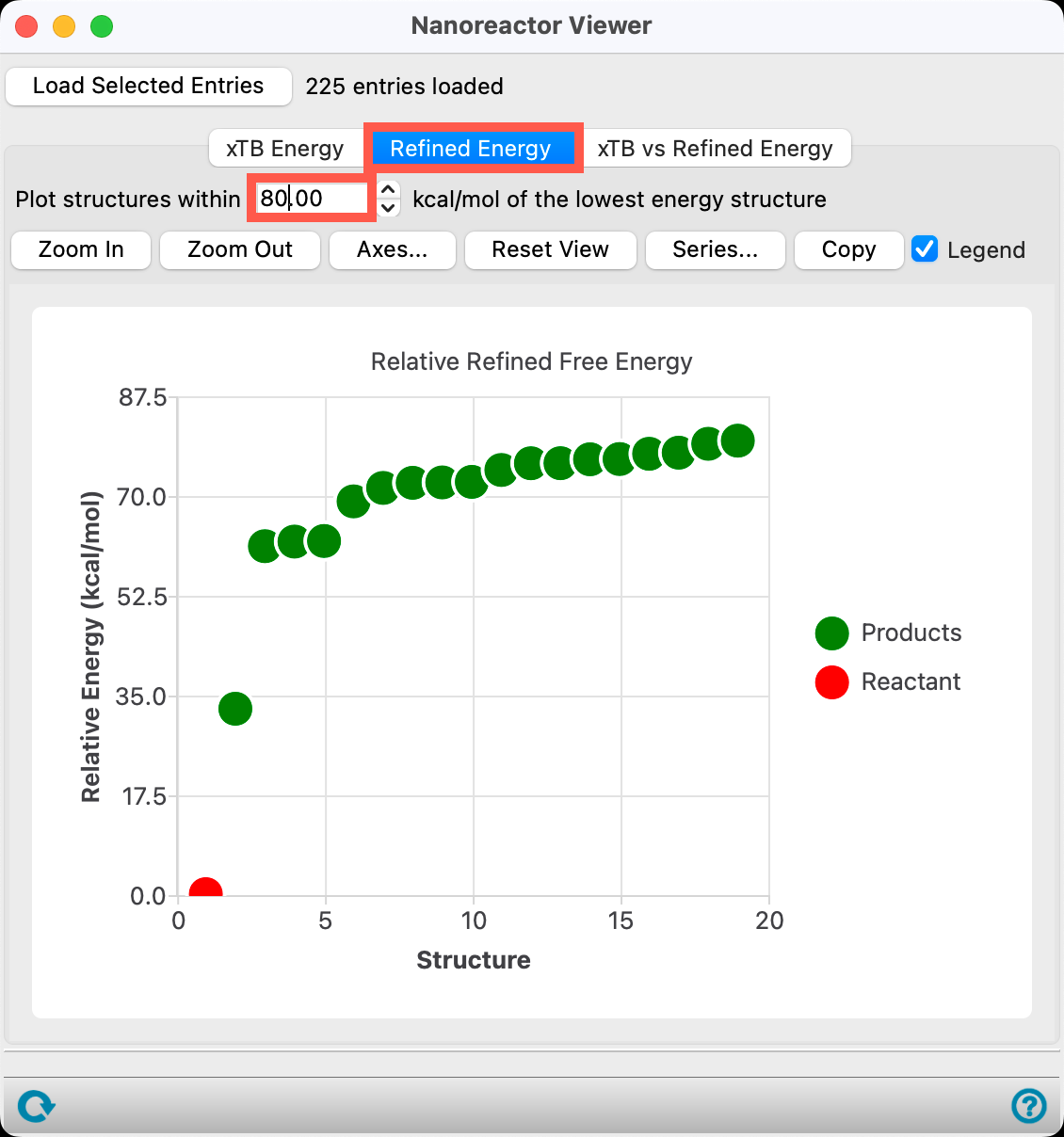
- Go to the Refined Energy tab
- Set the Plot structure within 80.00 kcal/mol of the lowest energy structure
- The number used for this value is flexible and can be adjusted by the user, as needed
- A threshold of 80 kcal/mol was chosen to illustrate a wide range of potential products
As a reminder, Nanoreactor only accounts for thermodynamics and not kinetics. It is especially useful when a user does not know the potential products of a reaction. To learn more about the kinetics of reactions of interest, refer to additional reactivity capabilities centering around transition states, such as those demonstrated in the Locating Transition States: Part 1 and Design of Asymmetric Catalysts with Reaction Network Enumeration Profiler tutorials.
8. Conclusion and References
In this tutorial, we learned how to use the Nanoreactor panel to predict thermal decomposition products of acetic acid and benzene without any prior knowledge of the reaction and study the ERN of 3-hydroperoxypropanal.
For further learning:
For introductory content, focused on navigating the Schrödinger Materials Science interface, an Introduction to Materials Science Maestro tutorial is available. Please visit the materials science training website for access to 100+ tutorials. For scientific inquiries or technical troubleshooting, submit a ticket to our Technical Support Scientists at help@schrodinger.com.
For self-paced, asynchronous, online courses in Materials Science modeling, including access to Schrödinger software, please visit the Schrödinger Online Learning portal on our website.
For some related practice, proceed to explore other relevant tutorials:
- Introduction to Geometry Optimizations, Functionals and Basis Sets
- Introduction to Multistage Quantum Mechanical Workflows
- Calculating Reaction Energetics for Molecular Systems
- Rigid and Relaxed Coordinate Scans
- Locating Transition States: Part 1
- Locating Transition States: Part 2
- R-group enumeration with the Materials Science Suite
- Design of Asymmetric Catalysts with Reaction Network Enumeration Profiler
- Microkinetic Modeling
- Organometallic Complexes
- Cheminformatics Machine Learning for Homogeneous Catalysis

For further reading:
- Help documentation on the Nanoreactor, Nanoreactor Results, and Reaction Network Viewer panels
- Fast and automated identification of reactions with low barriers: the decomposition of 3-hydroperoxypropanal. DOI: 10.21468/SciPostChem.1.1.003.
- Fast and automated identification of reactions with low barriers using meta-MD simulations. DOI: 10.7717/peerj-pchem.22
- Exploration of Chemical Compound, Conformer, and Reaction Space with Meta-Dynamics Simulations Based on Tight-Binding Quantum Chemical Calculations. DOI:10.1021/acs.jctc.9b00143
- High-temperature kinetics of thermal decomposition of acetic acid and its products. DOI:10.1002/kin.550160504
- Kinetics of Decomposition Reactions of Acetic Acid Using DFT Approach. DOI:10.2174/1874123101812010014
- GFN2-xTB—An Accurate and Broadly Parametrized Self-Consistent Tight-Binding Quantum Chemical Method with Multipole Electrostatics and Density-Dependent Dispersion Contributions. DOI:10.1021/acs.jctc.8b01176
- Discovering chemistry with an ab-initio nanoreactor. DOI:10.1038/nchem.2099
- Unimolecular Reaction Pathways of a γ-Ketohydroperoxide from Combined Application of Automated Reaction Discovery Methods. DOI: 10.1021/jacs.7b11009
- On Benchmarking of Automated Methods for Performing Exhaustive Reaction Path Search. DOI: 10.1021/acs.jctc.8b01182
- Automated Transition State Search and Its Application to Diverse Types of Organic Reactions. DOI:10.1021/acs.jctc.7b00764
- Inorganic Benzene Valence Isomers. DOI: 10.1002/asia.202000535
9. Glossary of Terms
Entry List - a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
Included - the entry is represented in the Workspace, the circle in the In column is blue
Project Table - displays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data
Recent actions - This is a list of your recent actions, which you can use to reopen a panel, displayed below the Browse row. (Right-click to delete.)
Scratch Project - a temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project
Selected - (1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Working Directory - the location where files are saved
Workspace - the 3D display area in the center of the main window, where molecular structures are displayed