Optoelectronics
Tutorial Created with Software Release: 2025-2
Topics: Energy Capture & Storage , Organic Electronics , Polymeric Materials
Methodology: Molecular Quantum Mechanics
Products Used: Jaguar , MS Maestro
|
1.0 MB |
This tutorial is written for use with a 3-button mouse with a scroll wheel.
Words found in the Glossary of Terms are shown like this: Workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Abstract:
In this tutorial, we will discuss the basic workflows in the optoelectronics panels of MS Maestro. First, we will calculate the oxidation potential, reduction potential, hole reorganization energy and triplet energy for two prototypical molecules: anthracene and phenylpyridine. Then, a larger data set will be provided and we will explore the utility of the optoelectronics results panel for visualizing data and identifying potential molecules of interest.
Tutorial Content
1. Introduction to Optoelectronics Workflows
The discovery and development of optoelectronic materials are of great interest, particularly with respect to organic light-emitting diodes (OLEDs) and organic photovoltaics (OPVs). When identifying optimal materials for optoelectronic applications, various properties are often considered: internal and condensed-phase properties, efficient charge injection and transport, and chemical and thermophysical stability, to name a few. The optoelectronics capabilities in the Materials Science Suite are designed to leverage rapid screening to complement experimental development by elucidating molecular properties and informing future synthetic targets.
To assess internal properties of optoelectronics materials, the Optoelectronics Calculations panel allows the user to implement efficient screening modes using the MS Jaguar quantum mechanics engine, minimizing computational expense while delivering valuable, accurate data.
For detailed descriptions of the various optoelectronics properties discussed herein, please refer to the Optoelectronic Properties help documentation.
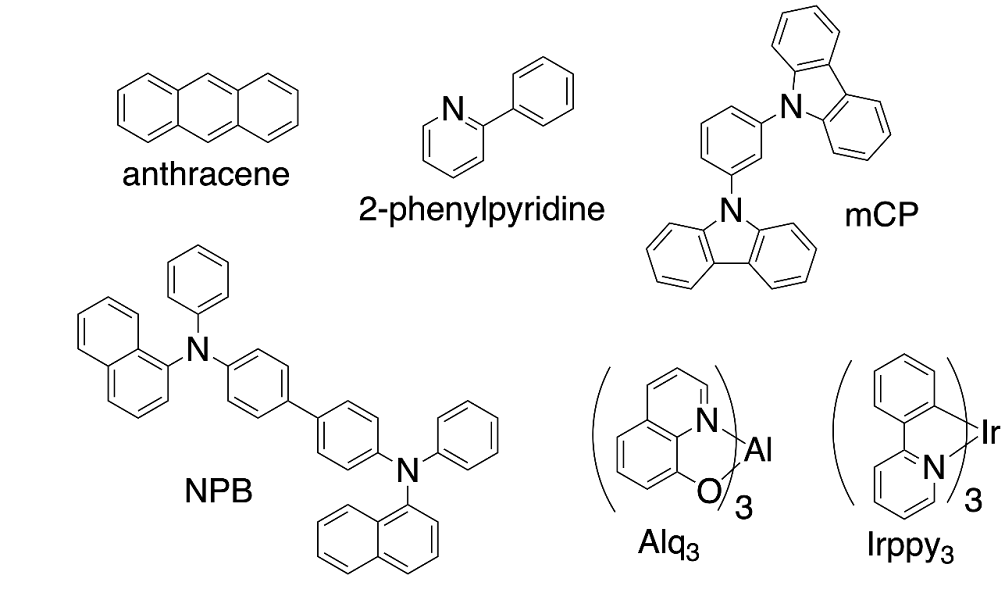
In this tutorial, we will first calculate the oxidation potential, reduction potential, hole reorganization energy, and triplet energy for two prototypical organic compounds (anthracene and 2-phenylpyridine, included in Scheme 1 below).
Scheme 1: Some prototypical molecules and ligands of relevance to optoelectronics
Subsequently, a larger data set will be provided to practice the process of using the optoelectronics results tools to efficiently sift through large data sets to identify promising candidates from a screen.
Finally, we note that while these tools are designed with optoelectronics in mind, the workflows are not limited to optoelectronic screening. General chemical analysis of any molecule (or family of molecules) can be conveniently performed using the tools described here. Relatedly, other complementary tools exist in the Materials Science Suite that might be useful when analyzing optoelectronics candidate molecules such as the Bond and Ligand Dissociation and Reaction Energetics Enumeration panels.
2. Creating Projects and Importing Structures
At the start of the session, change the file path to your chosen Working Directorythe location where files are saved in MS Maestro to make file navigation easier. Each session in MS Maestro begins with a default Scratch Projecta temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project, which is not saved. A MS Maestro project stores all your data and has a .prj extension. A project may contain numerous entries corresponding to imported structures, as well as the output of modeling-related tasks. Once a project is saved, the project is automatically saved each time a change is made.
Structures can be built in MS Maestro or can be imported using File > Import Structures (or drag-and-dropped), and are added to the Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data. The Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion is located to the left of the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed. The Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data can be accessed by Ctrl+T (Cmd+T) or Window > Project Table if you would like to see an expanded view of your project data.
- Double-click the Materials Science icon
- (No icon? See Starting Maestro)
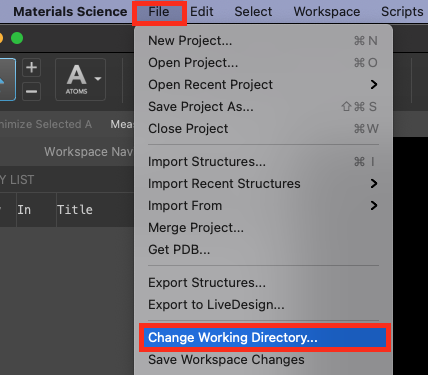
- Go to File > Change Working Directory
- Find your directory, and click Choose
- Pre-generated input and results files are included for running jobs or examining output. Download the zip file here: schrodinger.com/sites/default/files/s3/release/current/Tutorials/zip/optoelectronics.zip
- After downloading the zip file, unzip the contents in your Working Directorythe location where files are saved for ease of access throughout the tutorial
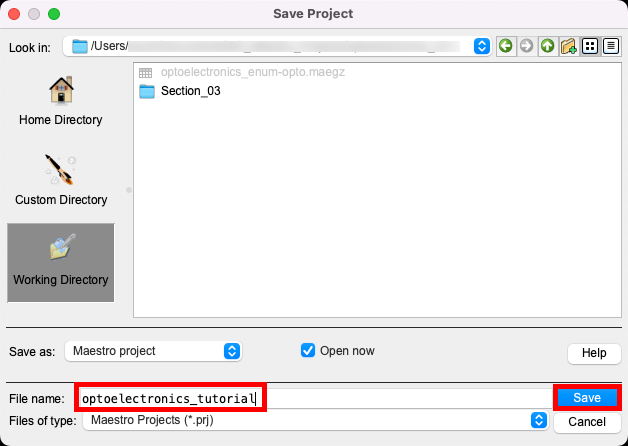
- Go to File > Save Project As
- Change the File name to optoelectronics_tutorial, click Save
- The project is now named
optoelectronics_tutorial.prj
- The project is now named
3. Optoelectronics Screening for Anthracene and Phenylpyridine
We will proceed to draw two organic molecules which will then be used for a standard optoelectronics calculation.
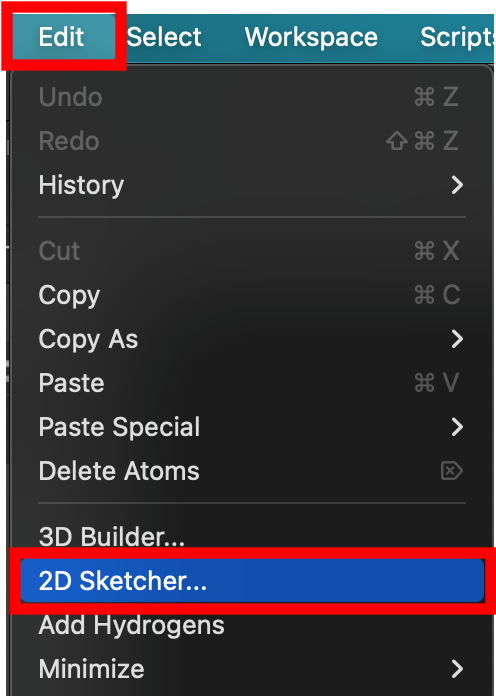
- From the main menu, go to Edit > 2D Sketcher
- The 2D sketcher opens
Note: To skip manual drawing, go to Import ( ) > Paste in Text and use the following strings for anthracene and 2-phenylpyridine: C1=CC=C2C=C3C=CC=CC3=CC2=C1 and C1=CC=C(C=C1)C2=CC=CC=N2. Name the entries anthracene and ppy and proceed to Step 5.
) > Paste in Text and use the following strings for anthracene and 2-phenylpyridine: C1=CC=C2C=C3C=CC=CC3=CC2=C1 and C1=CC=C(C=C1)C2=CC=CC=N2. Name the entries anthracene and ppy and proceed to Step 5.
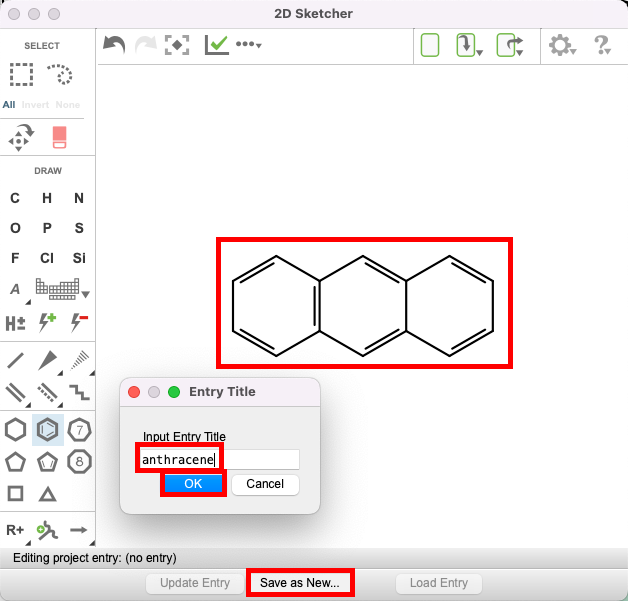
- Draw anthracene
- Click Save as New and name the entry: anthracene, then click OK.
- The 3D anthracene molecule will be both includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed and selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
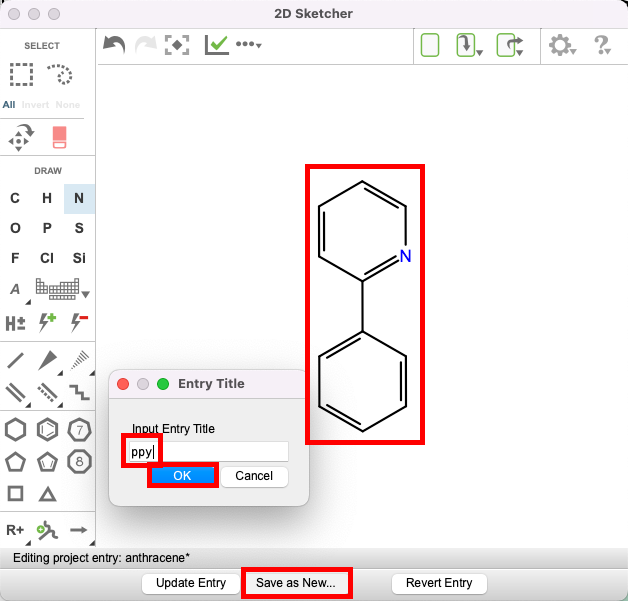
- Keep the 2D sketcher open, erase the anthracene molecule and sketch 2-phenylpyridine (shown in the Figure)
- To do so, use the eraser, the benzene ring, the line tool, and the N atom
- Click Save as New and name the entry: ppy, then click OK
Note: Clicking Update Entry will overwrite the anthracene molecule. While this can be a useful way for editing a molecule in the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed, in this instance, we want to draw another molecule, so Save as New should be used
- Close the 2D Sketcher.
Now the 3D ppy molecule will be includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed and selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
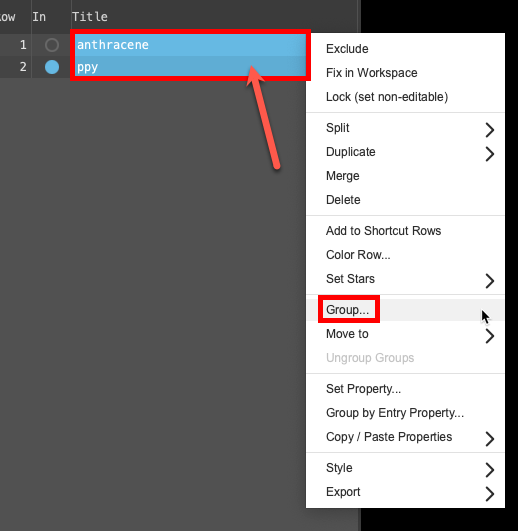
- Select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries both molecules in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion (Shift + click) and right click on either name in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- A menu opens
- From the menu, click Group
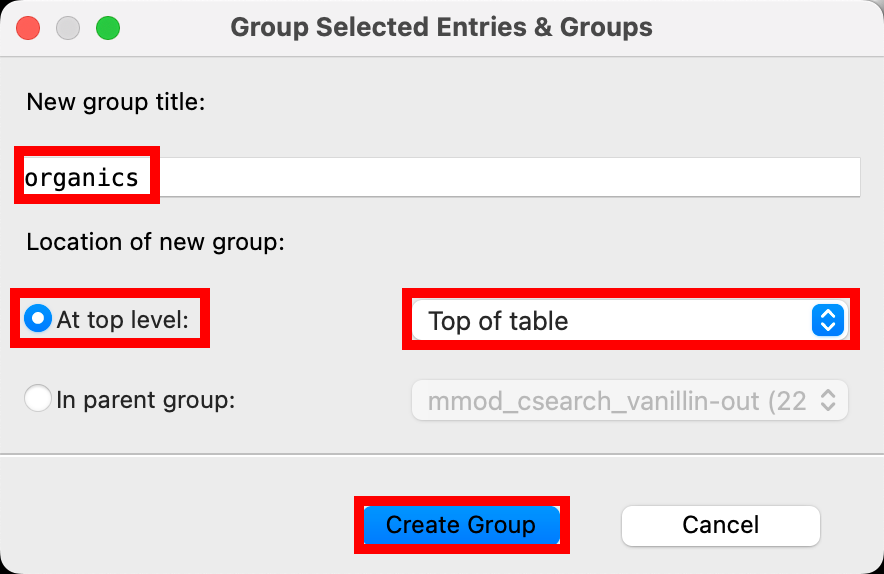
- The Group Selected Entries & Groups panel opens
- Change New group title to organics
- Select At top level for the Location of new group
- In the corresponding option menu select Top of table
- Click Create Group
- The two molecules in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion are grouped under a common group called organics (2)
Note: While this grouping step is by no means required or necessary, it is a convenient way to keep your entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion organized and to make multiple selections later, especially if you are developing a large project
Often at this step, it would be recommended to perform a Conformational Search. This is particularly important because several optoelectronics properties are quite sensitive to conformation. However, in this case, the starting geometries are suitable to proceed given the limited conformational space associated with these simple molecules.
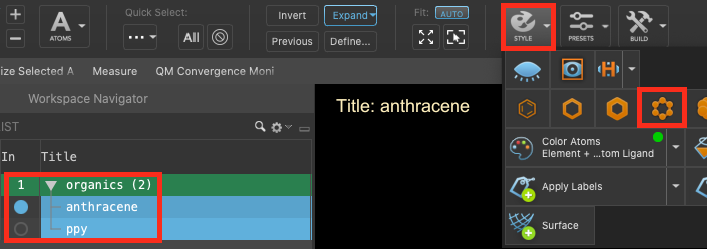
- Ensure that the organics (2) group is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Quantum Mechanics > Optoelectronics > Optoelectronics Calculations
- The Optoelectronics Calculations panel opens
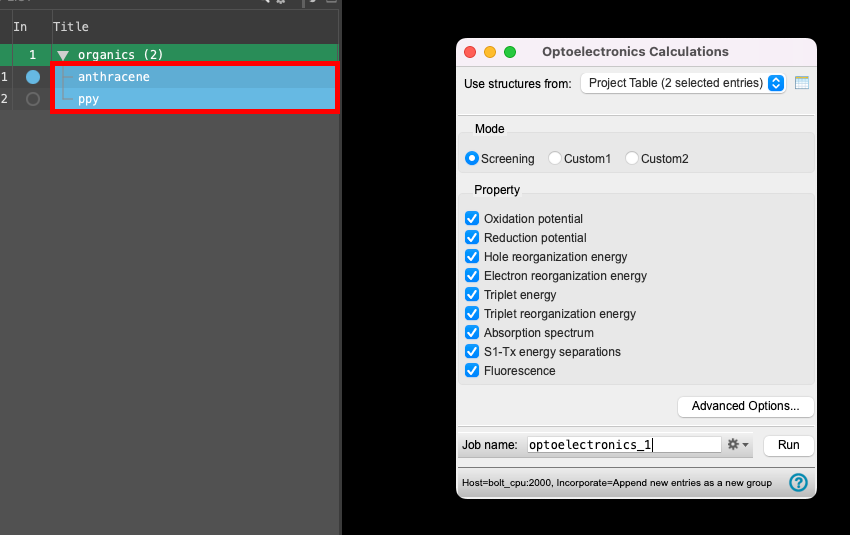
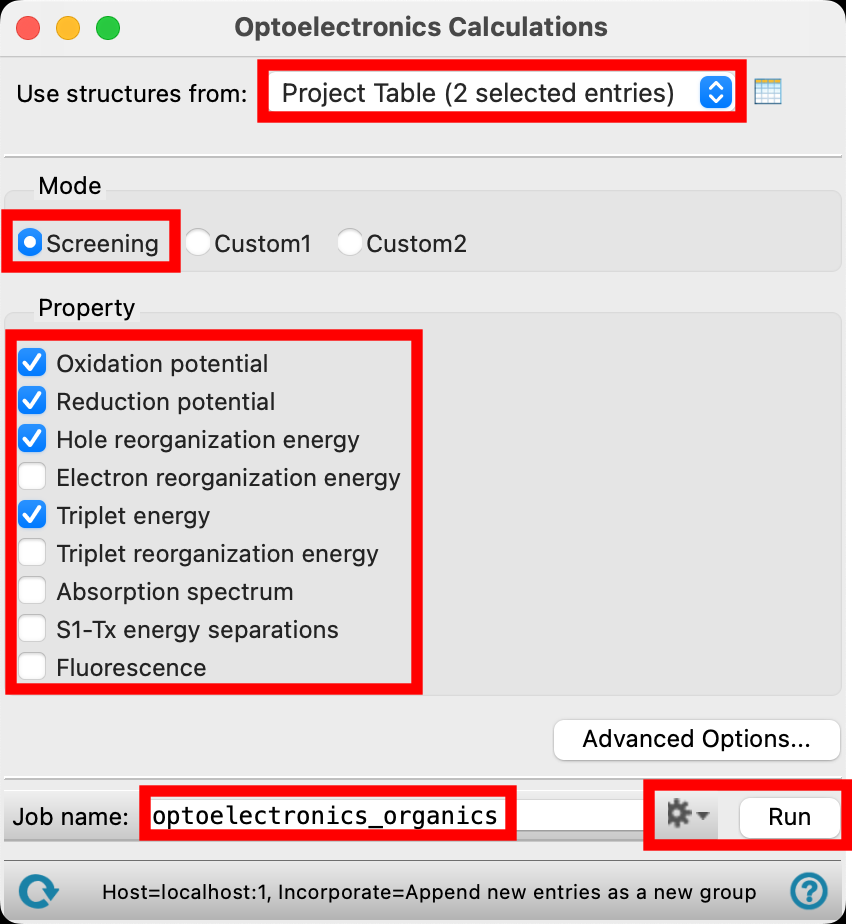
- For Use structures from, choose Project Table (2 selected entries)
- For Mode, select Screening
- Choose the following from the Property section: oxidation potential, reduction potential, hole reorganization energy, and triplet energy
- Change the Job name to optoelectronics_organics
- Adjust the job settings depending on your available resources, and click Run
- This job takes ~6 minutes on a 12 CPU host
- Pre-generated results are also available for proceeding
- Close the Optoelectronics Calculations panel
Screening mode is highly recommended for efficiently and accurately running optoelectronics calculations. To use a Custom mode or see the parameterization associated with Screening mode, click Advanced Options
For detailed descriptions of the various optoelectronics properties, please refer to the Optoelectronic Properties help documentation.
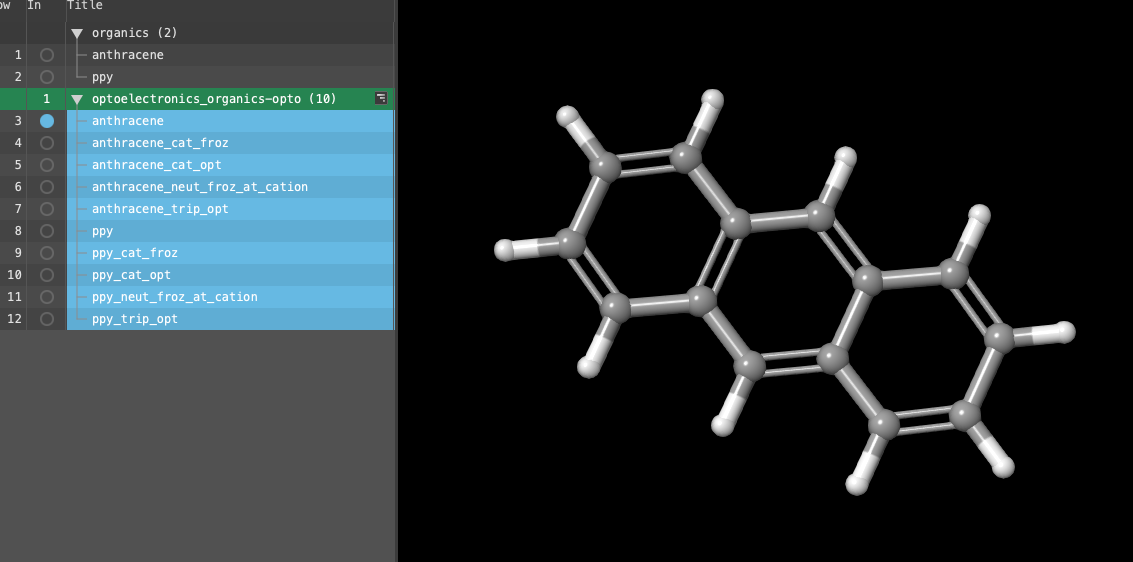
- If you ran the job, select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries the entry group from the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion: optoelectronics_organics-opto1 (10) and proceed to Step 22
- Otherwise, to import the pre-generated results, from the main menu, choose File > Import Structures
- Navigate to where you downloaded the tutorial files and choose the
optoelectronics_organics-opto.maegzfile - Click Open
- A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion. The entry is automatically selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Note: Ten outputs should be in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion (five for each molecule). The requested optoelectronics properties are calculated from a minimum number of calculations with relatively inexpensive but accurate basis sets due to the efficiency of Screening mode
In Section 4 of the tutorial, we will practice using the more advanced optoelectronics analysis tools. For now, let’s simply look at some of the data for anthracene and 2-phenylpyridine in the Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data.
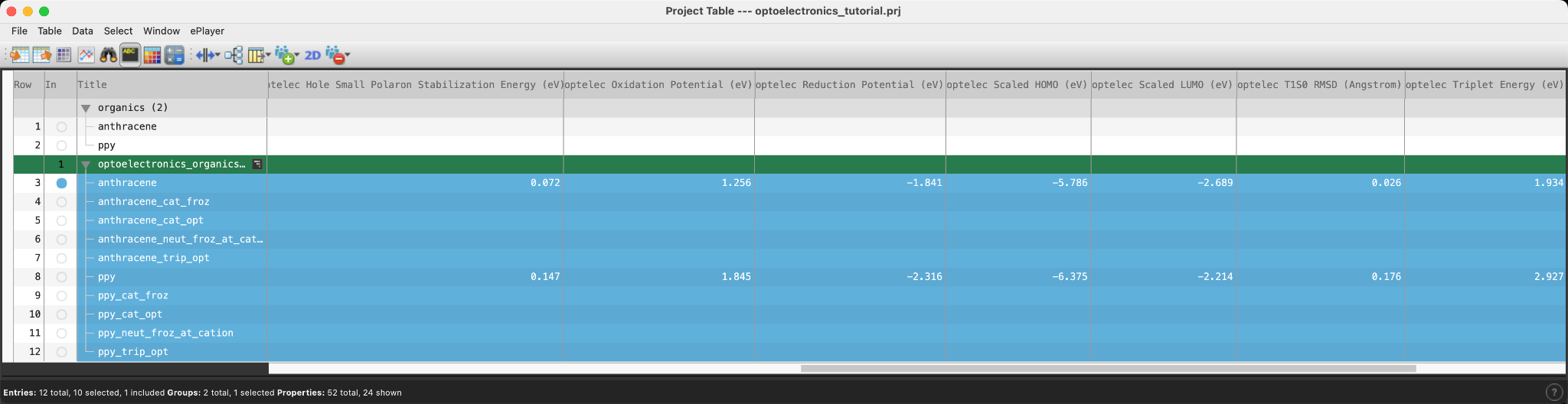
- Open the Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data

- Data from the job is available for perusal. Notice that in addition to the requested values, some additional output is provided, including dipole moment (D)
4. Importing and Analyzing a Larger Data Set
We will now learn a bit more about the analysis tools associated with optoelectronics calculations. For this section, a larger data set of 165 molecules has been provided. In this case, the molecules were designed using enumeration techniques starting with 2-phenylpyridine (see: R-Group Enumeration with the Materials Science Suite). Otherwise, the same steps that we just implemented for calculating optoelectronics properties for anthracene and 2-phenylpyridine were repeated.
- From the main menu, choose File > Import Structures
- Navigate to where you downloaded the tutorial files and choose the provided
Section_04 > optoelectronics_enum-opto.maegzfile - Click Open
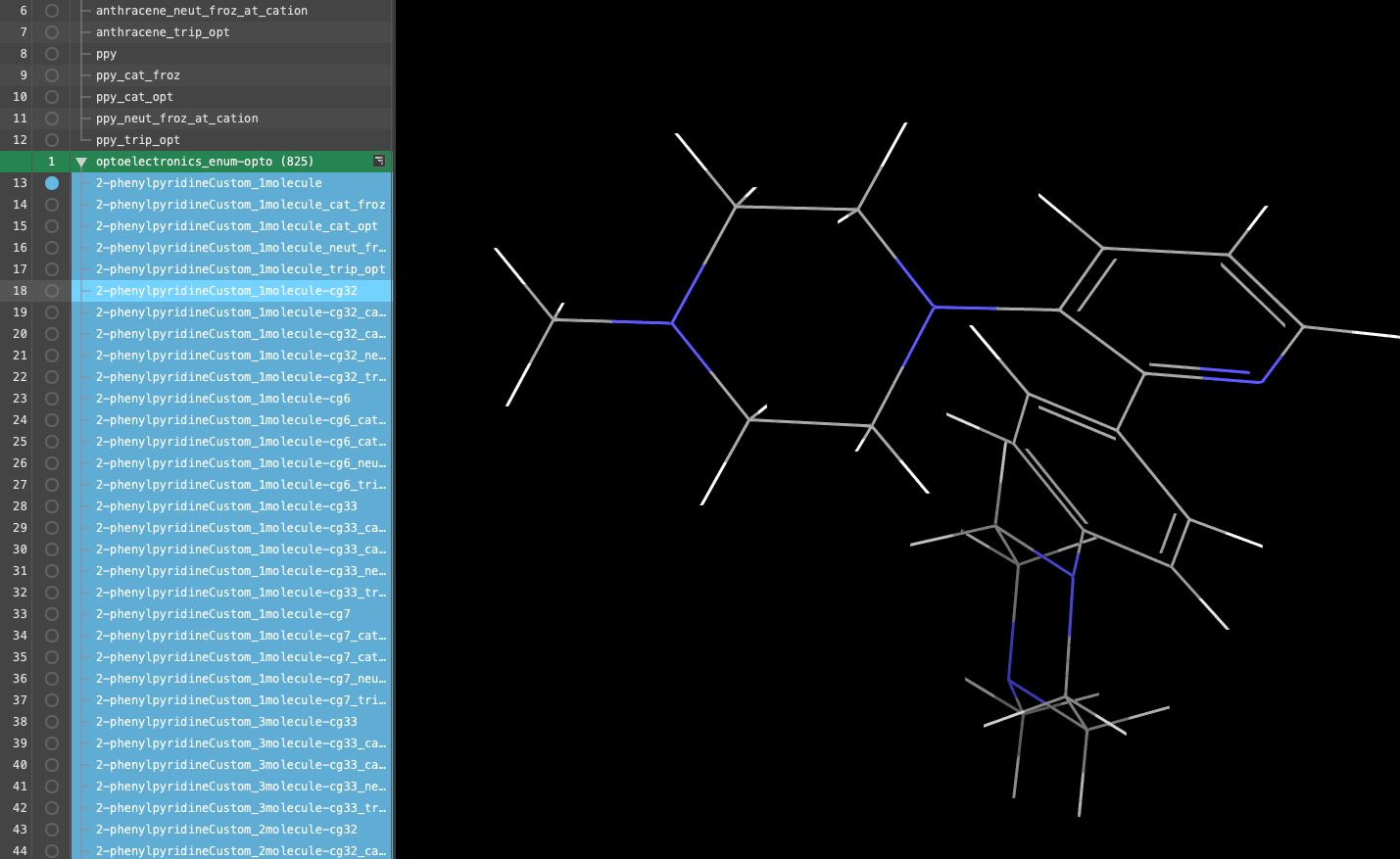
- A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion. The group is automatically selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
- The group is titled optoelectronics_enum-opto (825)
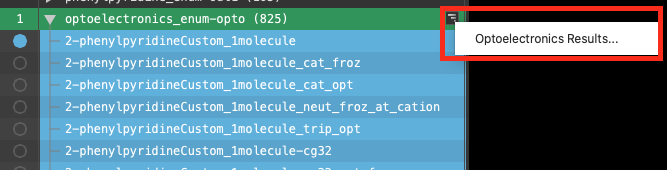
The most convenient method to access the Optoelectronics Results panel is by using the Workflow Action Menu (WAM) button  which appears next to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion.
which appears next to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion.
- Click on the WAM button and select Optoelectronics Results
- The Optoelectronics Results panel opens
Optoelectronics data for the 165 enumerated compounds is presented in the table. In addition to the requested properties, several additional outputs are provided that are generated from the calculation at no additional computational expense
Note: The results panel can also be accessed from the tasks menu via Tasks > Materials > Quantum Mechanics > Optoelectronics > Optoelectronics Results
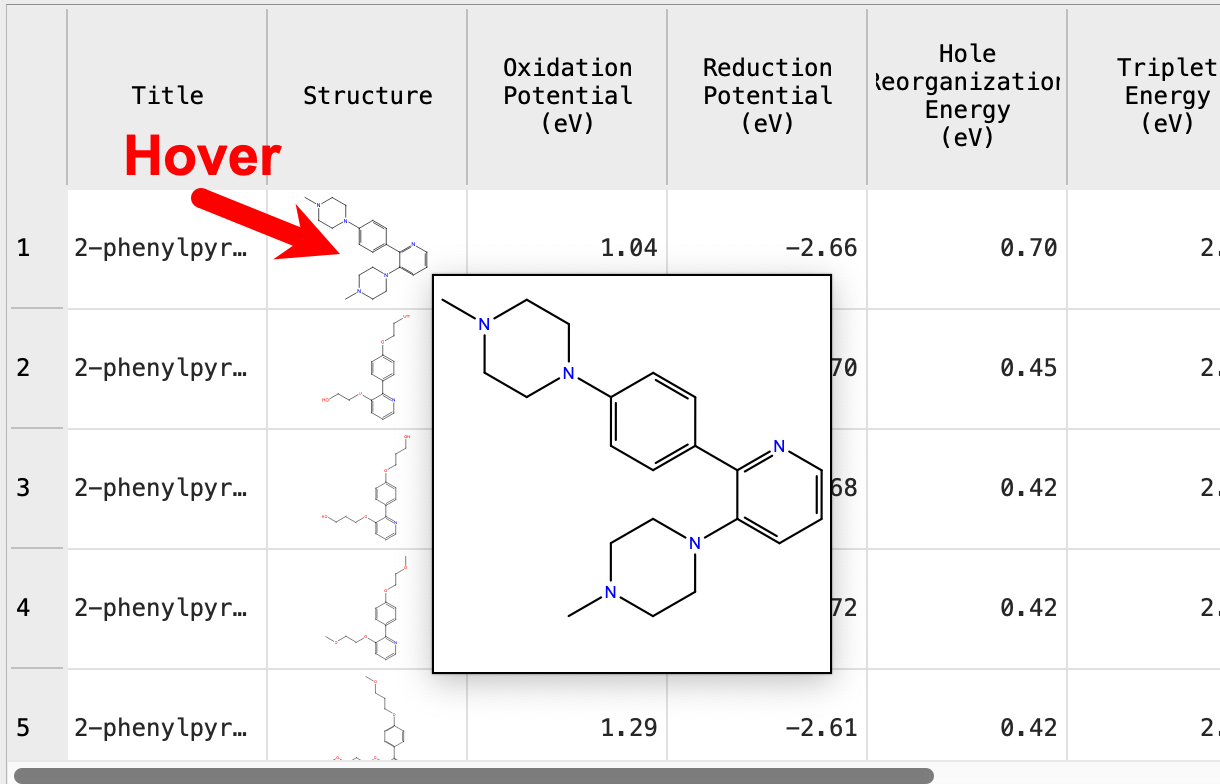
Hovering the mouse over any of the structures, you can see a larger 2D image
-
Hover the mouse over any of the structures
- A larger 2D inset appears
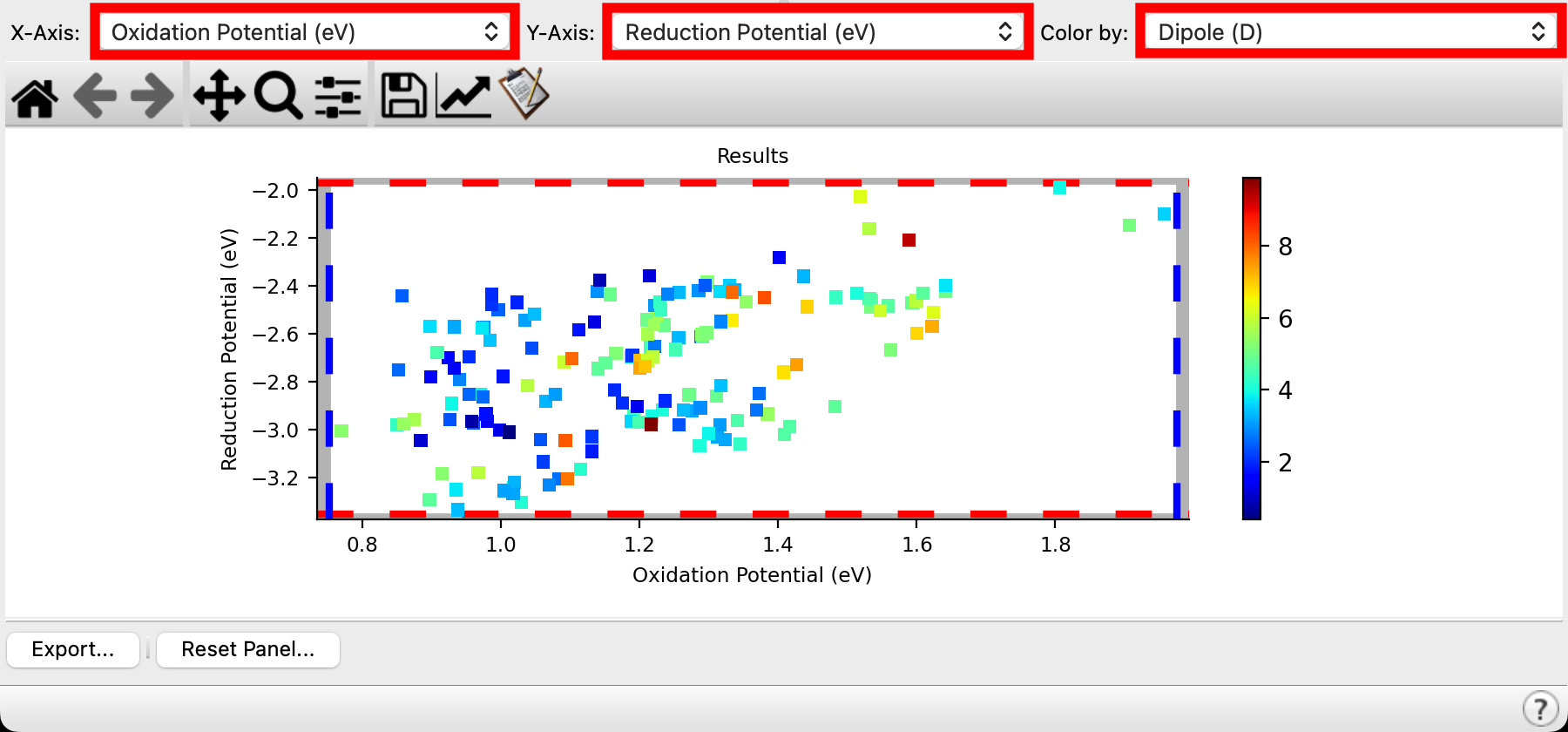
Figure 4-4. Optoelectronics results plot of oxidation potential, reduction potential and dipole moment.
A practical approach to sifting through a large data set is to use the plotting tool to visually identify target molecules. For this exercise, let’s suppose we are interested in identifying a 2-phenylpyridine derivative with the following attributes: oxidation potential between 1.35 and 1.50 eV, reduction potential above -2.6 eV, and a low dipole moment
- From the X-axis dropdown menu, select Oxidation Potential (eV)
- From the Y-axis dropdown menu, select Reduction Potential (eV)
- From the Color by dropdown menu, select Dipole (D)
- A results plot is immediately generated with the requested data
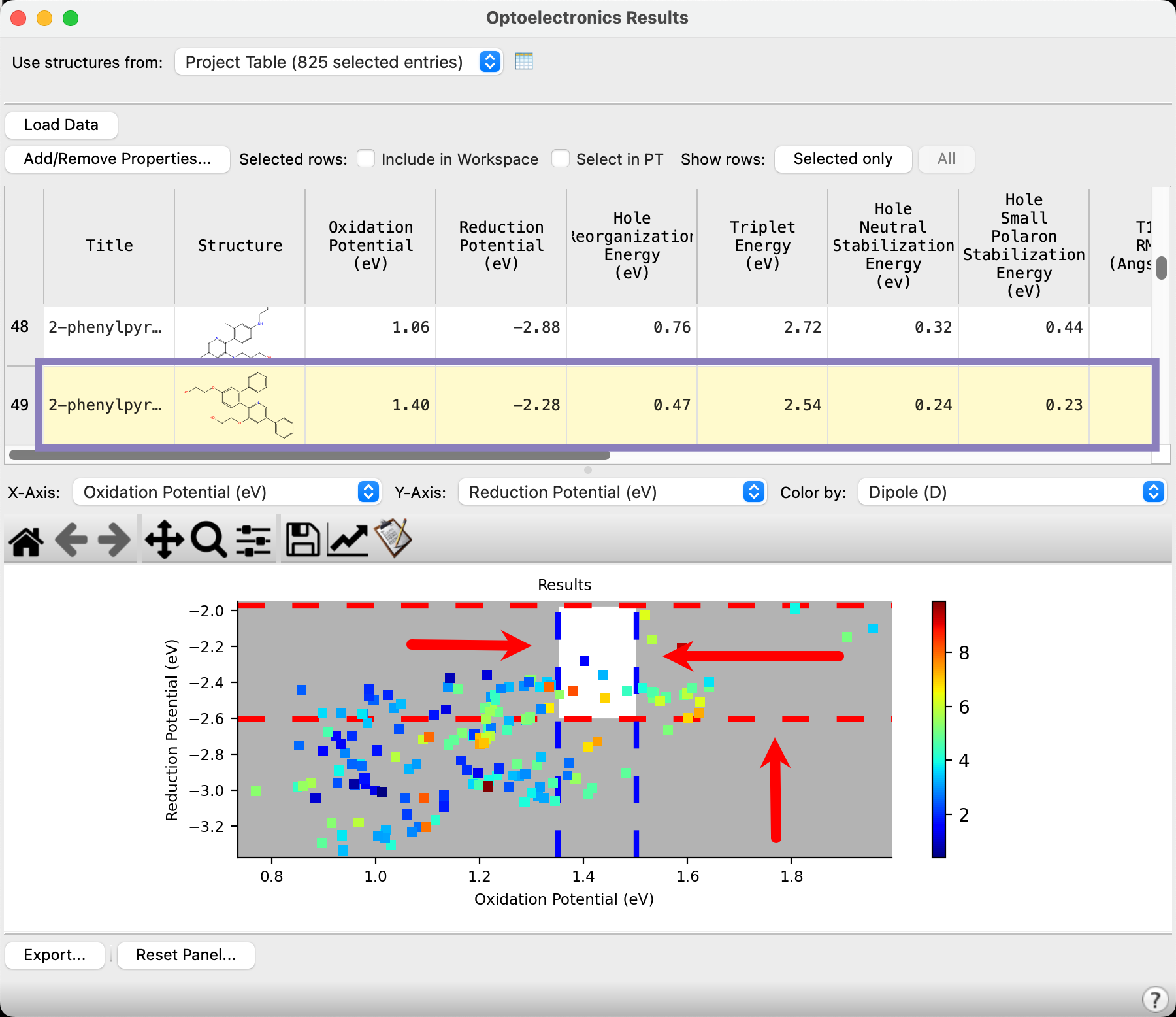
The dashed bars can be used to filter a set of target compounds.
- Click and drag on the blue vertical dashed lines to approximately enclose the target oxidation potential region of 1.35-1.50 eV
- The range outside of the dashed lines turns grey
- Click and drag on the bottom red horizontal dashed line to approximately enclose the target reduction potential above -2.6 eV
- The range outside of the dashed lines turns grey
- Scroll in the above table and you will notice that the compounds matching the gating criteria are highlighted in yellow
Six compounds appear to fit the criteria based on oxidation and reduction potentials. Analyzing the dipole moment coloring indicates that four of the compounds have relatively lower dipole moments
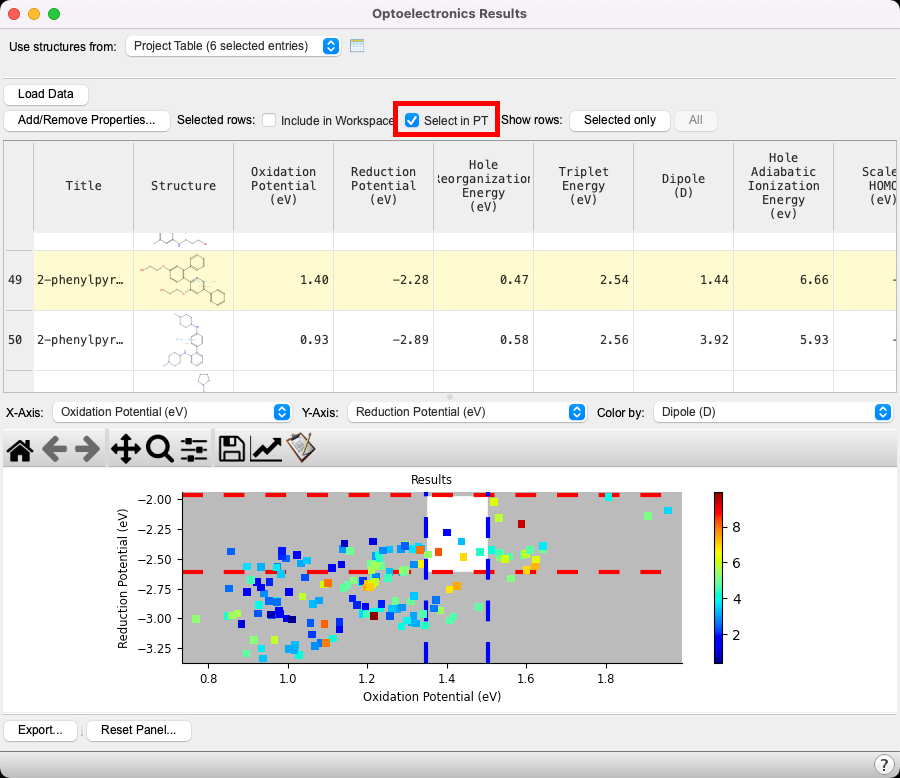
- Click on the Select in PT (Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data) checkbox
- The six compounds that fit the criteria are selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries in the entry list. These entries are highlighted in blue in the entry list
- Close the Optoelectronic Results panel
Note: The Include in Workspace and Select in PT checkboxes are convenient tools for communicating between the workspacethe 3D display area in the center of the main window, where molecular structures are displayed, the project tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data and the optoelectronics results
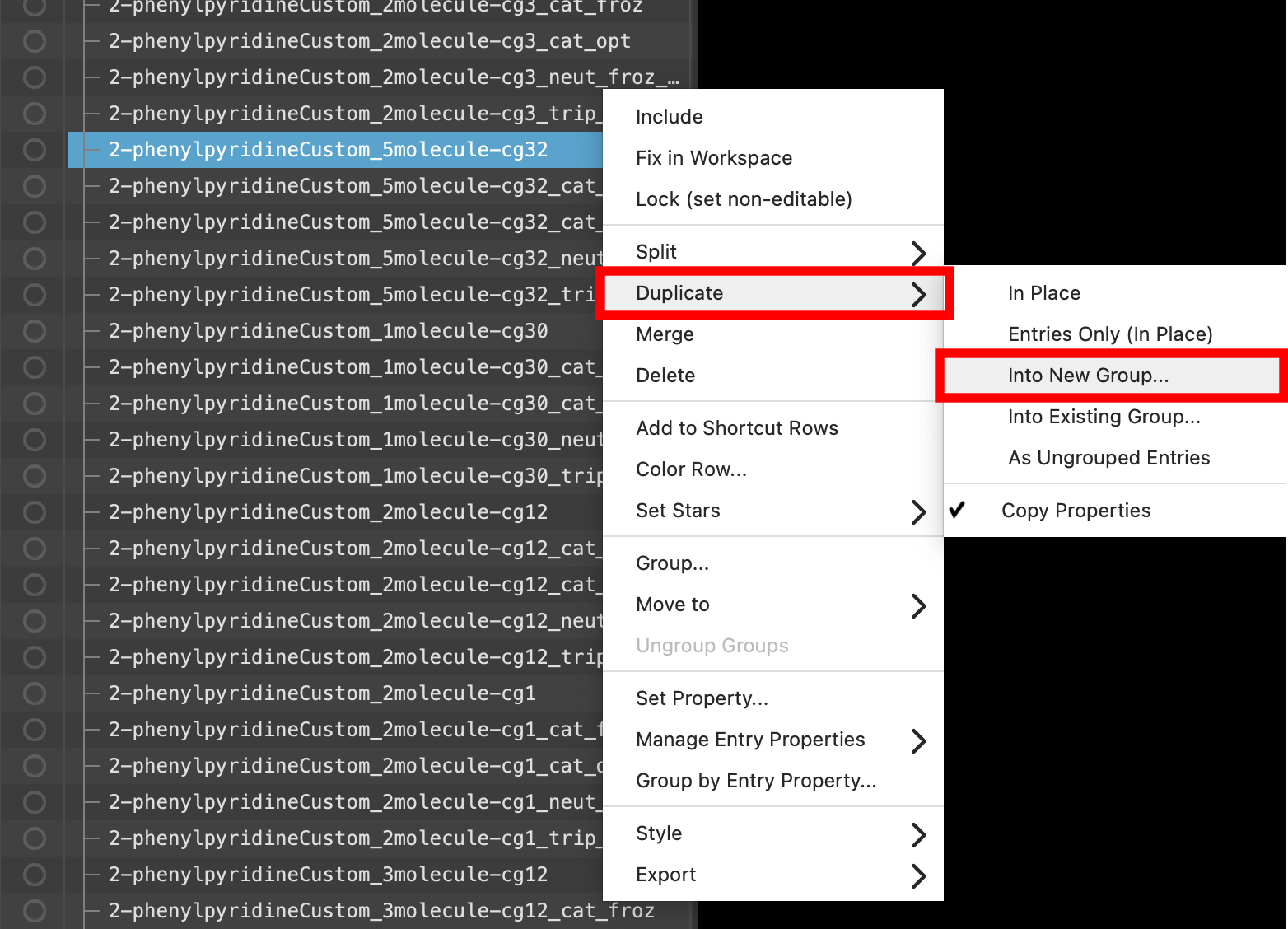
- Back in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion, right click on any of the six selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries compounds (highlighted in blue)
- Click Duplicate > Into New Group
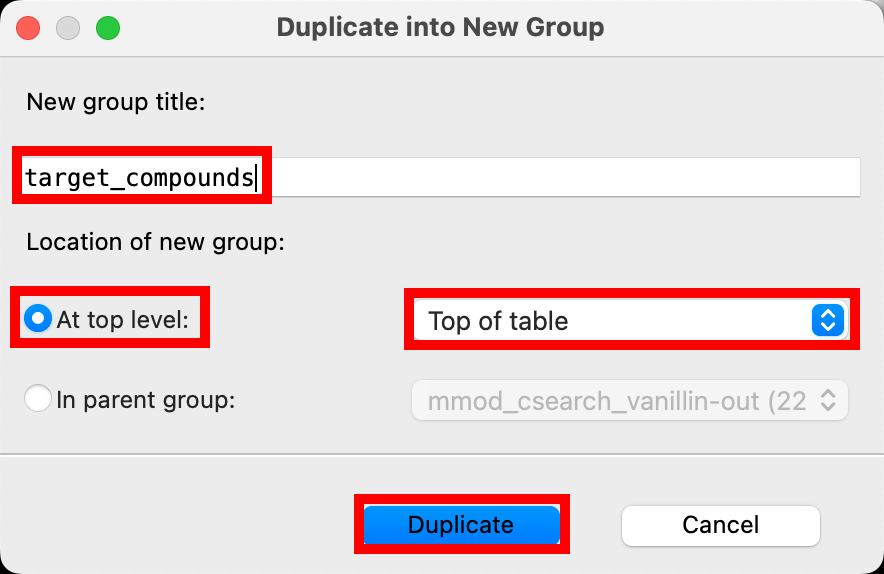
- The Duplicate into New Group panel opens
- Set the group title to target_compounds
- Click the At top level radio button
- In the corresponding option menu select Top of table
- Click Duplicate
- A new group is created in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion titled target_compounds containing the six molecules of interest
- The group is both selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Includethe entry is represented in the Workspace, the circle in the In column is blue all 6 molecules in the target_compounds group in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
To see the exact values for the parameters of interest, you can reopen the Optoelectronics Results panel or the Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data at any time and cross-reference the molecule name.
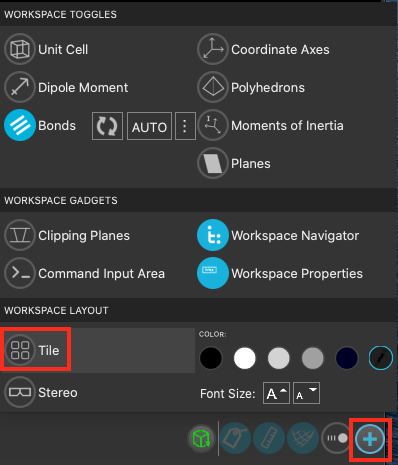
Alternatively, you may wish to include some properties in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed directly.
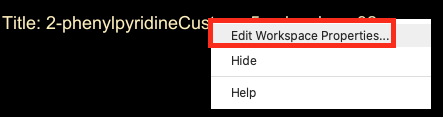
- Right click on the Title in the upper-left corner of the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
- Click Edit Workspace Properties
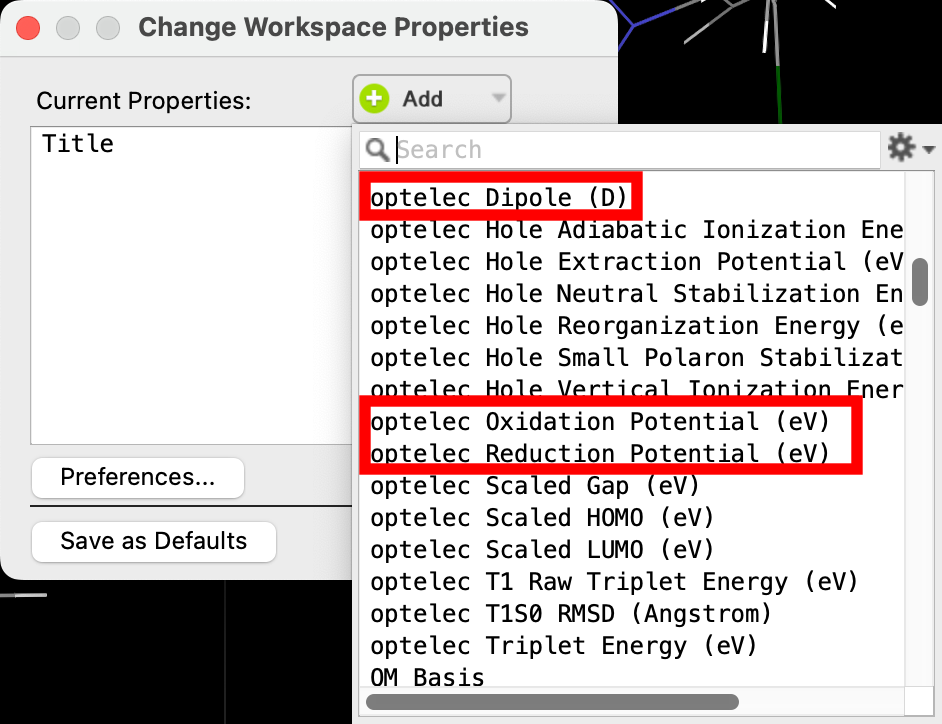
- Change Workspace Properties appears
- Click Add and select optelec oxidation potential, optelec reduction potential and optelec Dipole (D)
- Click Close
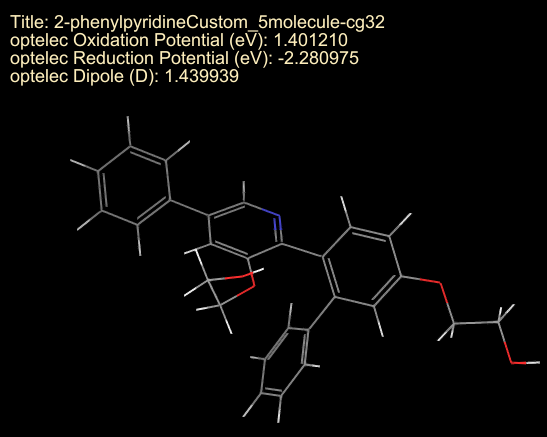
- Each tile now contains the title as well as the three properties of interest
Based on the hypothetical criteria, the substituted 2-phenylpyridine with the lowest dipole moment (1.44) within the ideal oxidation and reduction potential ranges is identified and might be considered for further exploration
5. Conclusion and References
In this tutorial, we utilized the optoelectronics panels in MS Maestro to calculate several key properties for two prototypical molecules: anthracene and phenylpyridine. Then, a larger data set was analyzed to practice visualizing data and identifying potential molecules of interest from a screening output.
For further learning:
For introductory content, focused on navigating the Schrödinger Materials Science interface, an Introduction to Materials Science Maestro tutorial is available. Please visit the materials science training website for access to 70+ tutorials. For scientific inquiries or technical troubleshooting, submit a ticket to our Technical Support Scientists at help@schrodinger.com.
For self-paced, asynchronous, online courses in Materials Science modeling, including access to Schrödinger software, please visit the Schrödinger Online Learning portal on our website.
For some related practice, proceed to explore other relevant tutorials:
- Introduction to Geometry Optimizations, Functionals and Basis Sets
- Introduction to Multistage Quantum Mechanical Workflows
- Optoelectronic Device Designer
- Locating Transition States: Part 1
- Locating Transition States: Part 2
- R-group enumeration with the Materials Science Suite
- Organometallic Complexes
- Band Shape
- Excited State Analysis
- Singlet-Triplet Intersystem Crossing Rate
- Optoelectronics Active Learning
- Calculating Transition Dipole Moments
- Bond and Ligand Dissociation Energy
- Molecular Deposition
- Liquid Electrolyte Properties: Part 1
- Liquid Electrolyte Properties: Part 2
For further reading:
- Organic Electronics overview from the Schrödinger website
- Help documentation on Optoelectronic Properties
- Atomistic-scale Simulation for the Analysis, Optimization and Accelerated Development of Organic Optoelectronic Materials. DOI:10.11370/isj.54.561
- Estimation of charge carrier mobility in amorphous organic materials using percolation corrected random-walk model. DOI:10.1016/j.orgel.2015.11.021
- Virtual screening for OLED materials. DOI:10.1117/12.2066565
- High-throughput quantum chemistry and virtual screening for OLED material components. DOI:10.1117/12.2025092
- Virtual screening of electron acceptor materials for organic photovoltaic applications. DOI:10.1088/1367-2630/15/10/105029
6. Glossary of Terms
Entry List - a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
Included - the entry is represented in the Workspace, the circle in the In column is blue
Project Table - displays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data
Recent actions - This is a list of your recent actions, which you can use to reopen a panel, displayed below the Browse row. (Right-click to delete.)
Scratch Project - a temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project
Selected - (1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Working Directory - the location where files are saved
Workspace - the 3D display area in the center of the main window, where molecular structures are displayed