Evaporation
Tutorial Created with Software Release: 2026-1
Topics: Consumer Packaged Goods , Organic Electronics , Polymeric Materials , Thin Film Processing
Methodology: All-Atom Molecular Dynamics , Coarse-Grained Modeling
Products Used: Desmond , MS Maestro
|
2.1 GB |
This tutorial is written for use with a 3-button mouse with a scroll wheel.
Words found in the Glossary of Terms are shown like this: Workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Abstract:
In this tutorial, we will learn to use the evaporation calculation and viewer panels to remove solvent molecules from all-atom and coarse-grained systems and equilibrate the systems using molecular dynamics simulations.
Tutorial Content
1. Introduction to the Evaporation Workflow
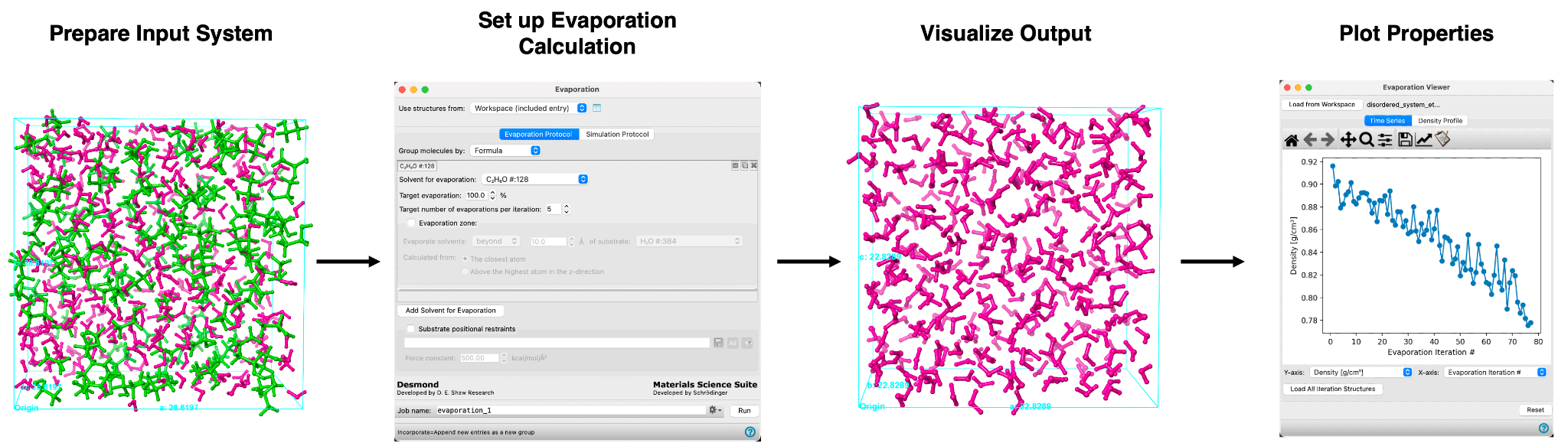
The Evaporation Calculation and Viewer panels allow you to simulate the removal of solvent molecules from a condensed system (e.g. polymer or molecular solid). In Schrödinger’s Materials Science (MS) Maestro suite, solvent molecules can be removed from an all-atom or coarse-grained periodic condensed system following a straightforward workflow as summarized here:
The calculation involves two stages that proceed iteratively. First, unless otherwise specified, a random subset of molecules are removed from anywhere in the unit cell as defined by the user. If an evaporation zone is specified, molecules are randomly removed within the boundaries of the set evaporation region. In the second stage, the system is relaxed using molecular dynamics (MD) simulation. These stages are performed iteratively until a target evaporation is reached. After the calculation is complete, a viewer panel is utilized to analyze the simulation results.
Evaporation calculations are useful for a variety of applications, for example, the drying of polymeric materials or the drying of thin films for organic electronics applications. The impact of changes in solvent composition on solution processed structures and their properties can be better understood using this tool.
In this tutorial, we will study three examples of evaporation; (1) in Section 3 we will remove all of the ethanol molecules from an equilibrated amorphous cell of ethanol-water, (2) in Section 4 we will remove chlorobenzene molecules from an equilibrated thin film system, and (3) in Section 5 we will evaporate multiple solvents from a coarse-grained structure.
For a more in depth explanation of building and equilibrating disordered systems, as provided in this tutorial, see the Disordered System Building and Molecular Dynamics Multistage Workflows tutorial.
2. Creating Projects and Importing Structures
At the start of the session, change the file path to your chosen Working Directorythe location where files are saved in MS Maestro to make file navigation easier. Each session in MS Maestro begins with a default Scratch Projecta temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project, which is not saved. A MS Maestro project stores all your data and has a .prj extension. A project may contain numerous entries corresponding to imported structures, as well as the output of modeling-related tasks. Once a project is saved, the project is automatically saved each time a change is made.
Structures can be built in MS Maestro or can be imported using File > Import Structures (or drag-and-dropped), and are added to the Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data. The Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion is located to the left of the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed. The Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data can be accessed by Ctrl+T (Cmd+T) or Window > Project Table if you would like to see an expanded view of your project data.
- Double-click the Materials Science icon
- (No icon? See Starting Maestro)
- Go to File > Change Working Directory
- Find your directory, and click Choose
- Pre-generated files are included for running jobs or examining output. Download the zip file here: schrodinger.com/sites/default/files/s3/release/current/Tutorials/zip/evaporation.zip
- After downloading the zip file, unzip the contents in your Working Directorythe location where files are saved for ease of access throughout the tutorial
- Go to File > Save Project As
- Change the File name to evaporation_tutorial, click Save
- The project is now named
evaporation_tutorial.prj
- The project is now named
For our first example, our structure of interest has been built in advance and is available for importing in the next step. We will import the equilibrated cell consisting of ethanol and water. This multi-component box contains 128 molecules of ethanol and 384 molecules of water. It has been equilibrated using a Compressive relaxation protocol followed by a 10 ns MD simulation.
If you are interested in building and equilibrating the system yourself, feel free to do so. For practice building and equilibrating multi-component systems, see the Disordered System Building and Molecular Dynamics Multistage Workflows tutorial. Machine Learning Force Fields (MLFF) are available as an alternative to OPLS force fields in the preparation process. Additional information regarding MLFF can be found in the help documentation or on our website.
- Go to File > Import Structures
- Navigate to where you downloaded the tutorial files (presumably your working directory) and choose
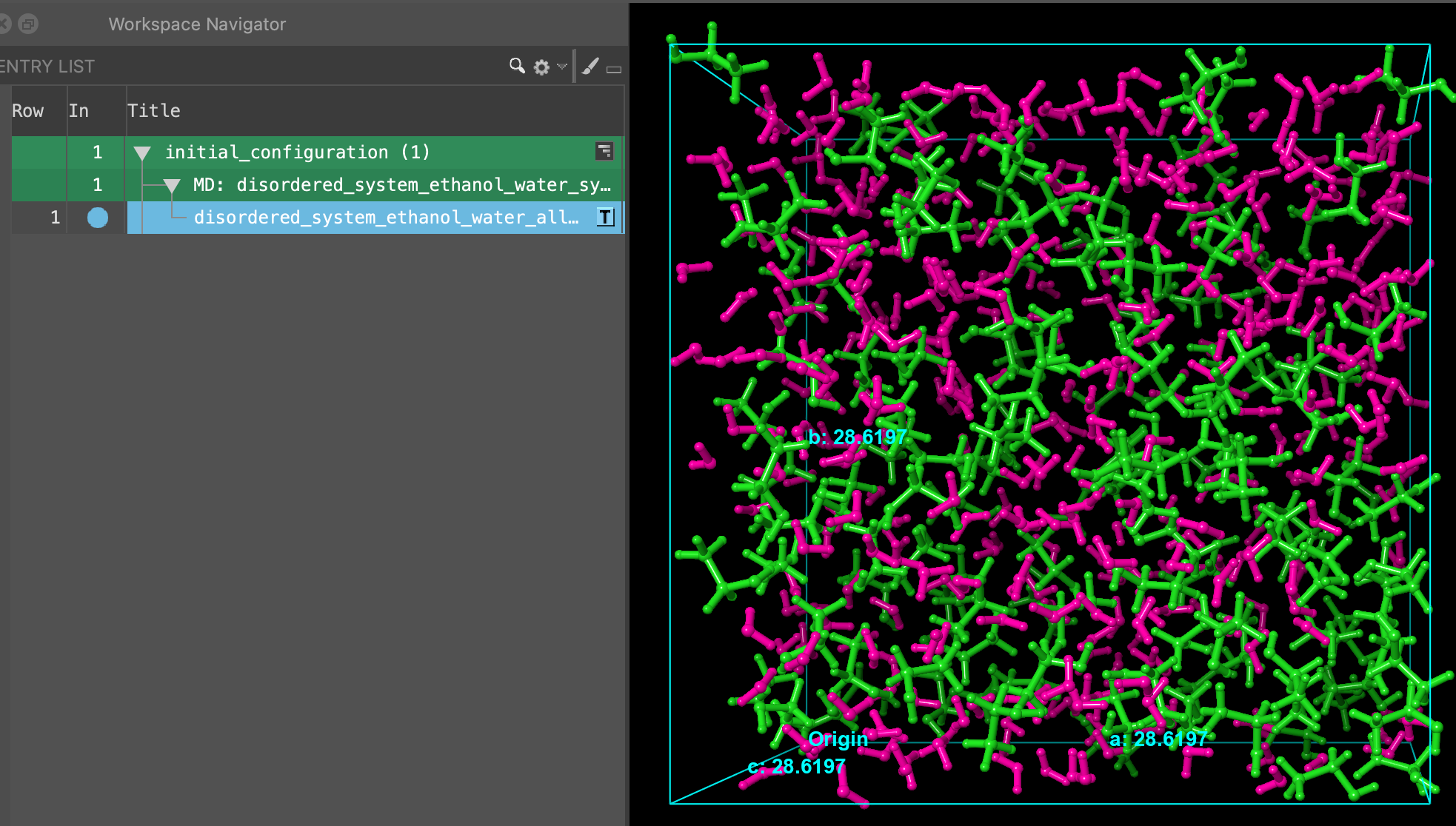
initial_configuration.cms. Click Open- A new entry group is added to the entry list containing an entry titled disordered_system_ethanol_water_all_components_amorphous
3. Evaporation from an All-Atom Multi-Component Box
In this section, we will use the Evaporation panel to remove all of the ethanol molecules from our ethanol-water box. The input system should be a well-relaxed MD output, as we have provided here. We will then visualize the output of the evaporation calculation more quantitatively in the Evaporation Viewer panel. If you would like to build and equilibrate such a system yourself, please see the Disordered System Building and Molecular Dynamics Multistage Workflows tutorial.
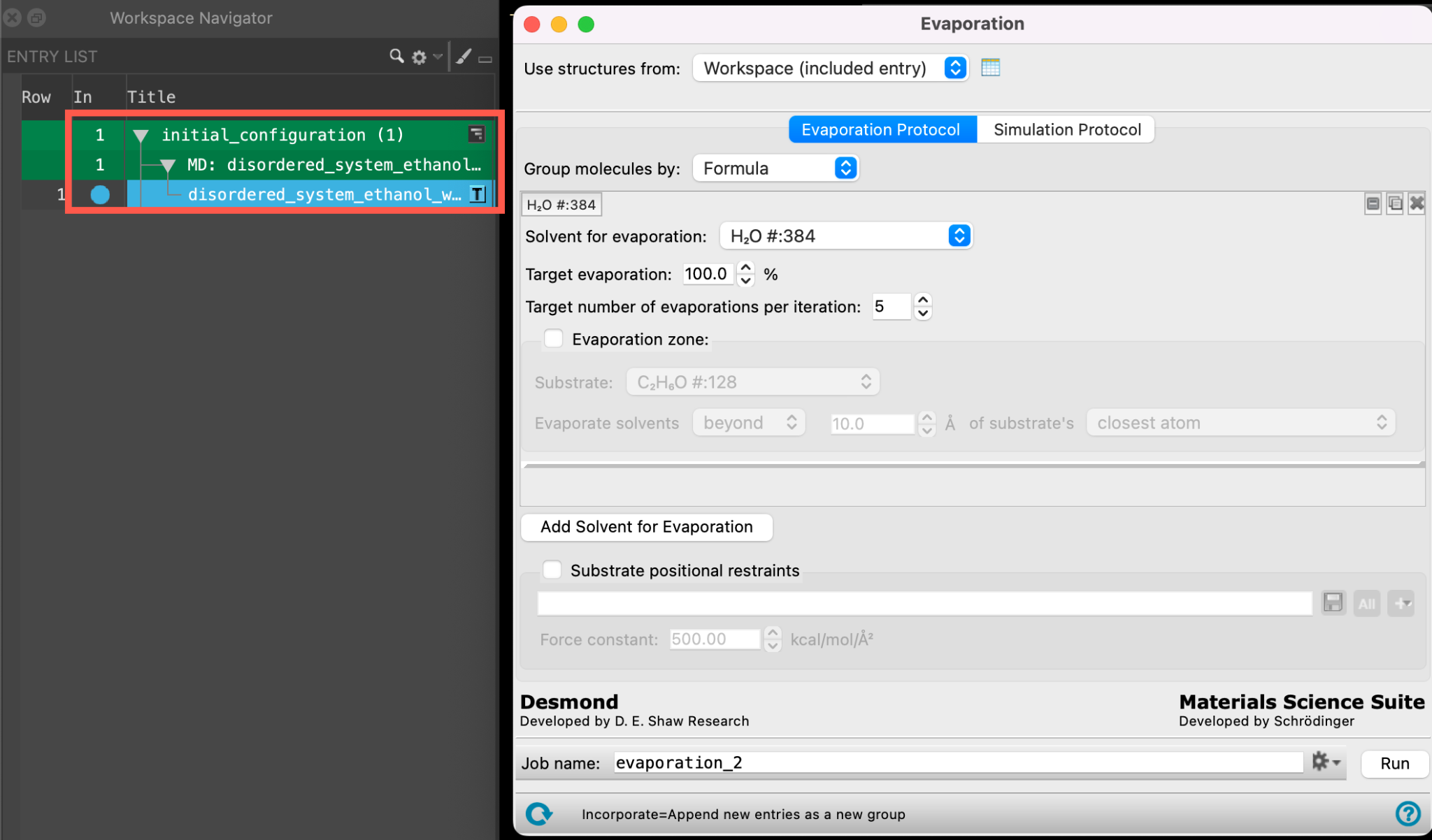
- Ensure that disordered_system_ethanol_water_all_components_amorphous is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Classical Mechanics > Evaporation > Evaporation Calculations
- The Evaporation panel opens
Let’s learn about the settings and capabilities of the Evaporation panel a bit more:
- In the Evaporation Protocol tab, parameters for the evaporation of a solvent(s) are specified. In most cases it is best to evaporate the solvent slowly from the system so that it has a chance to equilibrate after a few molecules are removed.
- Molecules that can be removed using the evaporation protocol can be grouped by Formula, SMILES, or Chiral SMILES.
- The Add Solvent for Evaporation button allows you to add additional solvents to remove from the system.
- For each solvent selected for evaporation, distance based evaporation can be enabled by selecting Evaporation zone. The Evaporation zone defines the regions of a unit cell from which to remove solvent molecules.
- Substrate positional restraints allow you to select atoms to restrain during the calculation using atomic specification language (ASL)
- The Simulation Protocol tab allows us to specify parameters for the MD simulations between evaporation iterations. The Simulation time sufficient to equilibrate the cell depends on how many molecules of solvent are being removed.
- The NPT Ensemble class will hold the pressure constant and allow the volume of the unit cell to change, which may be good for bulk systems that contract upon evaporation.
- The NVT Ensemble class will hold the volume constant which is useful for cases in which we do not want uniform shrinking of the cell, such as for a surface.
- Data is generated for each iteration of evaporating and equilibrating and is stored by default. The Remove subdirectories option can be toggled on and off to keep and remove the trajectory information from each iteration.
Visit the help documentation for a complete summary of the parameters.
Here we use the Evaporation panel to remove ethanol molecules from an ethanol-water box. It is important to note that we could remove water molecules instead of ethanol if we would like to. Note that the panel can also be used to evaporate multiple solvents from all-atom or coarse-grained systems, as demonstrated in Section 5.
Now, let’s specify the evaporation parameters for this example.
- Retain Formula for Group molecules by
- We will choose our solvent for evaporation in this example based on its molecular formula
- You can additionally identify molecules by their SMILES or Chiral SMILES for solvent selection
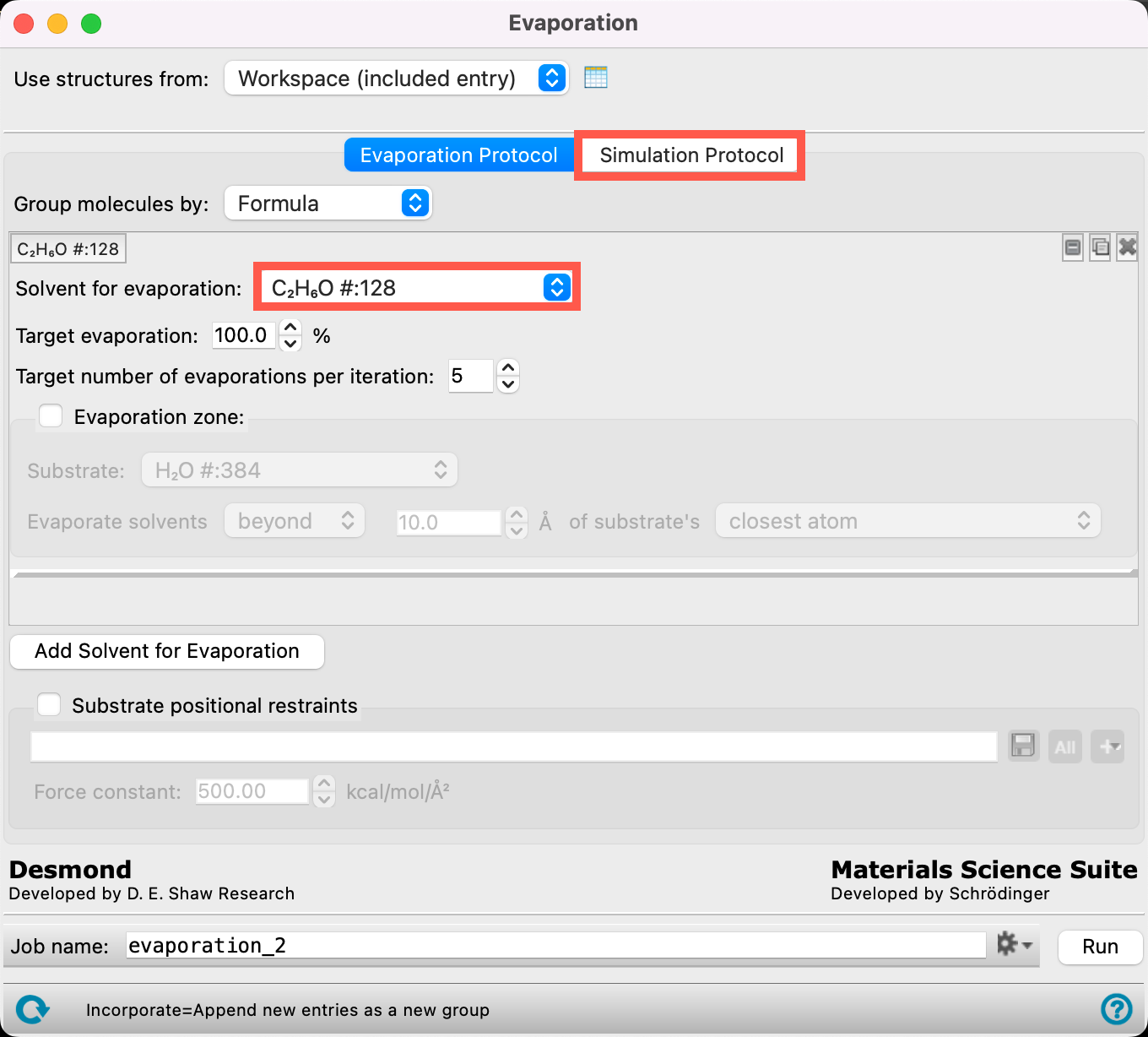
- From the Solvent for evaporation dropdown menu, select C2H6O #:128
- The 128 ethanol molecules have now been identified as the target molecules for the evaporation calculation
We will leave the defaults for Target evaporation and Target number of evaporations per iteration. This means that we will remove at most 5 ethanol molecules per iteration until there are no ethanol molecules left. Feel free to adjust these settings for your systems of interest.
- Go to the Simulation Protocol tab
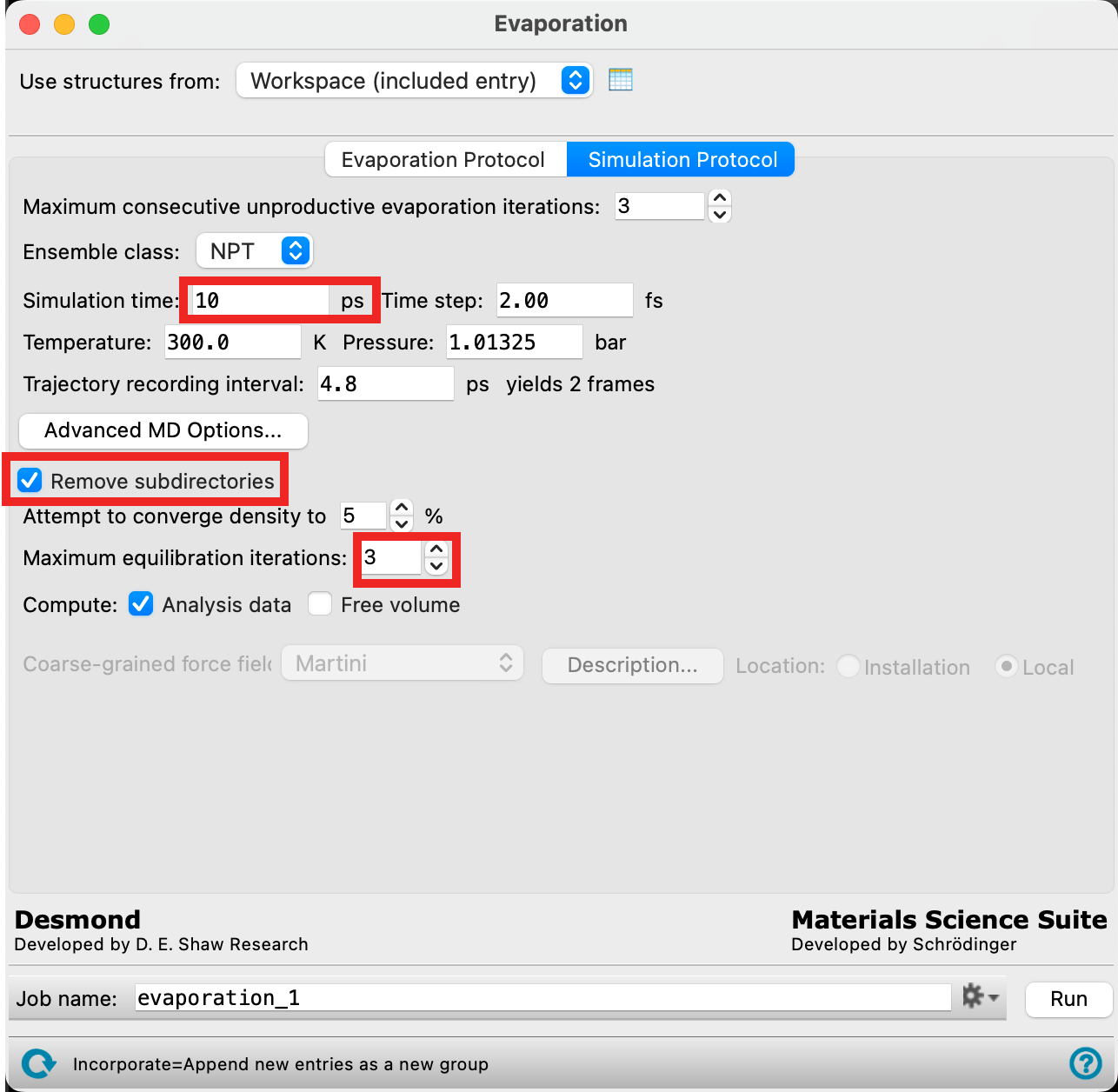
- Enter 10 ps for the Simulation time
- Check Remove subdirectories
- We will not be keeping the trajectory information for each iteration because the file sizes quickly become very large. If you would like to view the trajectory for each MD simulation, uncheck this box to ensure you have the relevant files
- For Maximum equilibration iterations enter 3
- After the molecules are removed from the system, the system will undergo equilibration. An attempt will be made to converge density to 5% as indicated in the panel, but if this fails, the equilibration procedure will be attempted again up to 3 times
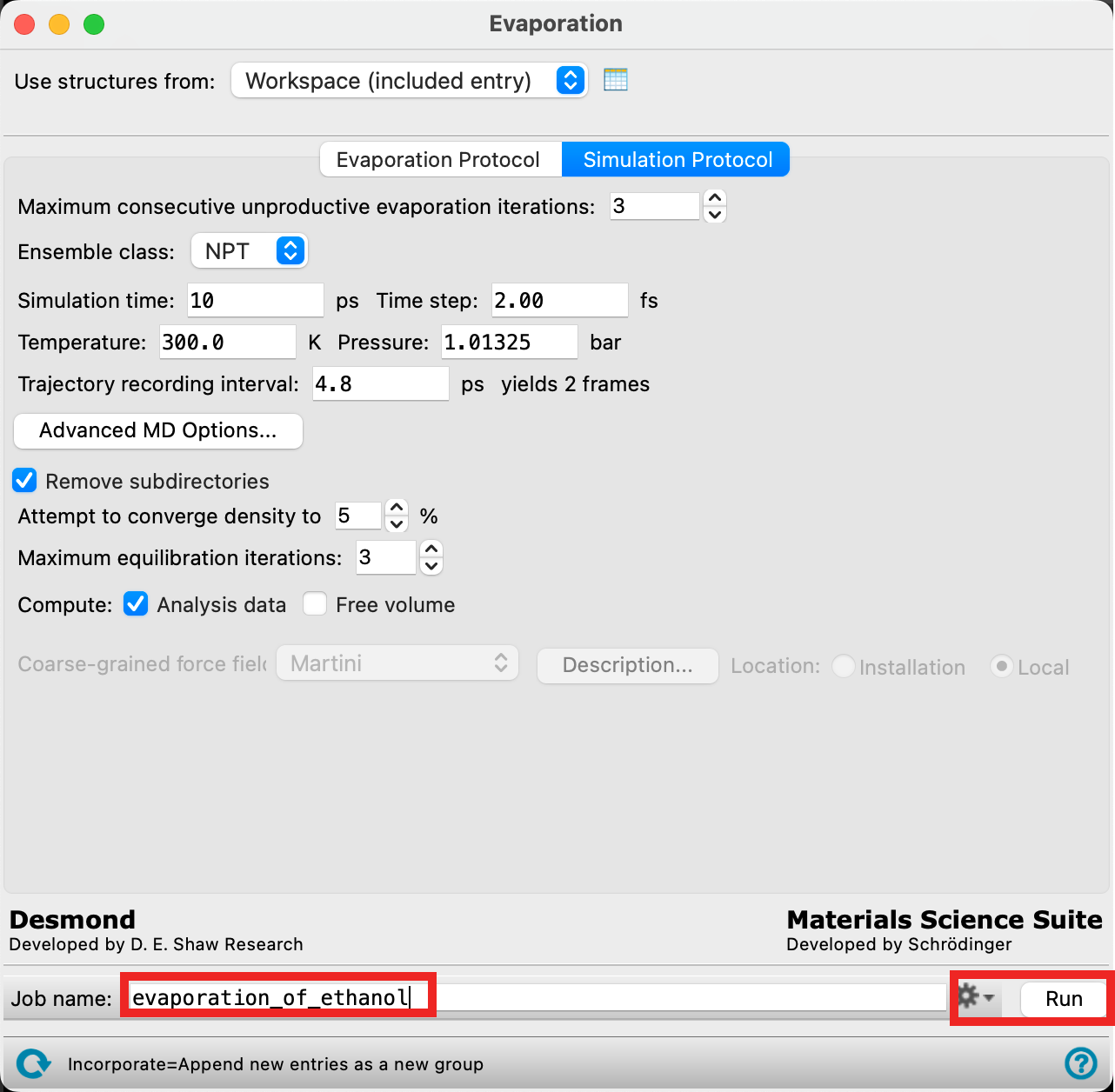
- Change the Job name to evaporation_of_ethanol
- Adjust the job settings (
 ) as needed
) as needed
- This job requires a GPU host. The job can be completed in one hour on a GPU host
- If you would like to run the job yourself, click Run. Otherwise, go to File > Import Structures, navigate to the provided tutorial files and Open
Section_03 > evaporation_of_ethanol > evaporation_of_ethanol-out.cms.gz - Close the Evaporation panel
- In general, always close the Evaporation panel after use. This panel is interactive with the workspace and leaving it open can cause slowdowns
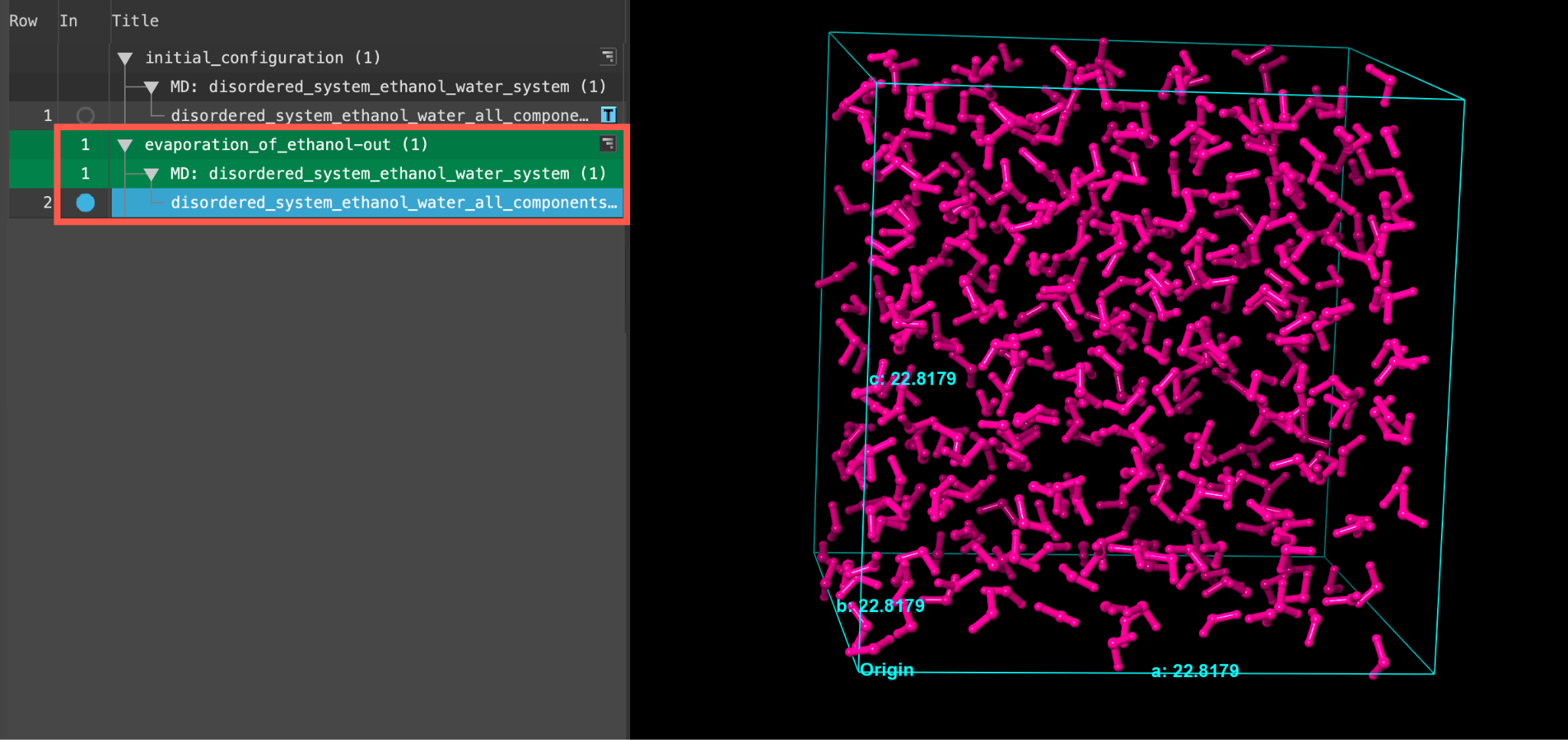
- When the job is finished or imported select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue the output disordered_system_ethanol_water_all_components_amorphous in the entry list.
- The displayed cell is from the last evaporation step. As shown in the Figure, only water atoms remain
- Feel free to stylize and visualize the output system in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
If you are interested in visualizing intermediate steps of the evaporation process, then you can manually import the relevant .maegz from the provided files.
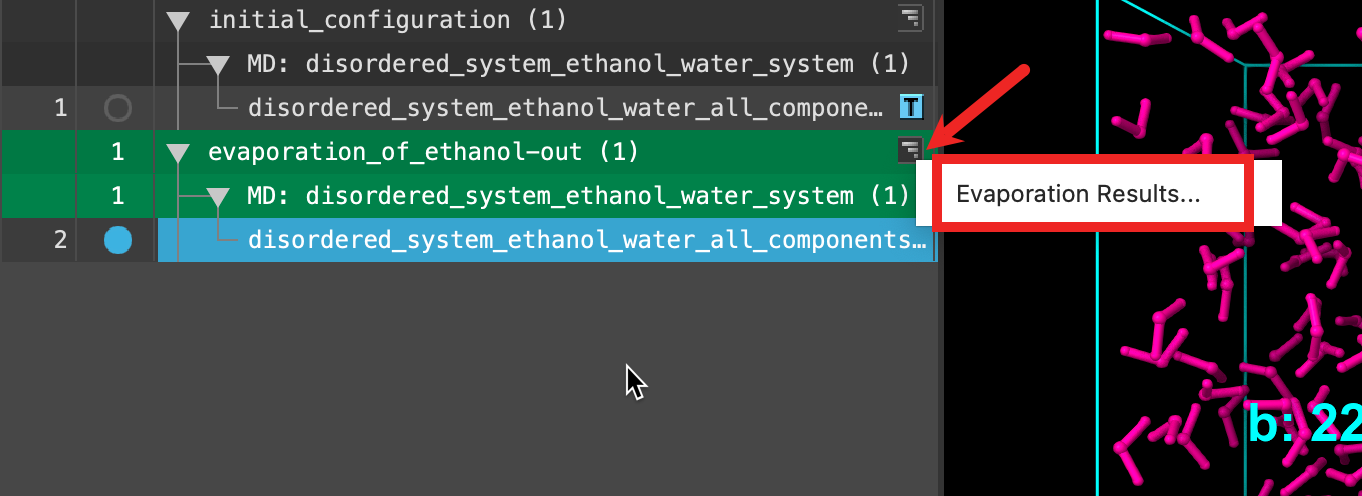
Now we will open the Evaporation Viewer panel to extract some quantitative metrics from our simulation. Despite the fact that only the last iteration is shown in the workspace, this panel provides analysis over the entire course of the calculation.
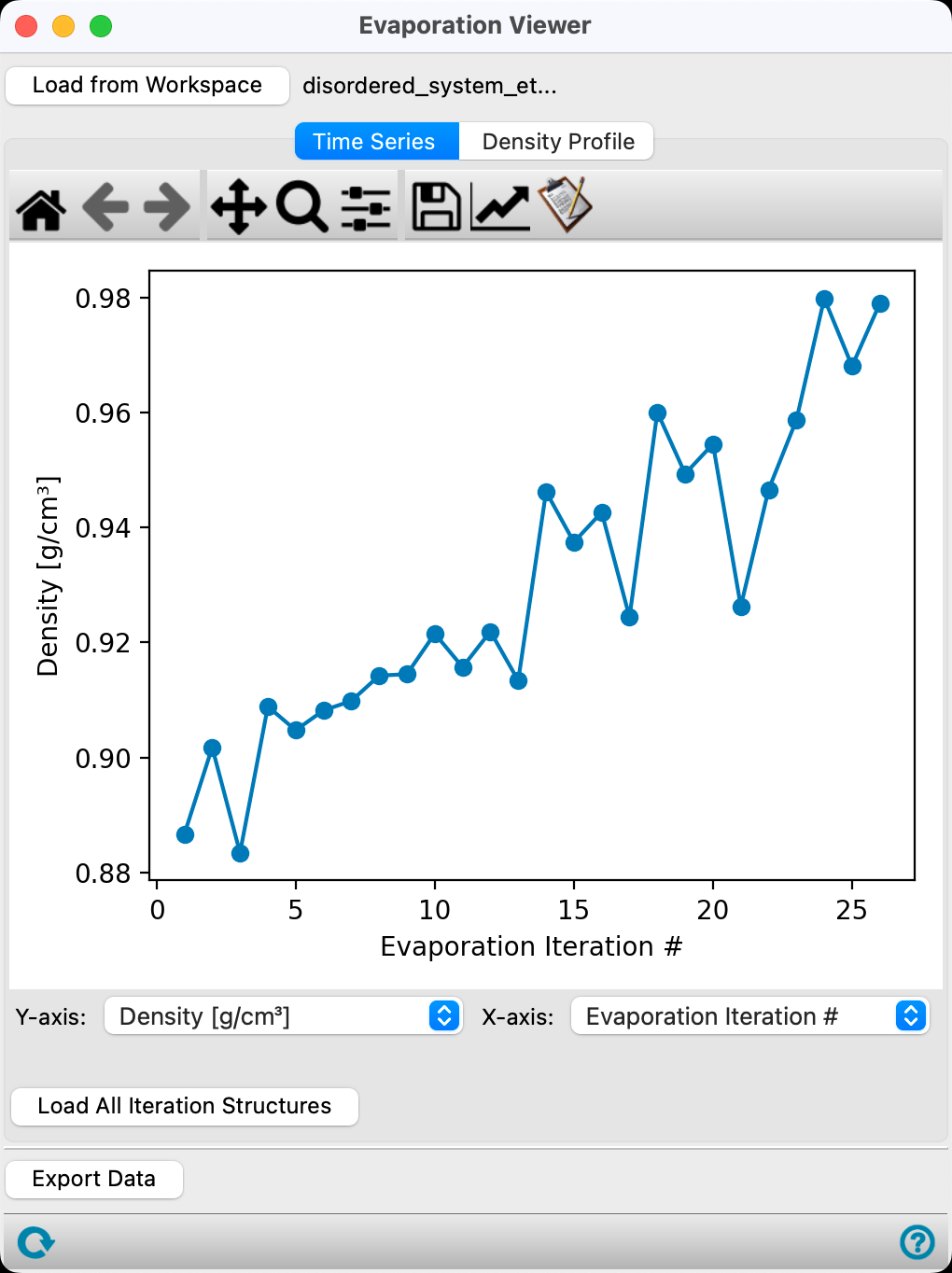
In the Time Series tab, the first plot provided is the Density (g/cm3) versus Evaporation iteration # for the calculation.
As expected, the density of the system increases as the less dense substance, ethanol, is removed. We approach the density of pure water as all the ethanol molecules are removed. Noise in the trend shown in the Figure can be reduced through increasing the equilibration time, decreasing the target number of evaporated molecules per iteration, and/or increasing the total system size.
Note: Clicking a data point in the plot will allow you to inspect the structure at that iteration of the evaporation calculation.
We can also analyze how the mass percent of a particular component changes with evaporation iteration. To do so,
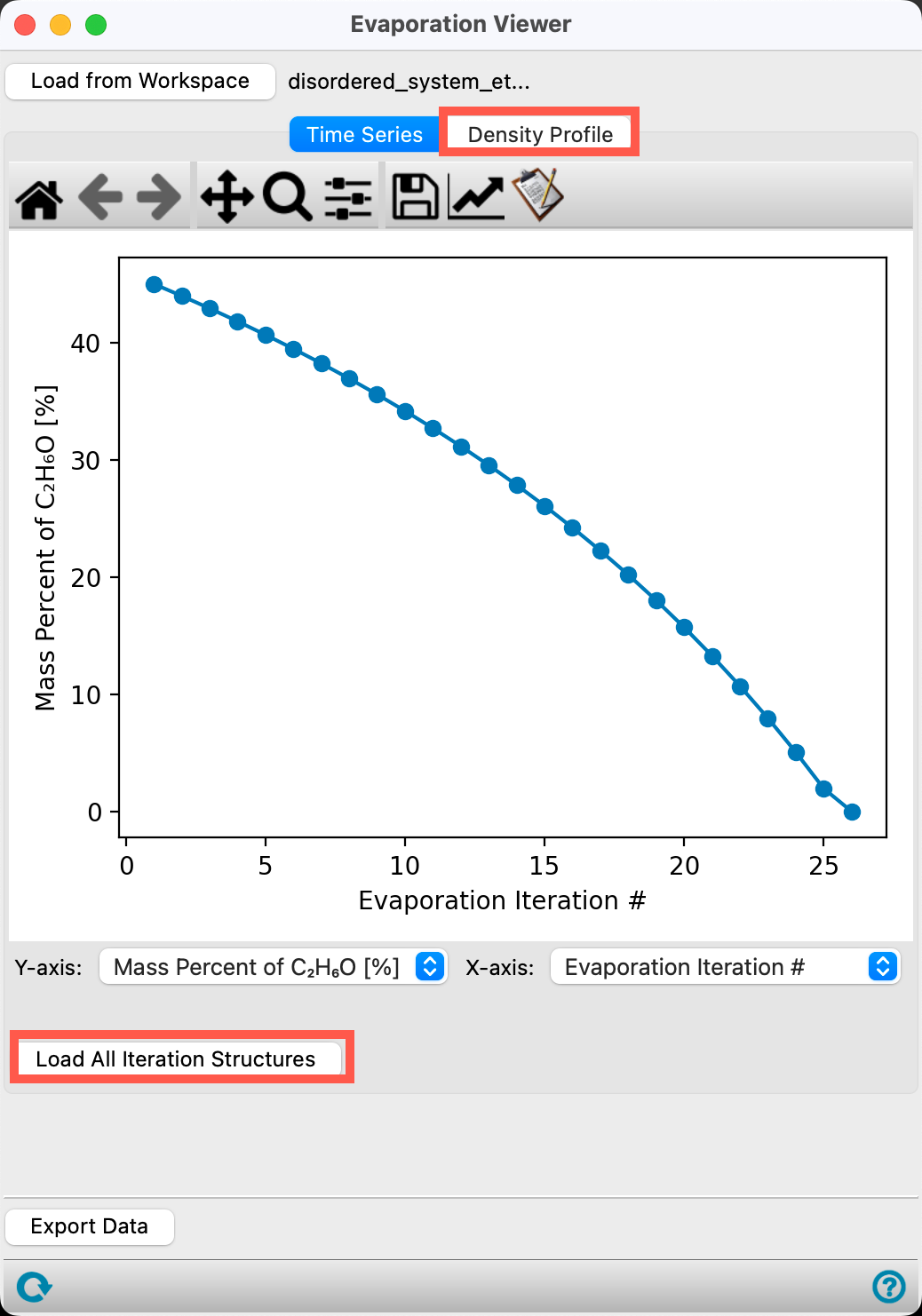
- Change the Y-axis to Mass Percent of C2H6O [%]
The plot updates to show the mass percent of C2H6O decreasing as a function of evaporation iteration. This is expected as the ethanol molecules are removed from the box.
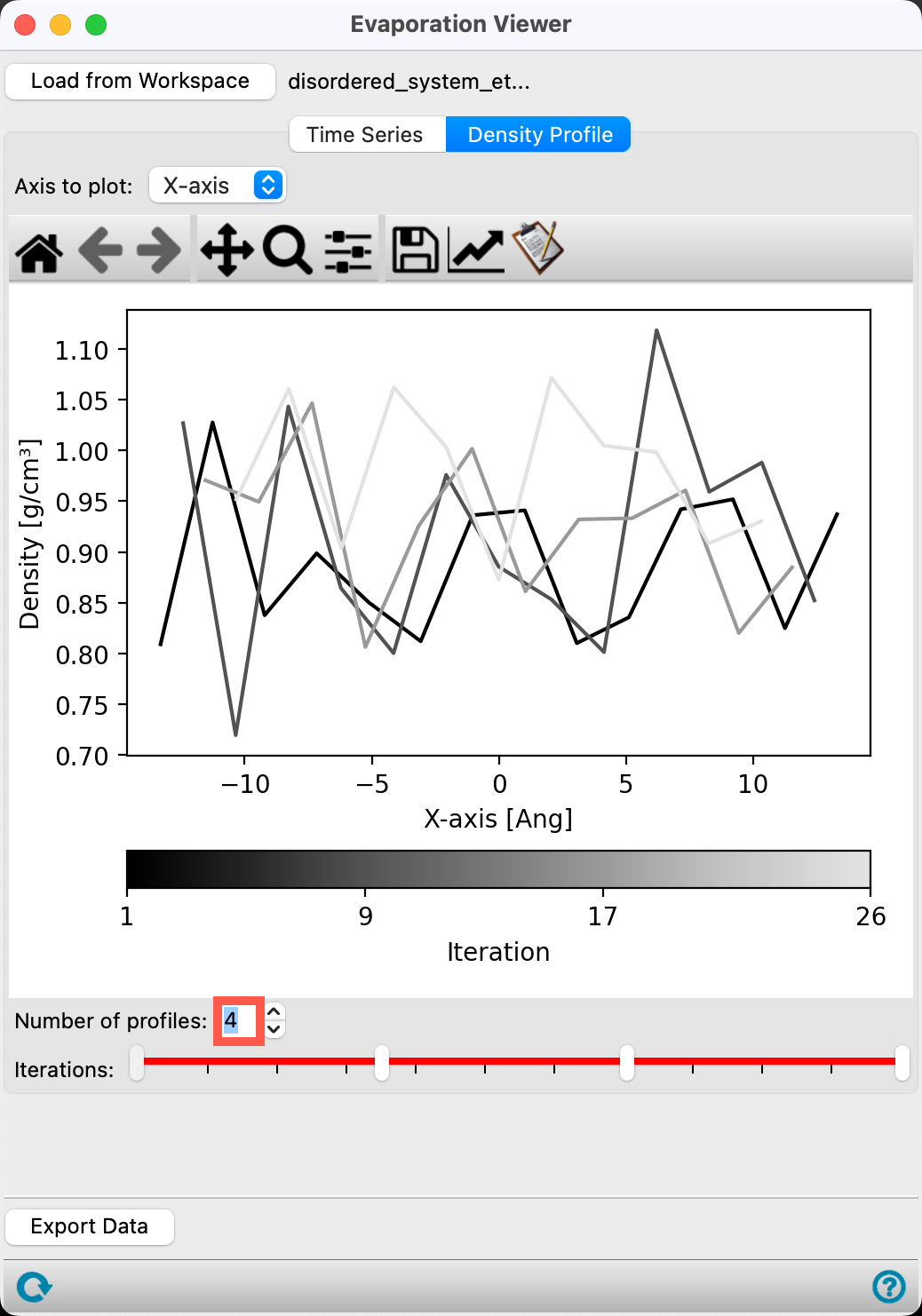
- Go to the Density Profile tab
In the Density Profile tab, the Density (g/cm3) is plotted versus the axis. The Axis to plot and Iterations to plot can be altered to view information of interest. By default, all iterations along the x-axis are plotted.
- Change the Number of profiles to plot to 4
- The density profiles for the 1st, 9th, 17th, and 26th (last) iterations are shown
Here, we study a simple two-component system. For a more complex system, such as a block copolymer, the Density Profile would allow you to visualize whether or not there are domains in the sample.
Feel free to explore the Viewer panel further.
4. Evaporation from a Thin Film
In this section, we will use the Evaporation panel to remove chlorobenzene molecules from a thin film system relevant in organic electronics. This system consists of a SiO2 substrate, a ~200 Å NPB (N,N′-Di(1-naphthyl)-N,N′-diphenyl-(1,1′-biphenyl)-4,4′-diamine) layer, and an emissive layer containing CBP (4,4′-Bis(N-carbazolyl)-1,1′-biphenyl ), Ir(ppy)3, and chlorobenzene. If you would like to build and equilibrate such a system yourself, please see the Building a Polymer-Polymer Interface Model tutorial.
- Go to File > Import Structures
- Navigate to where you downloaded the tutorial files (presumably your working directory) and choose
Section_04 > npb-cbp-ir-solvent_system-out.cms.Click Open- A new entry group is added to the entry list containing an entry titled npb-cbp-ir-solvent
- Ensure that npb-cbp-ir-solvent is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Classical Mechanics > Evaporation > Evaporation Calculations
- The Evaporation panel opens
- The Evaporation panel may take a few moments to load as this is a large system
Now, let’s specify the evaporation parameters for this example.
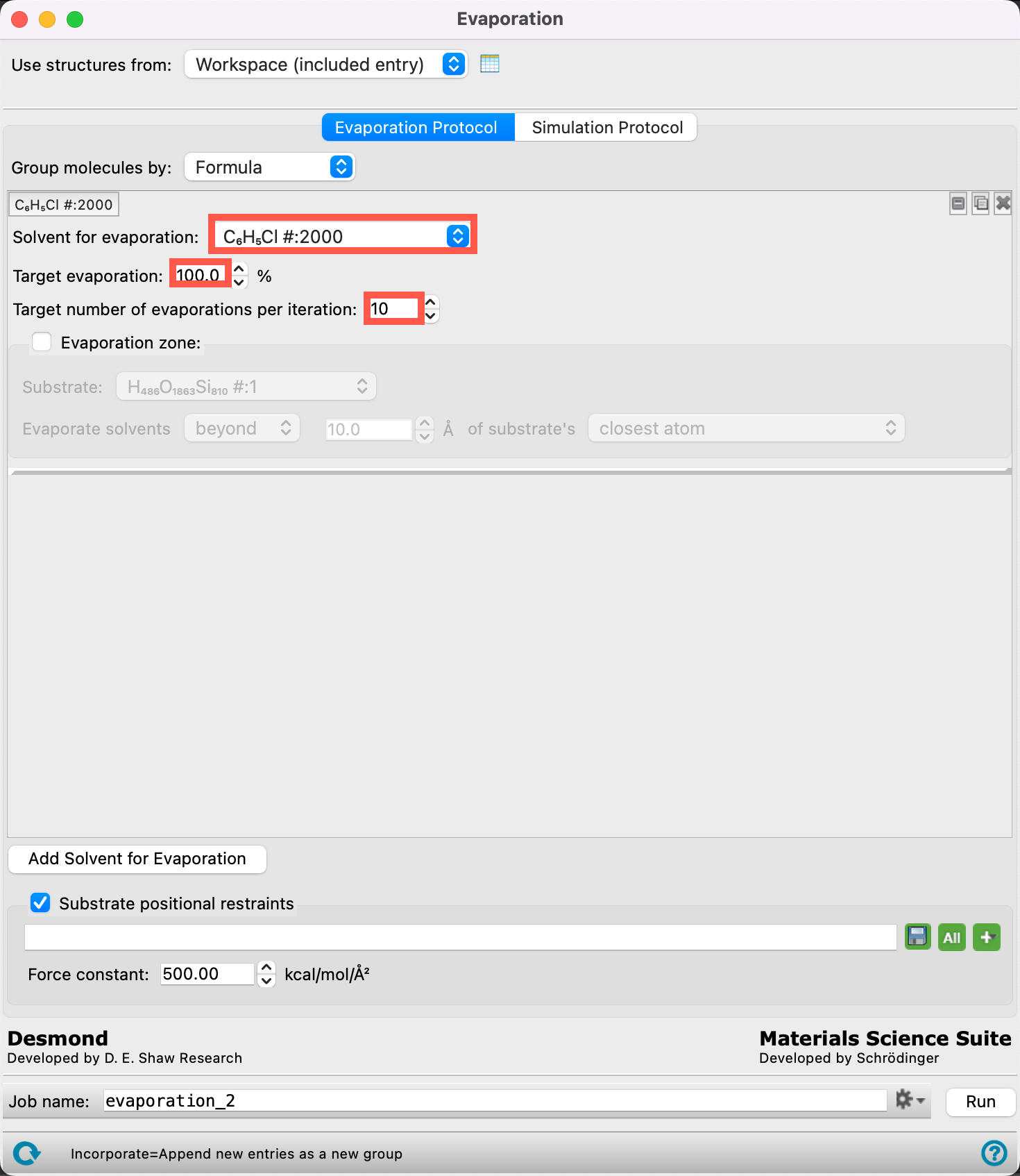
- Retain Formula for Group molecules by
- From the Solvent for evaporation dropdown menu, select C6H5Cl #:2000
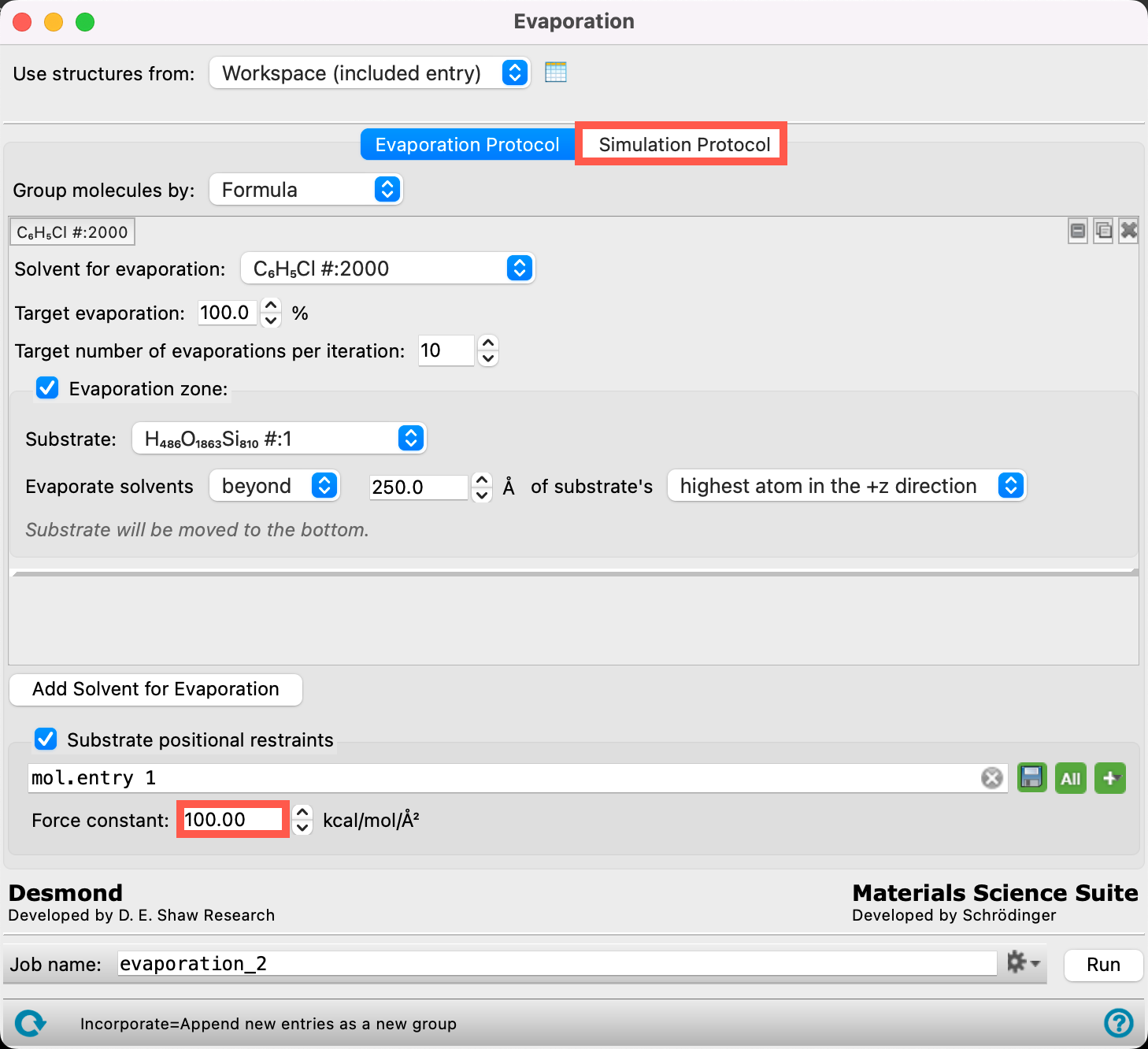
- The 2000 chlorobenzene molecules have now been identified as the target for the evaporation calculation
- Ensure the Target evaporation is 100%
- Change the Target number of evaporations per iteration to 10
- This means that we will remove at most 10 chlorobenzene molecules per iteration until there are no chlorobenzene molecules left.
Next, let’s define where in the structure we want to remove the chlorobenzene molecules from.
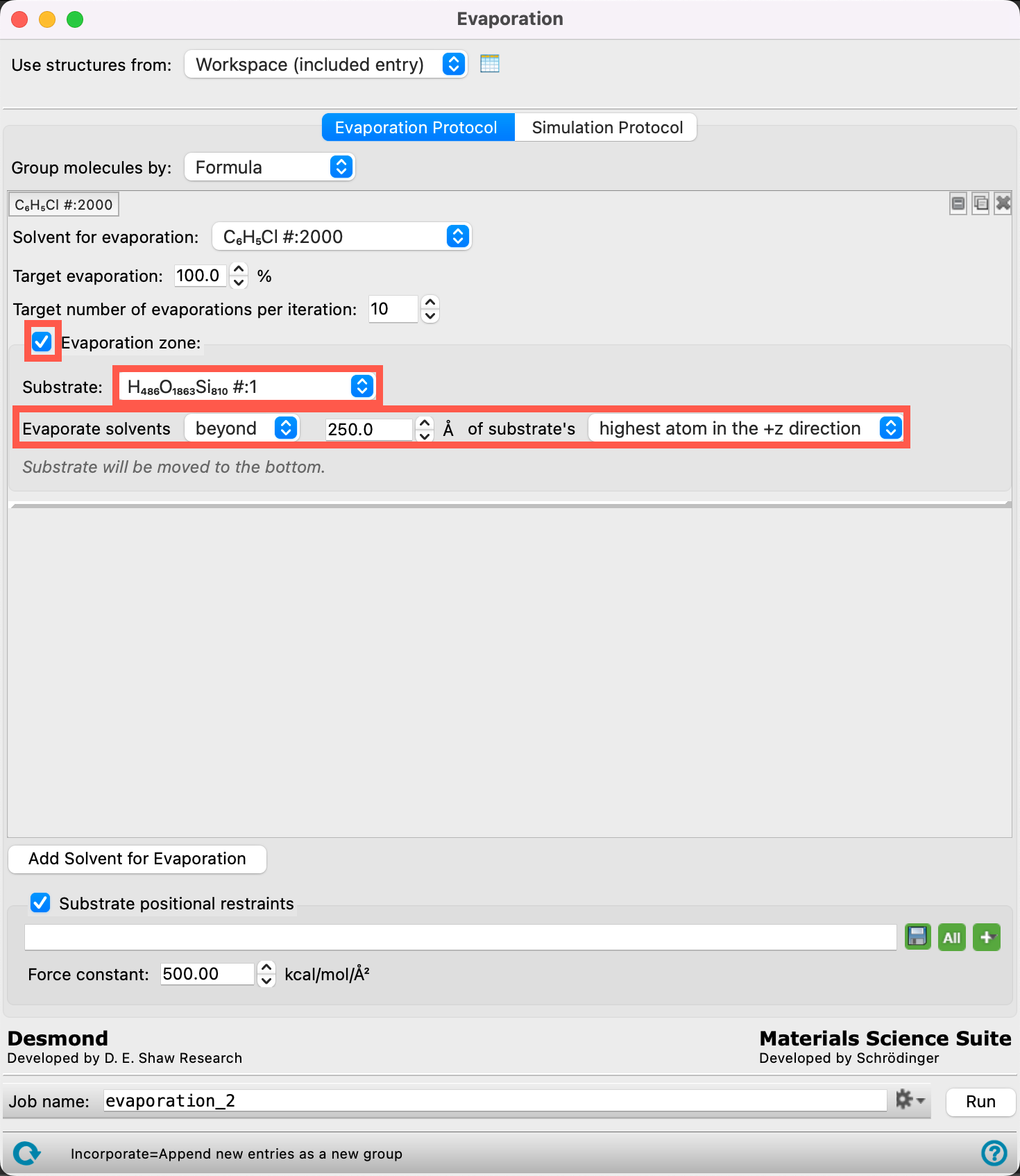
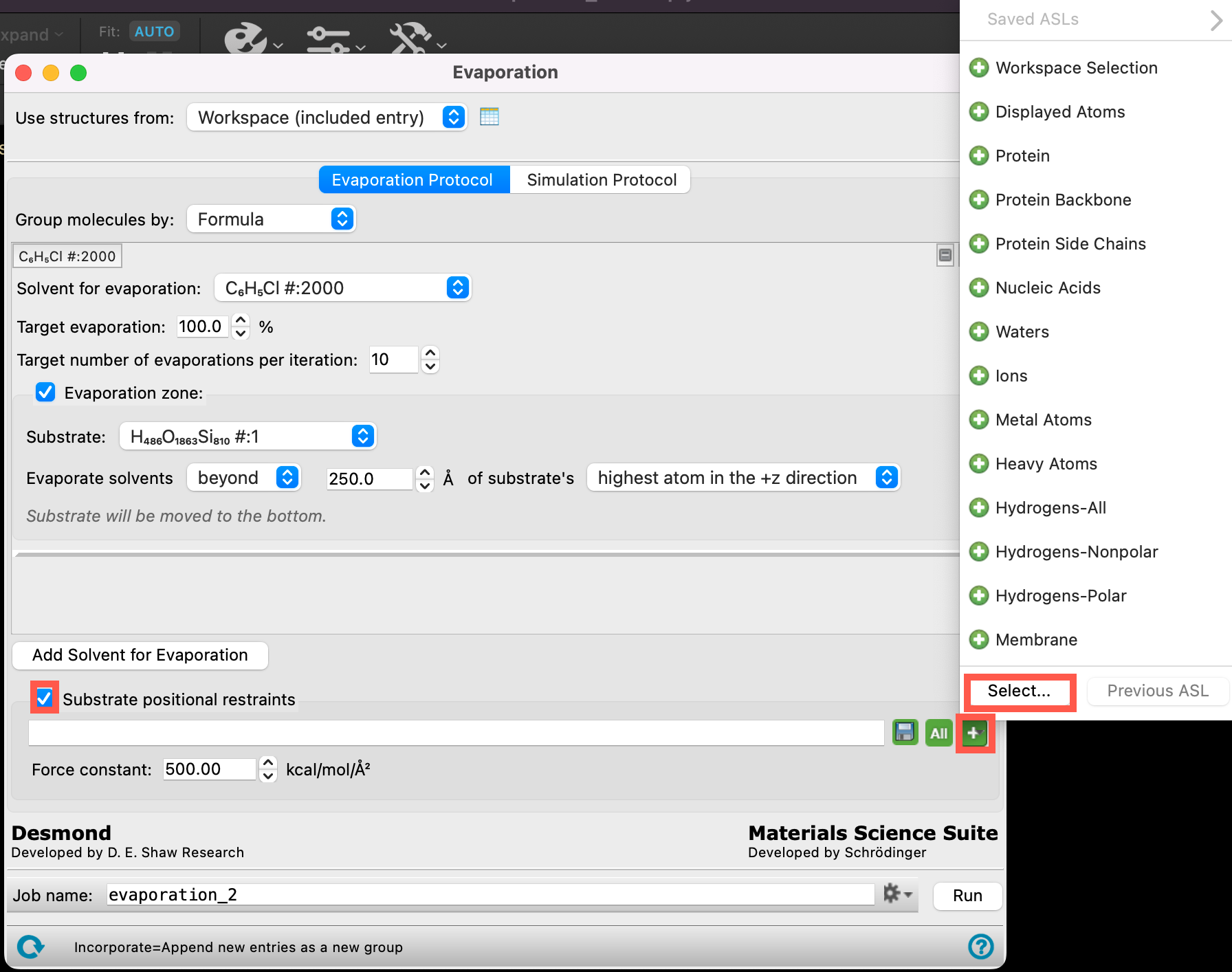
- Check Evaporation zone
-
Change the parameters to match those in the Figure:
- Substrate (H486O1863Si810#:1)
- Evaporate solvents beyond 250 Å of substrate’s highest atom in the +z direction
- We set the height of the evaporation zone as such because it is right above the thickness of the NPB layer + the estimated thickness of the emissive layer
During the evaporation protocol, there is potential for drifting (i.e. the whole SiO2 slab changing its position), particularly as more and more molecules are removed from the system. To prevent this, we can use the Substrate positional restraints section of the panel. Let’s restrain the SiO2 slab for our thin film system:
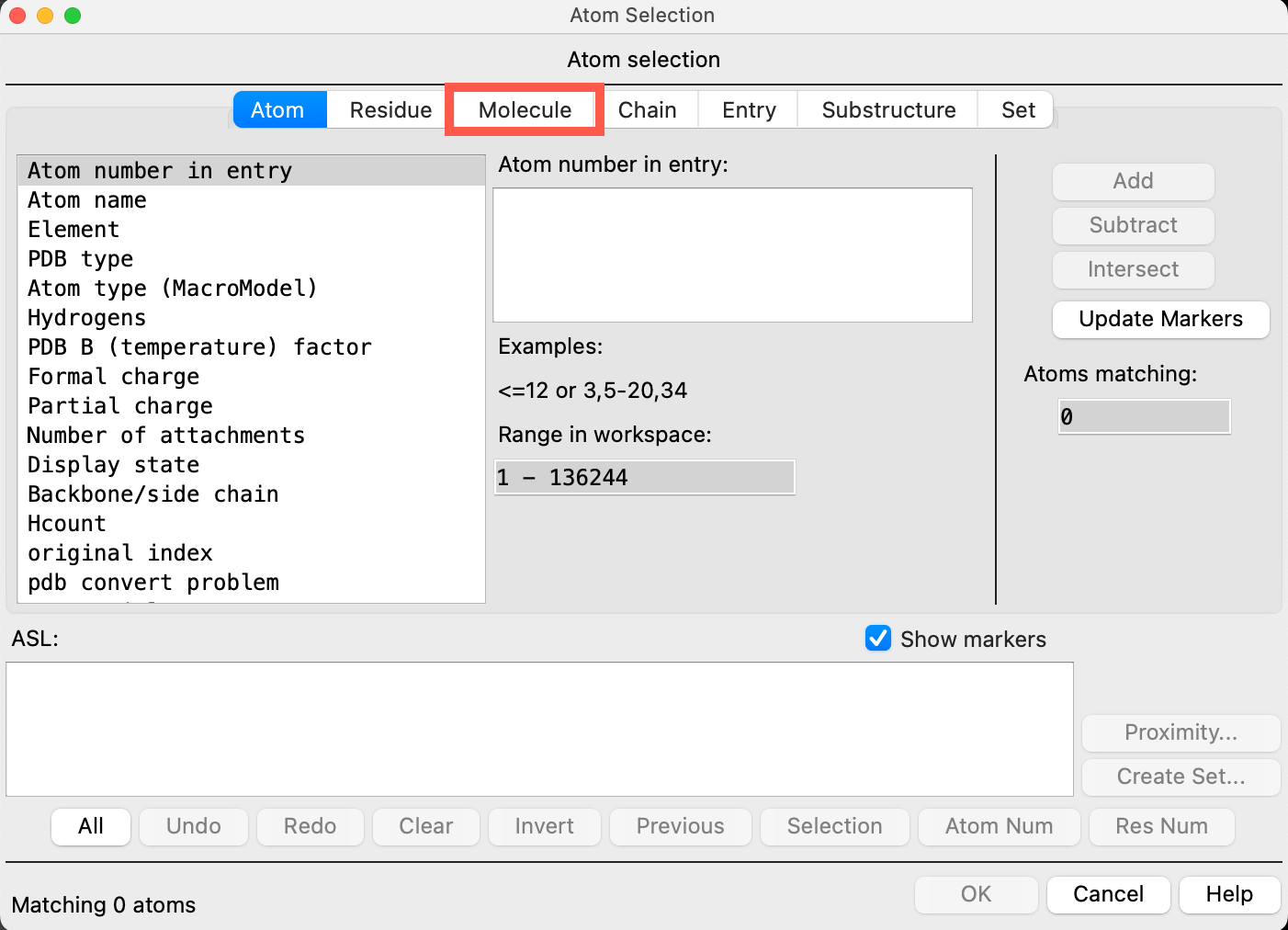
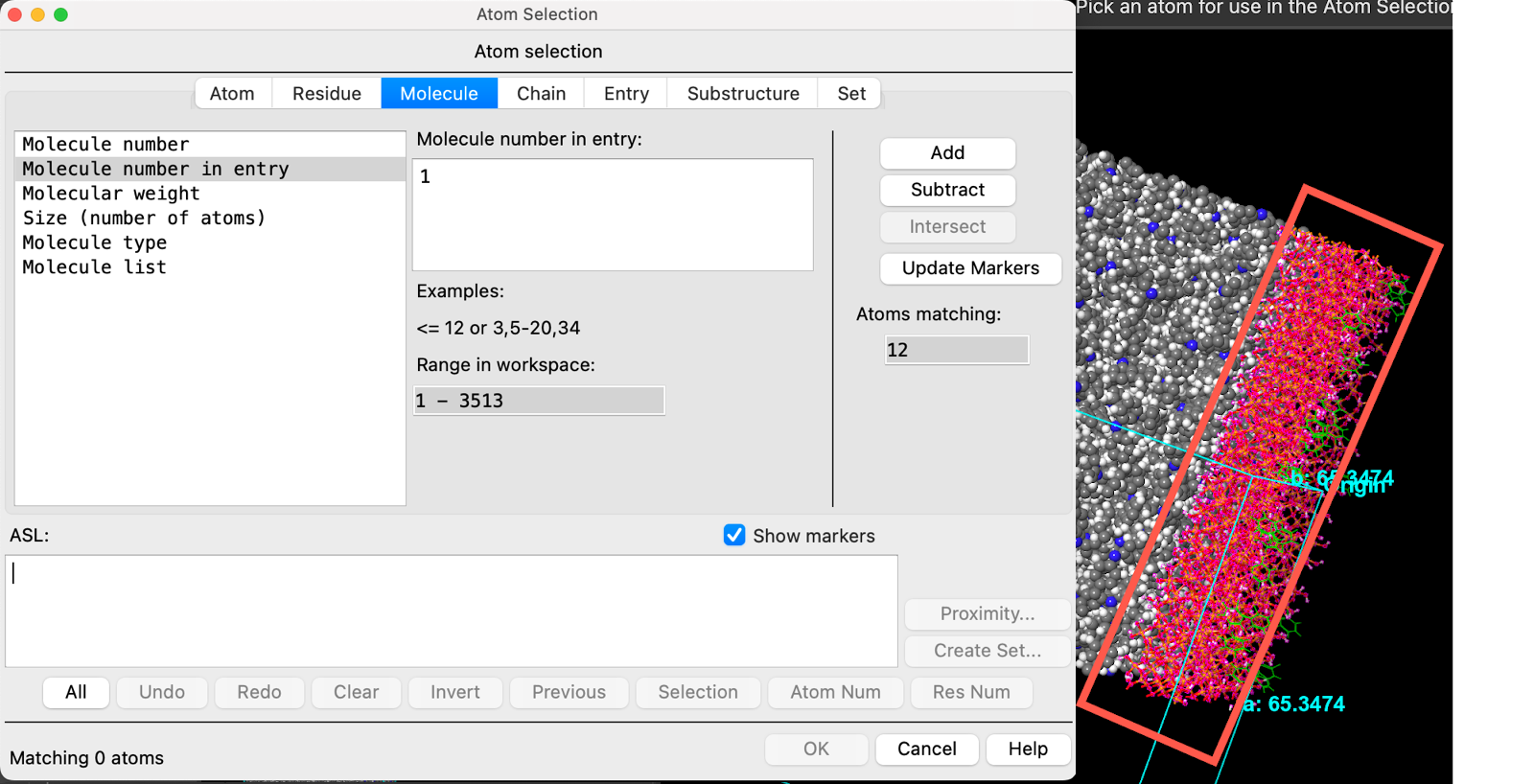
- Click on the SiO2 substrate in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
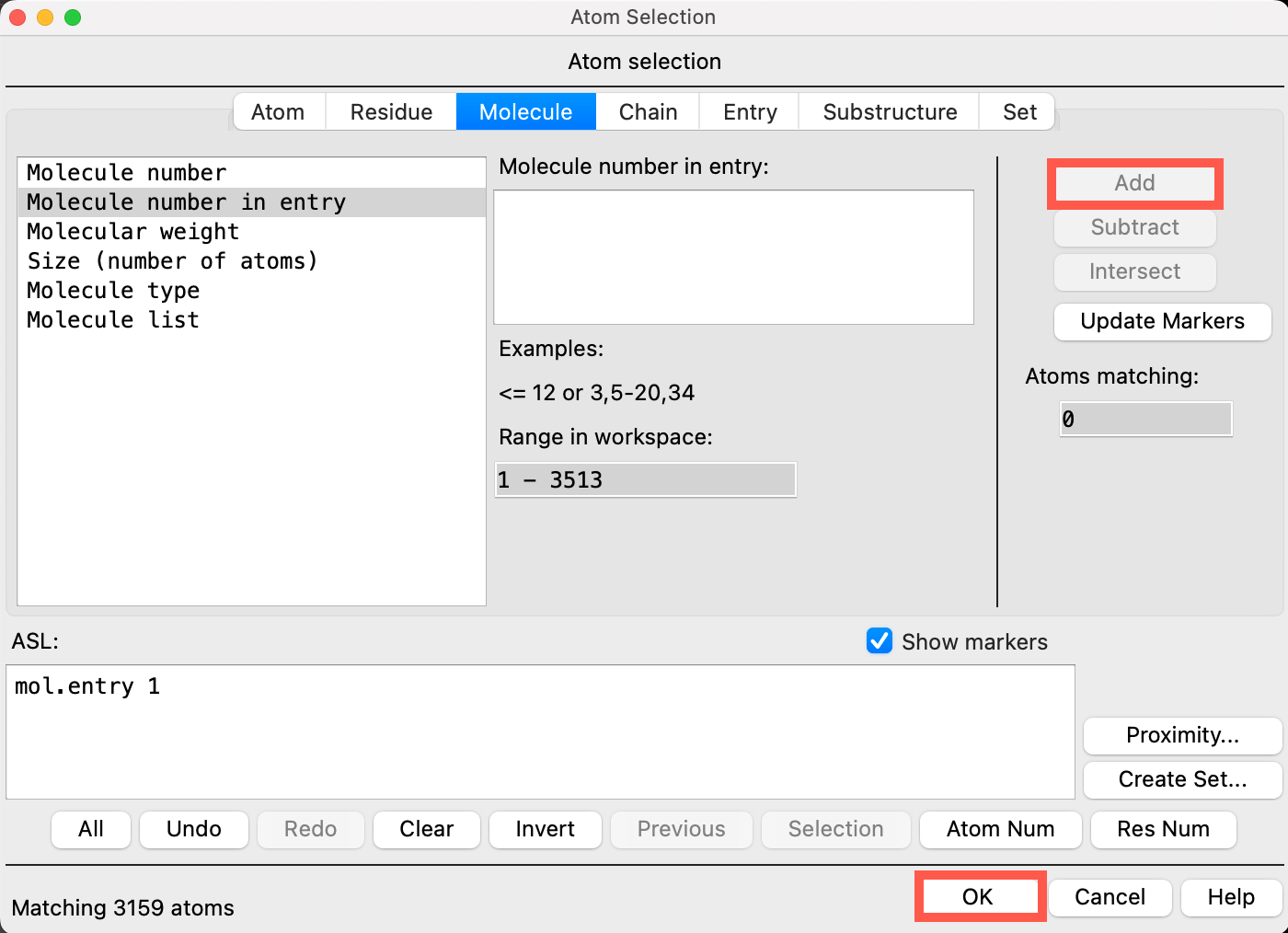
- The entire SiO2 workspace is recognized as molecule 1
- Click Add
- The SiO2 substrate is added to the ASL box at the bottom of the panel
- Click OK
- The SiO2 substrate will now be restrained during the evaporation calculation
- Change the Force constant to 100
- The force constant is associated with a positional restraint used to restrain a particle to a certain fixed position in the simulation volume. This restraint prevents the entire substrate from drifting in the simulation. 100 kcal/mol/Å2 is chosen, for this example, to be consistent with the Force constant used when this system was built using the Molecular Deposition workflow
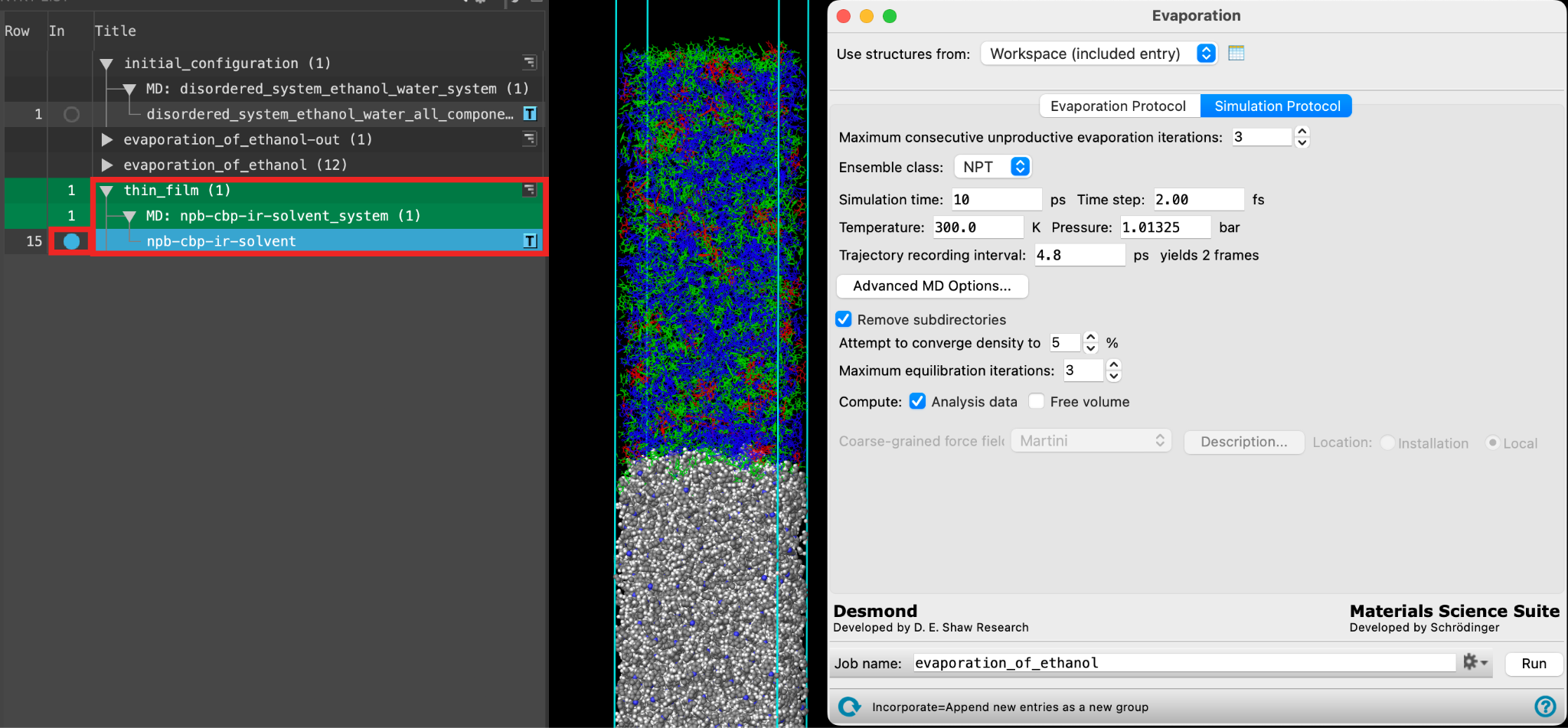
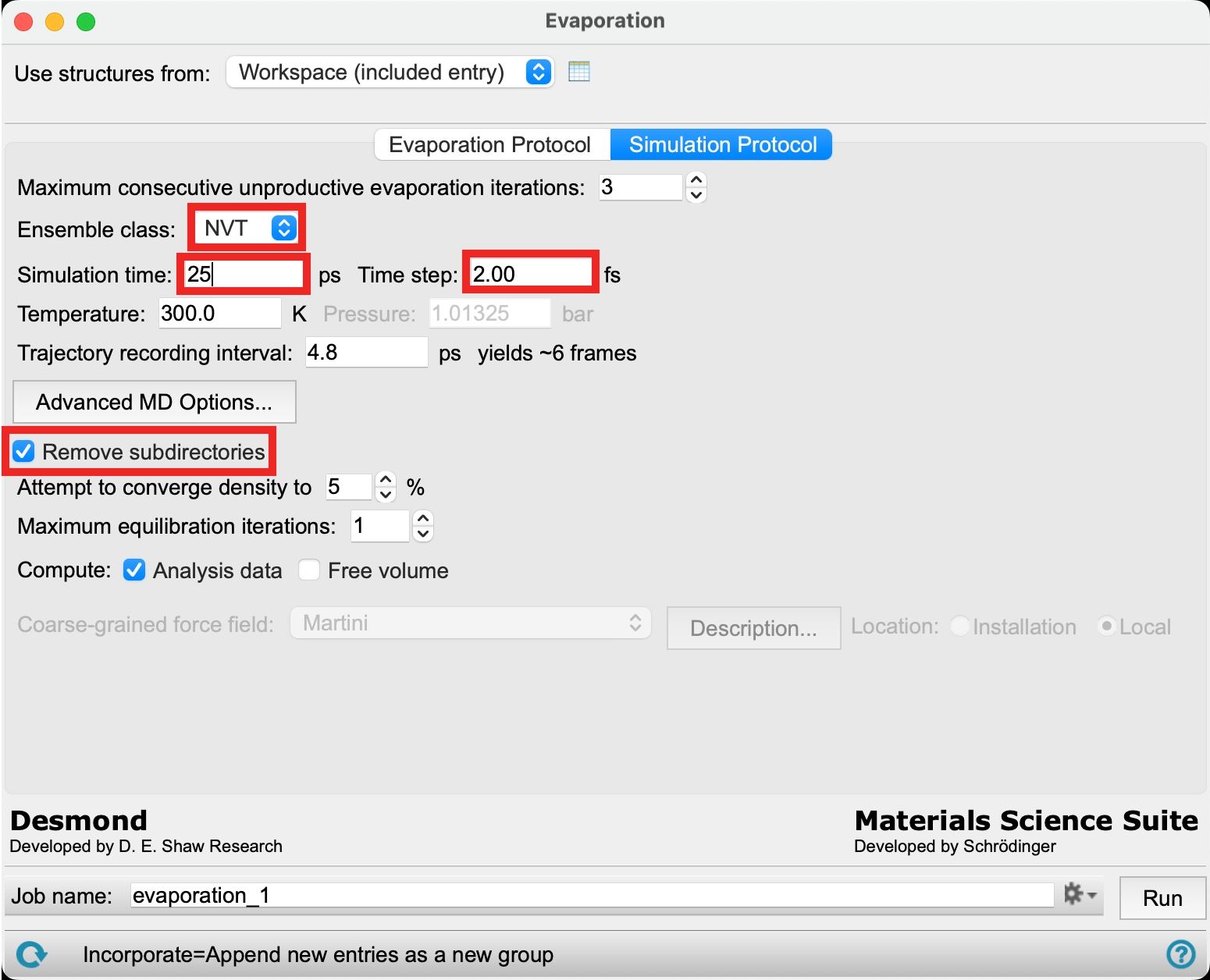
- Go to the Simulation Protocol tab
- Change the Ensemble class to NVT
- Enter 25 ps for the Simulation time
- For Time step enter 2 fs
- Check Remove subdirectories
- We will not be keeping the trajectory information for each iteration. If you would like to view the trajectory for each MD simulation, uncheck this box to ensure you have the relevant files
Note: The default Ensemble class of NPT is good for bulk systems, but NVT should be used for surfaces, as we do in this example.
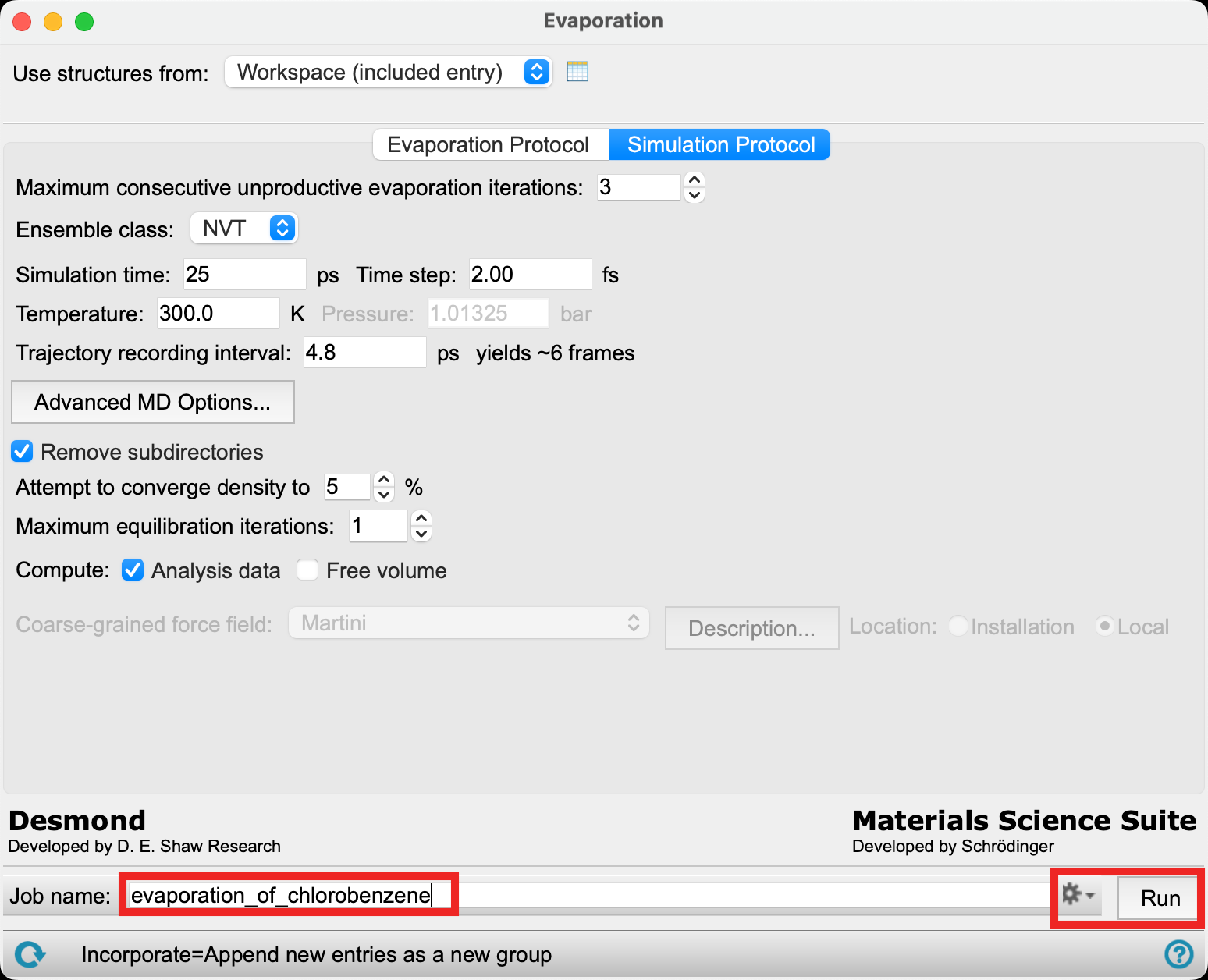
- Change the Job name to evaporation_of_chlorobenzene
- Due to the long calculation time for this job, we will proceed with the provided files. Go to File > Import Structures, navigate to the provided tutorial files and Open
Section_04 > evaporation_of_chlorobenzene > evaporation_of_chlorobenzene-out.cms.gz- This job requires a GPU host and can be completed in 1 day
- Close the Evaporation panel
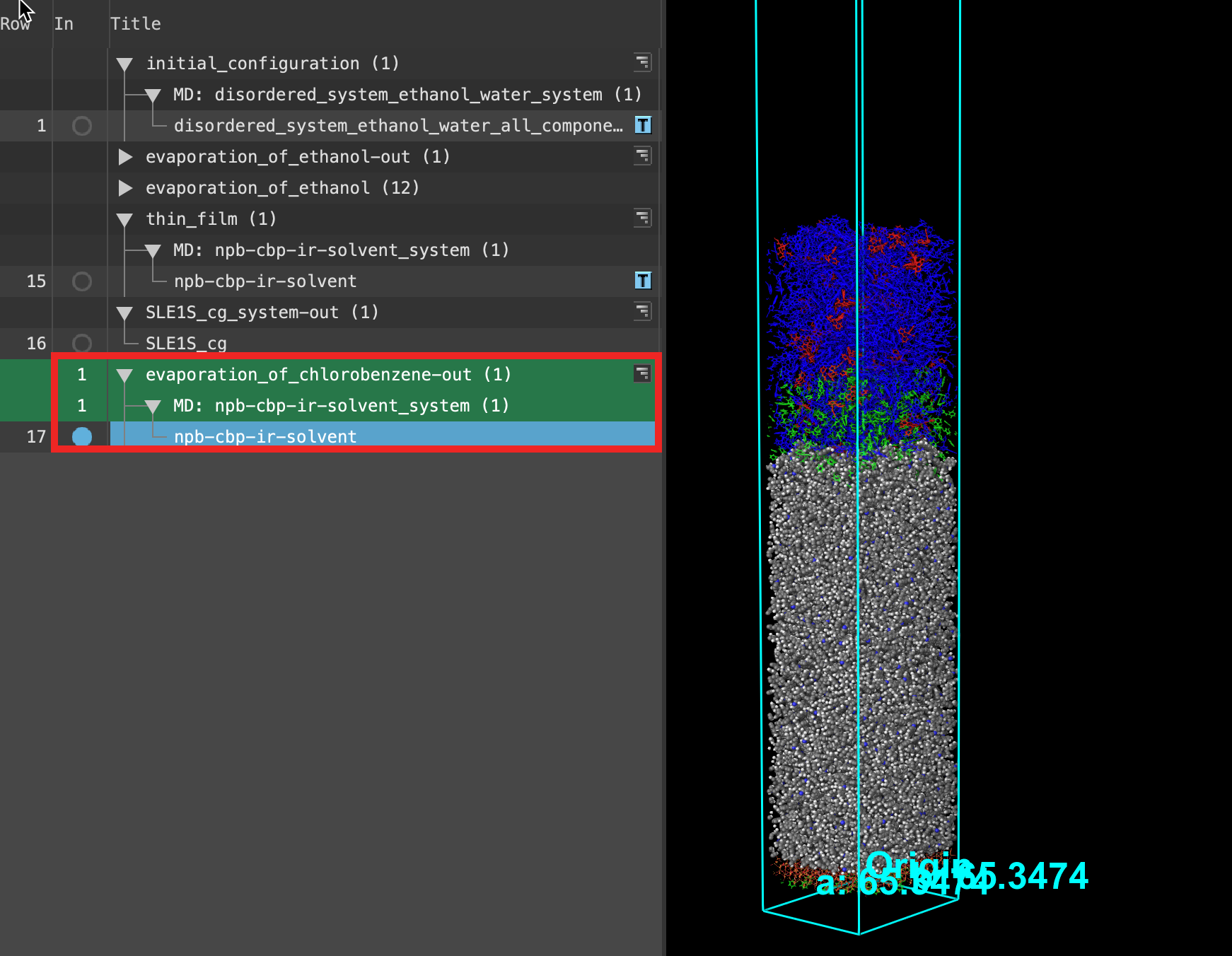
- When the job is finished or imported select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue the output npb-cbp-ir-solvent in the entry list.
- The displayed cell is from the last evaporation step. As shown in the Figure, the cell size is the same as the original input. This is because we used an NVT simulation, meaning the cell volume remained constant.
- Feel free to stylize and visualize the output system in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
There are some chlorobenzene molecules (green molecules) left near the NPB layer. The inability to remove all of the solvent from a system has been seen as a cause of efficiency loss and increased degradation.
Now we will open the Evaporation Viewer panel to extract some quantitative metrics from our simulation. Despite the fact that only the last iteration is shown in the workspace, this panel provides analysis over the entire course of the calculation.
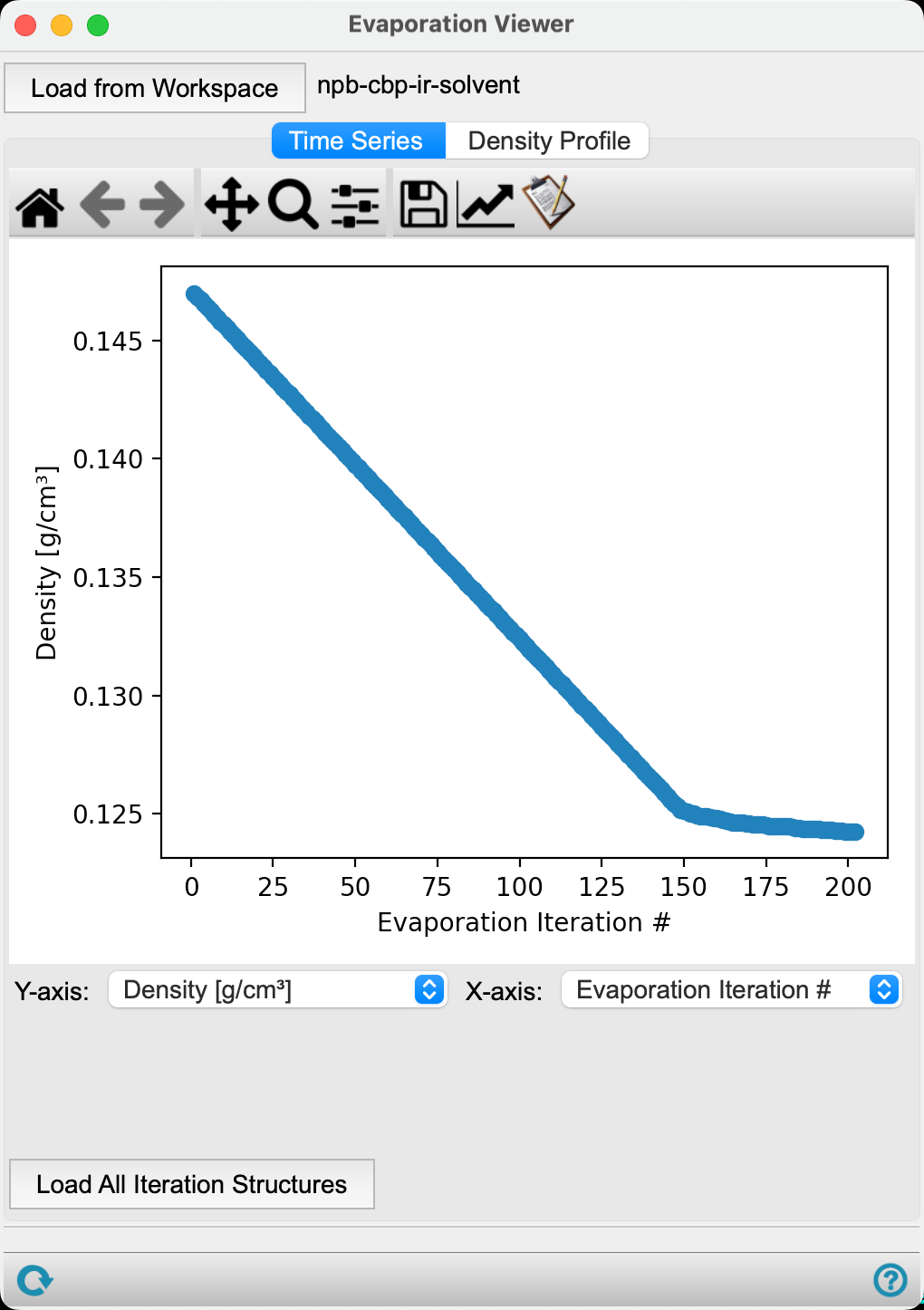
In the Time Series tab, the first plot provided is the Density (g/cm3) versus Evaporation iteration # for the calculation.
The density of the system decreases as the calculation proceeds. It is worth noting that this is not the most insightful metric, as molecules were being removed while the cell volume was being held constant, which necessitates a decrease in density.
We can also analyze how the mass percent of a particular component changes with evaporation iteration. To do so,
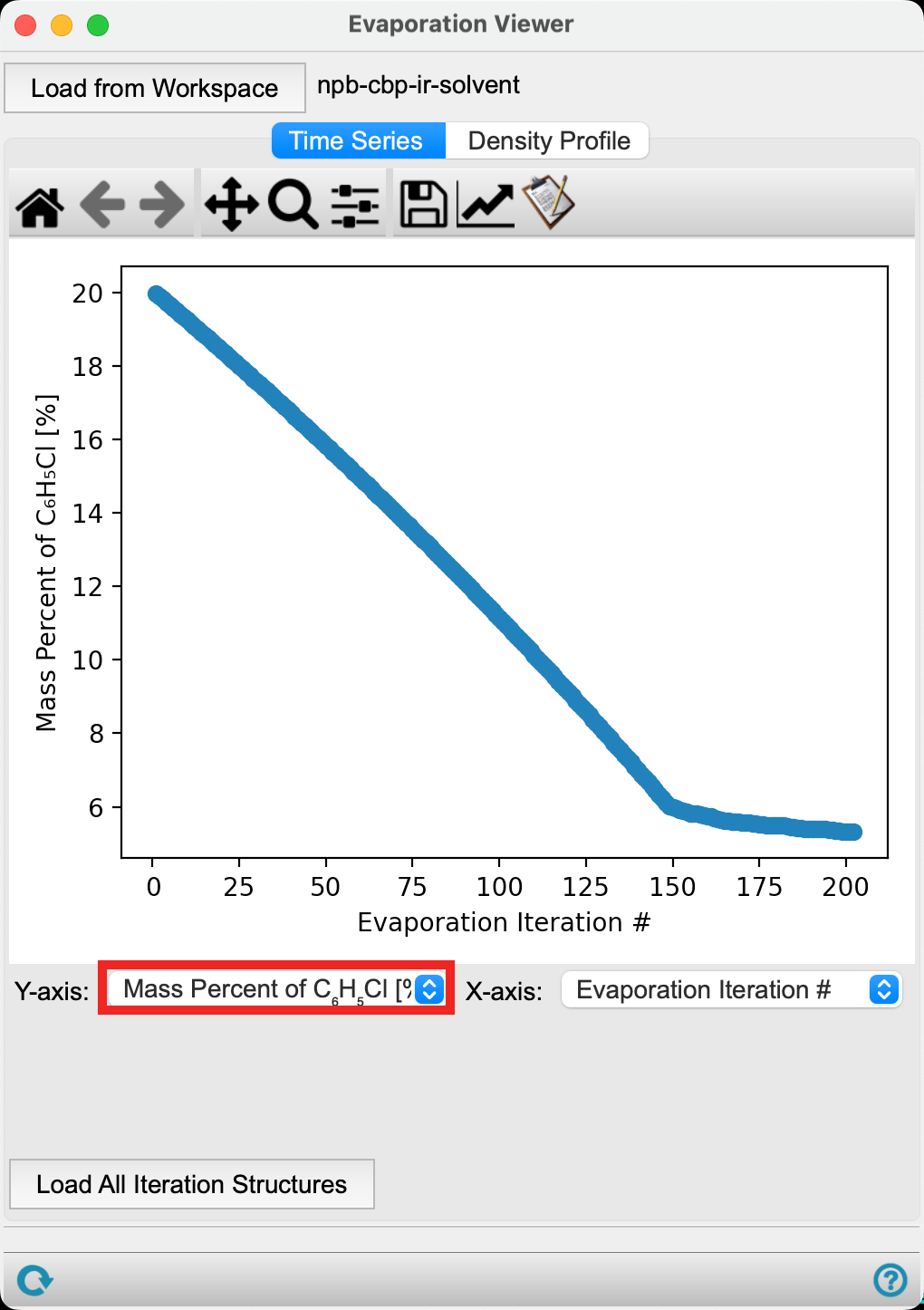
- Change the Y-axis to Mass Percent of C6H5Cl [%]
The plot updates to show the mass percent of chlorobenzene decreasing as a function of evaporation iteration. We see that fewer chlorobenzene molecules are removed after iteration 150. This is due to the solvent molecules being trapped in the organic layer.
- Close the Evaporation Viewer panel
5. Evaporation from a Coarse-Grained Multi-Component Box
Sodium lauryl ether sulfate (SLES) is an inexpensive surfactant, omnipresent in consumer packaged goods. In this section, we will use the Evaporation panel to remove water particles from our coarse-grained SLES box. This will require removing multiple solvents as there are two types of water particles in this coarse-grained system. The input system has already been equilibrated and prepared for further MD by applying a Martini coarse-grained force field.
- Go to File > Import Structures
- Navigate to where you downloaded the tutorial files (presumably your working directory) and choose
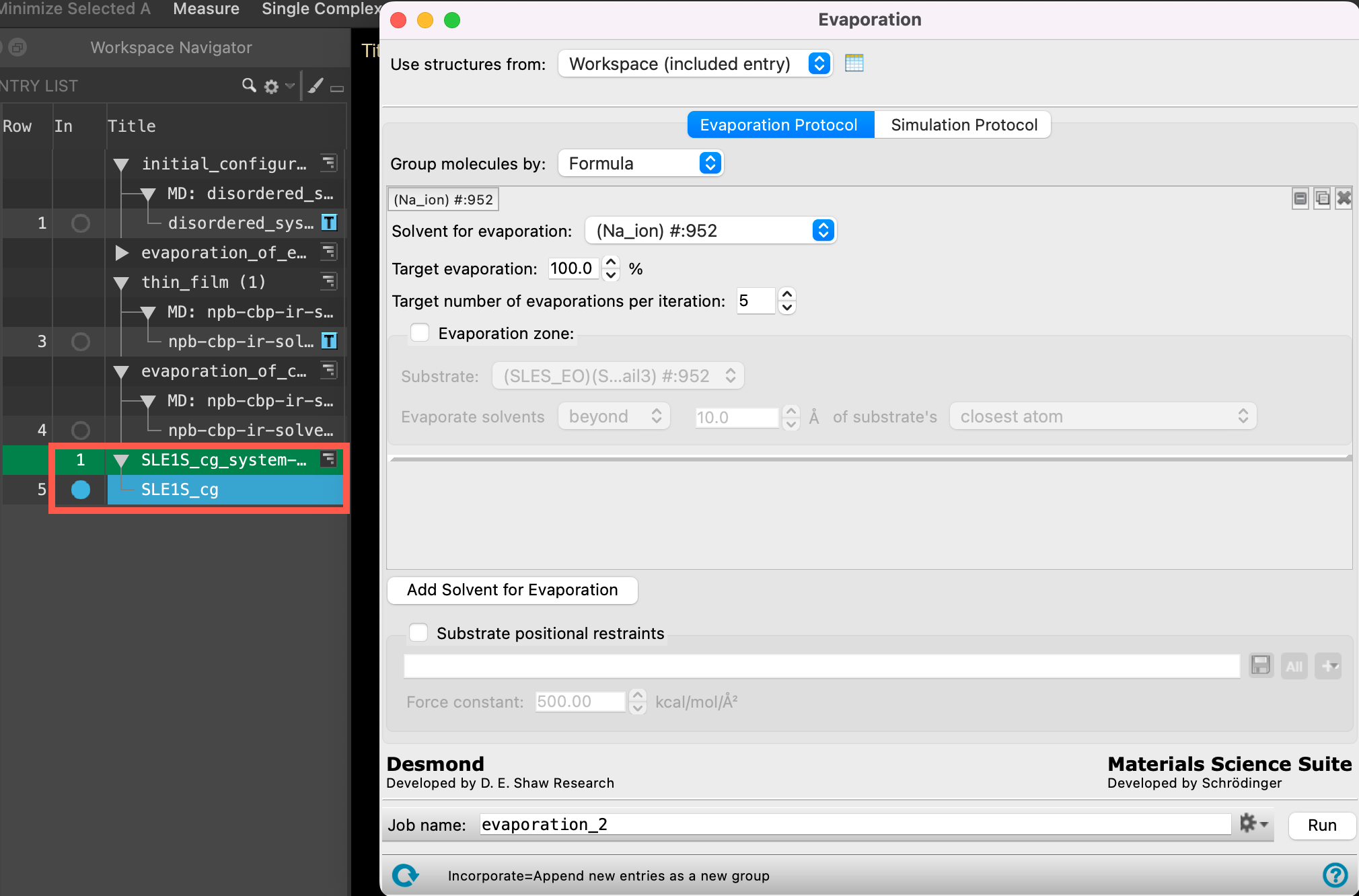
Section_05 > md_prep_SLE1S_cg > SLE1S_cg_system-out.cms. Click Open- A new entry group is added to the entry list containing an entry titled SLE1S_cg
- Ensure that SLE1S_cg is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Classical Mechanics > Evaporation > Evaporation Calculations
- The Evaporation panel opens
Before moving forward, we need to ensure the force field file for our SLE1S_cg structure is available to be used during the evaporation calculations. For this, we must move the force field file into the correct Schrödinger directory.
On Mac OS / Linux:
Move the provided force field file from its current location Section_05 > SLE1S_cgff.json to .schrodinger > matsci_templates > coarse_grain_force_field_parameters using the file explorer or the command line. Note that .schrodinger is a hidden folder.
On Windows:
Move the provided force field file from its current location Section_05 > SLE1S_cgff.json to AppData > Local > Schrodinger > matsci_templates > coarse_grain_force_field_parameters using the file explorer or the command line. Note that Schrodinger is a hidden folder.
There is no need to worry if you are unclear on how to proceed. If you need assistance performing this step, please email education@schrodinger.com. If you would like to continue the tutorial without performing this step, the results of the calculations are provided towards the end of this section.
Now, let’s specify the evaporation parameters for this example:
- Retain Formula for Group molecules by
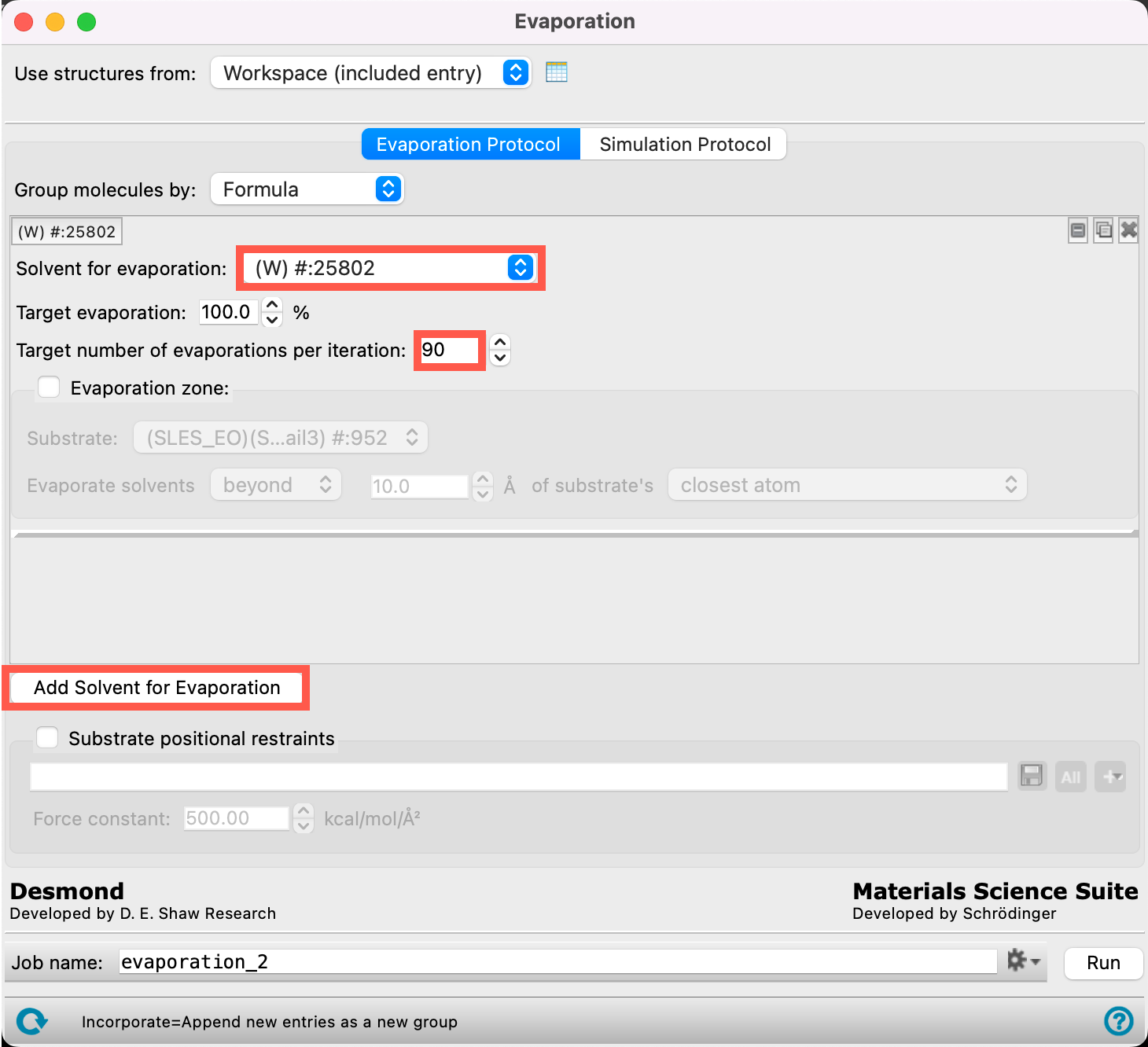
- From the Solvent for evaporation dropdown menu, select (W) #:25802
- The 25802 water particles of type W have now been identified as the target for the evaporation calculation
- Ensure the Target evaporation is 100%
- Change the Target number of evaporations per iteration to 90
- This means that we will remove at most 90 water particles per iteration until there are no water particles left.
- Click Add Solvent for Evaporation
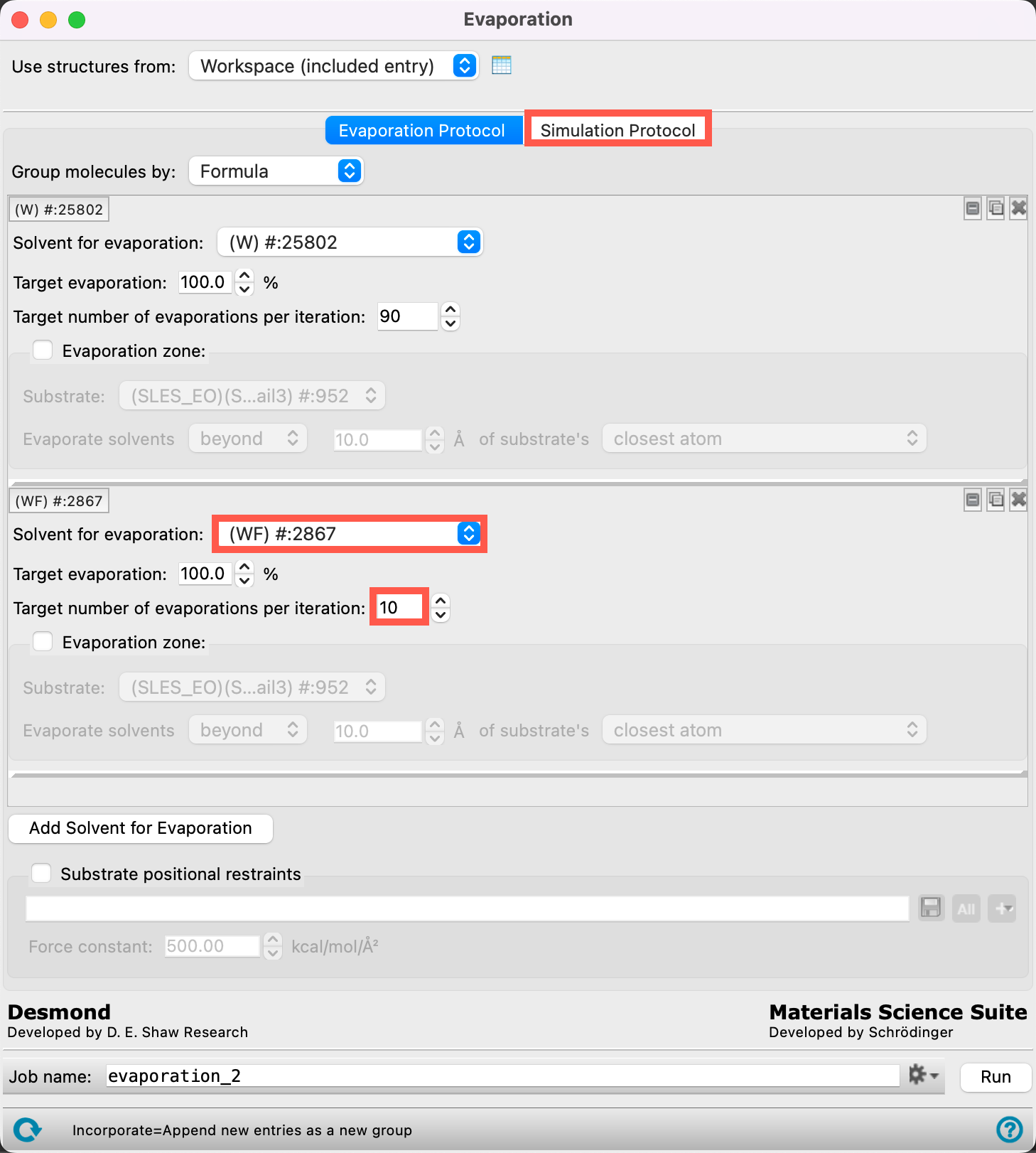
- We can now specify the evaporation parameters for WF
- From the Solvent for evaporation dropdown menu, select (WF) #:2867
- The 2867 water particles of type WF have now been identified as the target for the evaporation calculation
- Ensure the Target evaporation is 100%
- Change the Target number of evaporations per iteration to 10
- This means that we will remove at most 10 water particles per iteration until there are no water particles left.
Note: It is suggested to exercise caution when removing water from a coarse-grained system. Martini water is designed to contain a ratio of W:WF of 9:1. Removing one water type at a time can cause the system to freeze. Hence, we remove W and WF in a 9:1 ratio in this example.
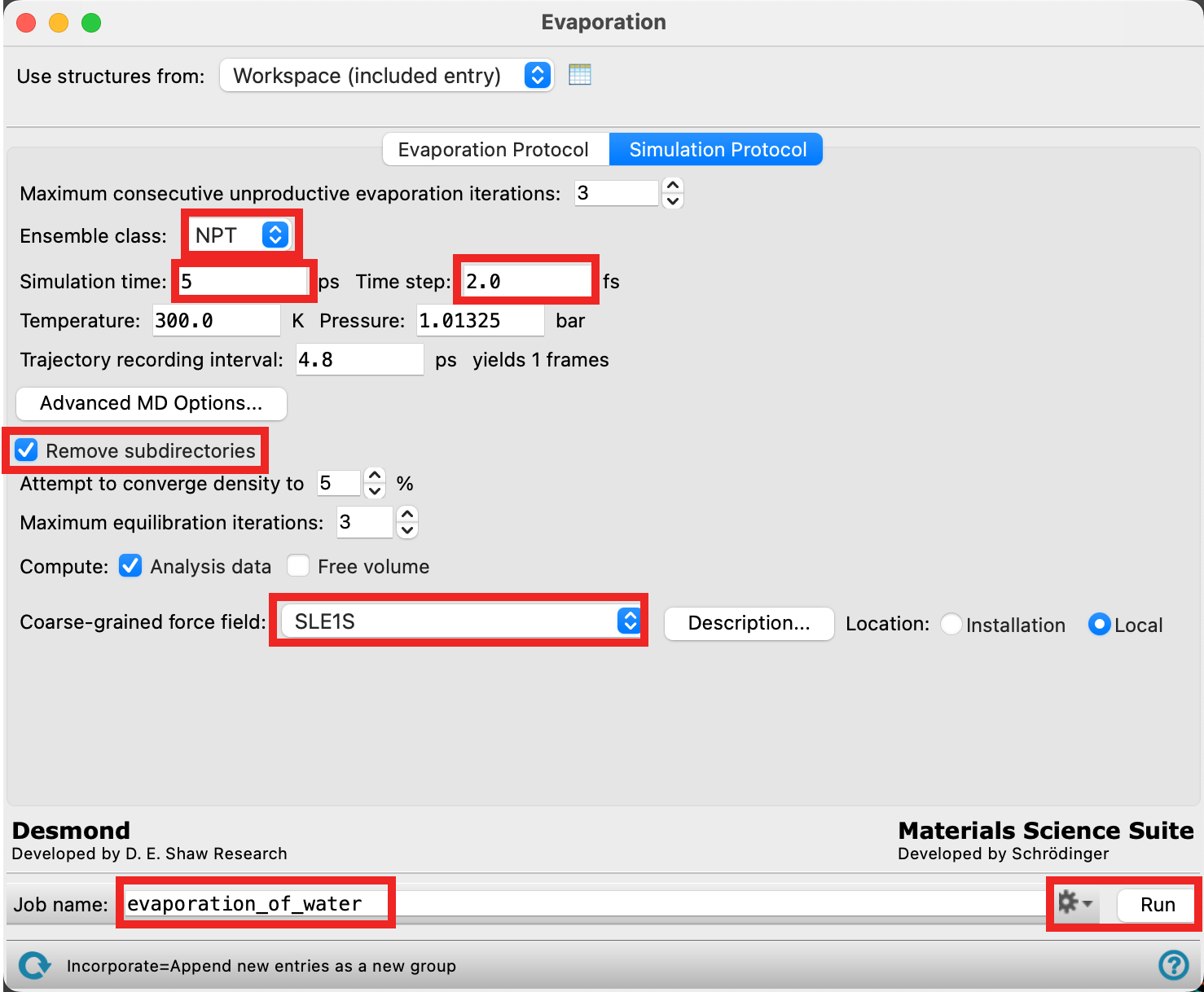
- Go to the Simulation Protocol tab
- Select NPT ensemble class.
- Enter 5 ps for the Simulation time
- For Time step enter 2 fs
- Check Remove subdirectories
- Ensure the Coarse-grained force field points to SLE1S
- If you do not see the force field file, it is likely that you did not place it in the correct location. Try to repeat the steps at the beginning of this section. If you still have trouble, please contact education@schrodinger.com or simply proceed with the provided files
- Change the Job name to evaporation_of_water
- Due to the long calculation time for this job, we will proceed with the provided files. Go to File > Import Structures, navigate to the provided tutorial files and Open
Section_05 > evaporation_of_water > evaporation_of_water-out.cms.gz- This job requires a GPU host and can be completed in 1 day
- Close the Evaporation panel
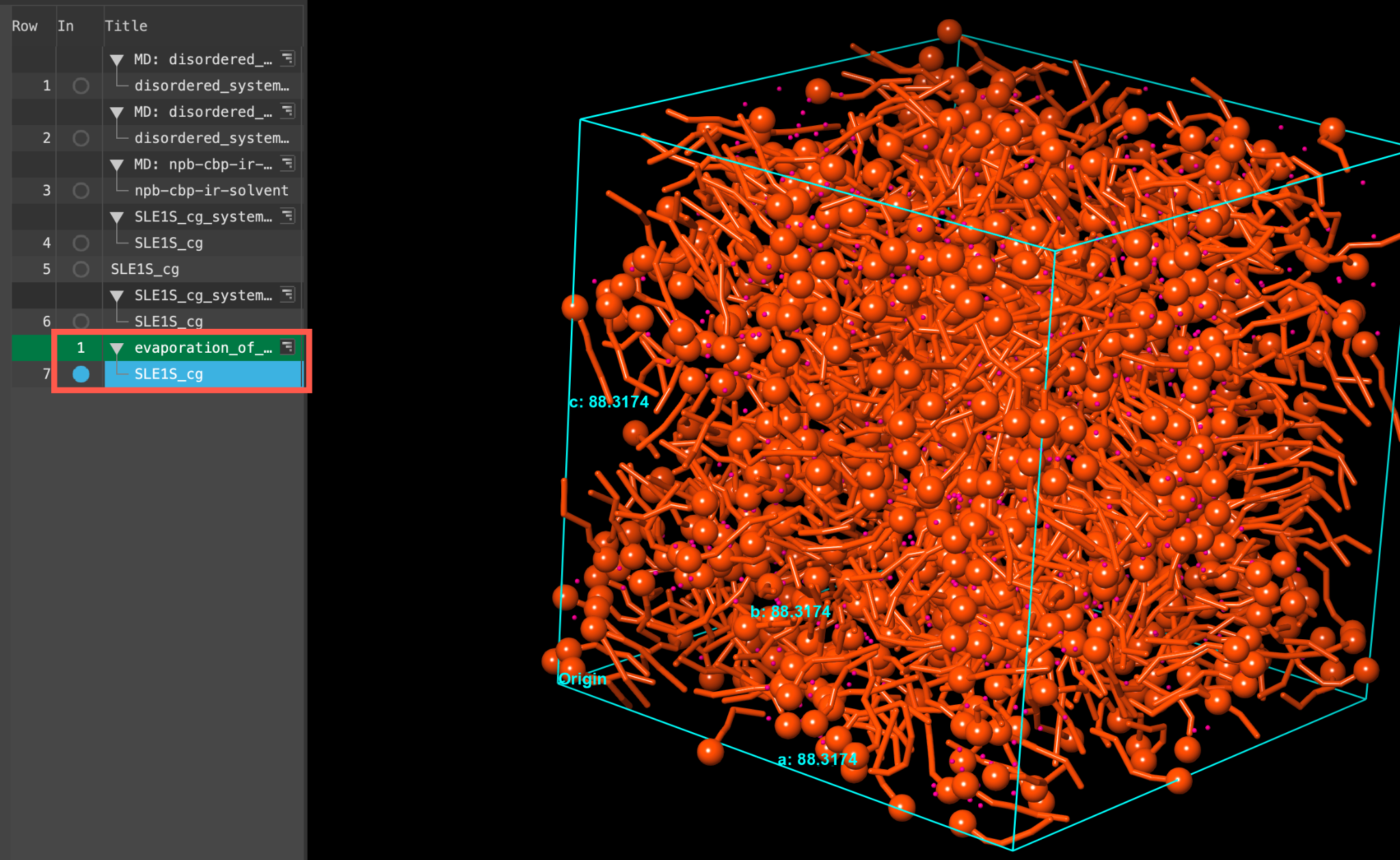
- When the job is finished or imported select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue the output SLE1S_cg in the entry list.
- The displayed cell is from the last evaporation step. As shown in the Figure, the cell size is significantly smaller than the original input as no water molecules remain
- Feel free to stylize and visualize the output system in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Now we will open the Evaporation Viewer panel to extract some quantitative metrics from our simulation.
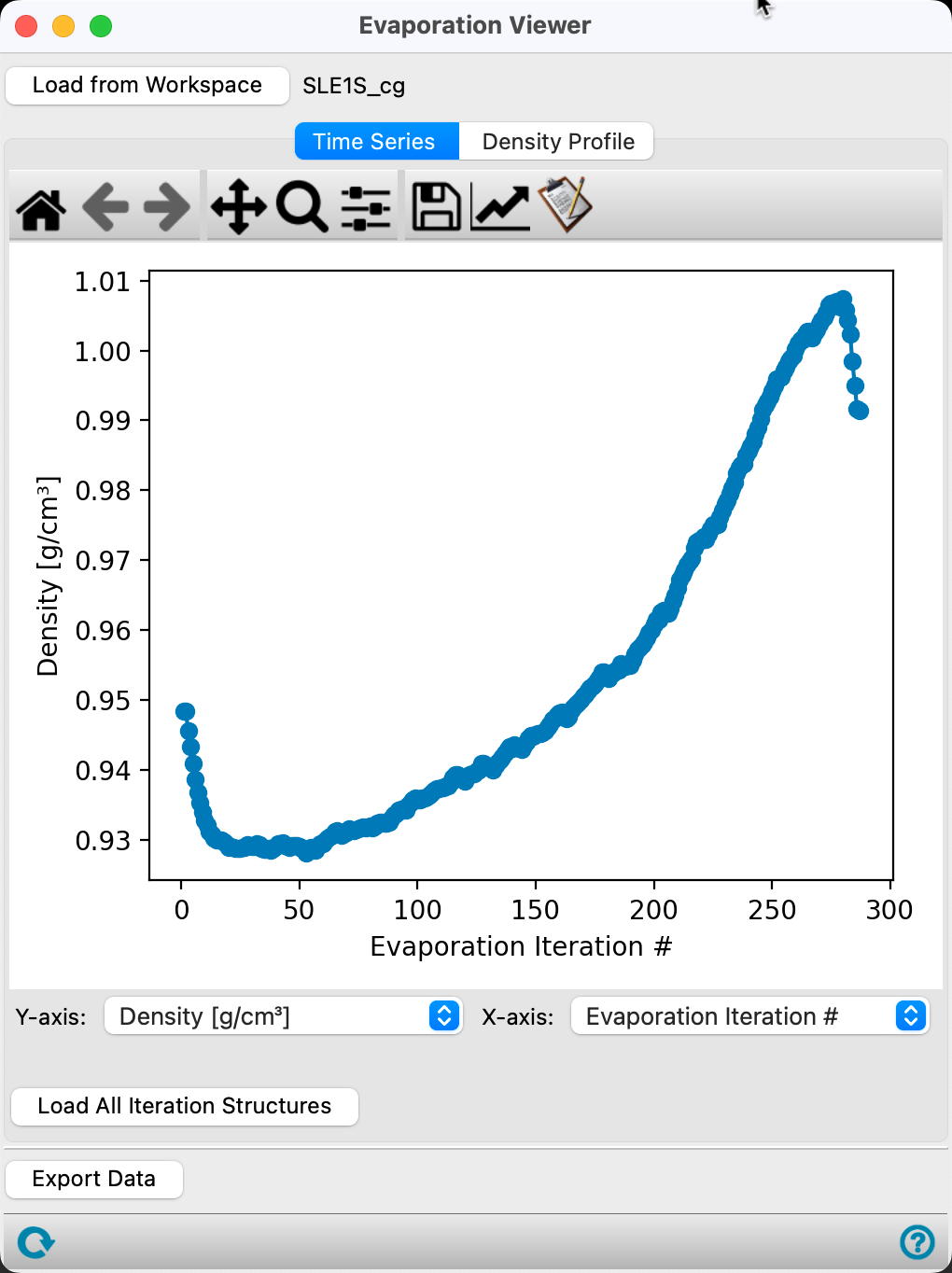
In the Time Series tab, the first plot provided is the Density (g/cm3) versus Evaporation iteration # for the calculation.
The density of the system increases significantly after the first 100 iterations. After most of the water is removed, the system reorganizes such that the density decreases.
We can also analyze how the mass percent of a particular component changes with evaporation iteration. To do so,
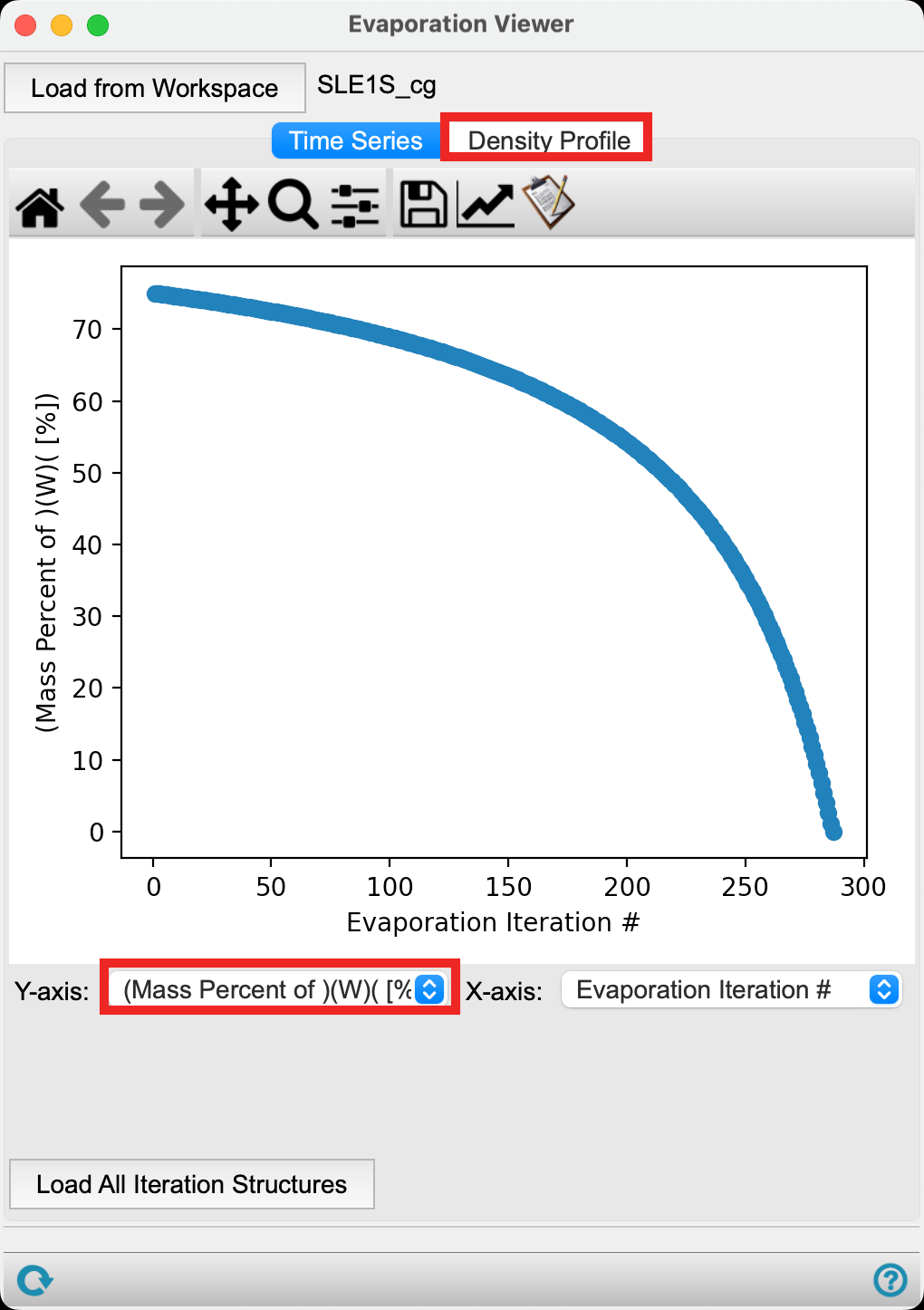
- Change the Y-axis to (Mass Percent of) (W) ([%])
The plot updates to show the mass percent of water of type W decreasing as a function of evaporation iteration. This is expected as the water particles are removed from the box.
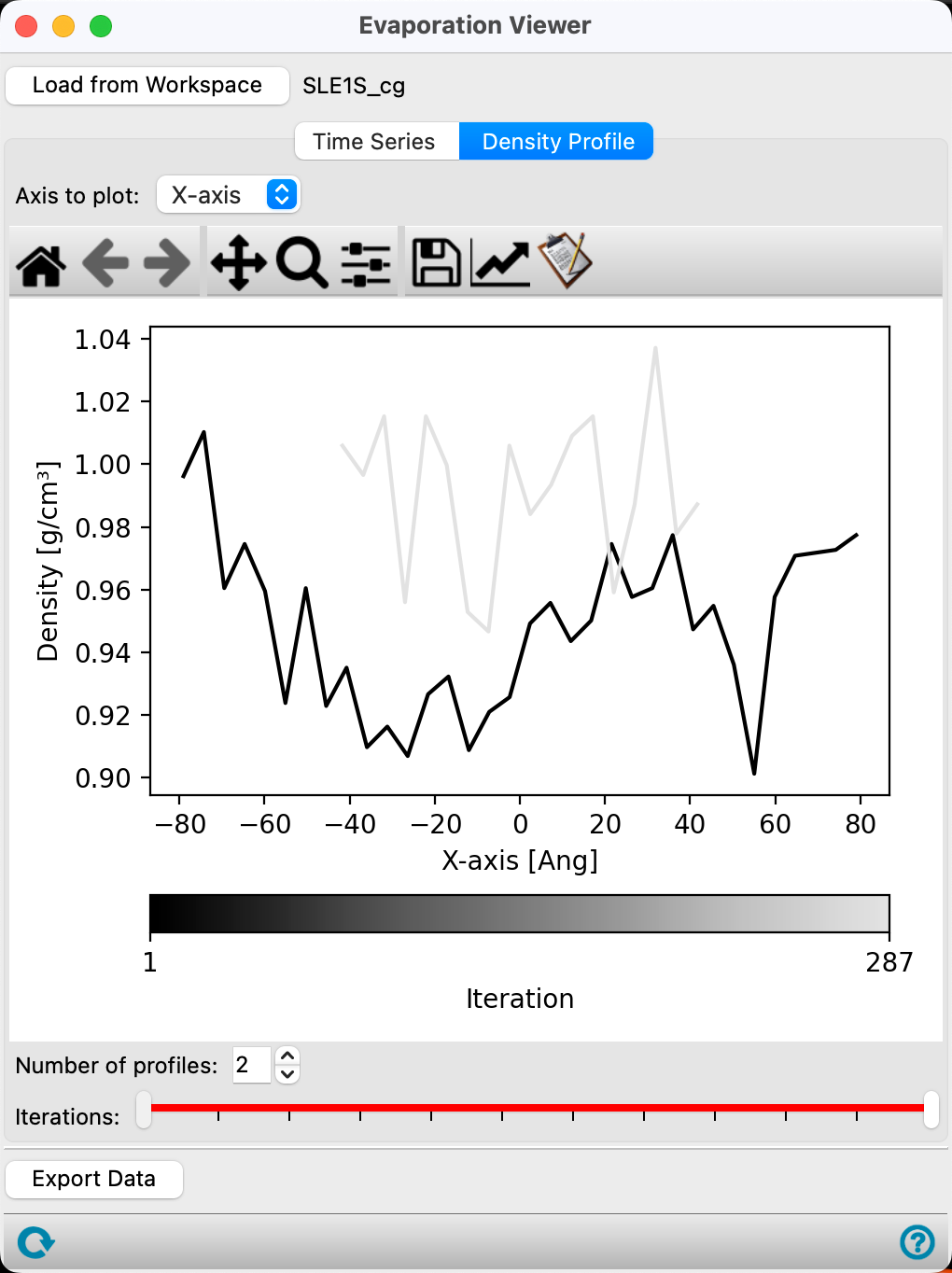
- Go to the Density Profile tab
This plot tells us the overall density of the cell has increased, as the gray curve is higher than the black. Additionally, the cell has shrunk significantly as the gray curve does not extend the entire x-axis as the black curve does. In this small simulation, there are distinct clusters that are also evident in the density profile. The reorganization of surfactants into larger macrostructures comes with an increased overall density.
Feel free to explore the Viewer panel further.
- Close the Evaporation Viewer panel
6. Conclusion and References
In this tutorial, we learned how to use the Evaporation calculation and viewer panels to remove compounds from an all-atom multi-component cell, a thin film on a substrate, and a coarse-grained multi-component cell.
For further learning:
For introductory content, focused on navigating the Schrödinger Materials Science interface, an Introduction to Materials Science Maestro tutorial is available. Please visit the materials science training website for access to 100+ tutorials. For scientific inquiries or technical troubleshooting, submit a ticket to our Technical Support Scientists at help@schrodinger.com.
For self-paced, asynchronous, online courses in Materials Science modeling, including access to Schrödinger software, please visit the Schrödinger Online Learning portal on our website.
For some related practice, proceed to explore other relevant tutorials:
- Disordered System Building and MD Multistage Workflows
- Building, Equilibrating and Analyzing Amorphous Polymers
- Building Solvated Systems
- Crosslinking Polymers
- Building a Semicrystalline Polymer
- Thermal Conductivity
- Polymer Property Prediction
- Cyclic Stress Strain
- Penetrant Loading
- Diffusion
- Molecular Dynamics Simulations for API Miscibility
- Cluster Analysis
- Electroporation
- Calculating Surfactant Tilt and Electrostatic Potential of a Bilayer System
- Building a Polymer-Polymer Interface
- Building a Carbohydrate Polymer
- Surface Tension
- Glass Transition Temperatures for APIs
- Viscosity
- Building a Coarse-Grained Surfactant Model with Martini Force Field
- Ibuprofen Cyclodextrin Inclusion Complexes with the Martini Coarse-Grained Force Field
For further reading:
- See the help documentation on the Evaporation Calculations and Viewer panels
- Structuring of Polymer Solutions upon Solvent Evaporation. DOI:10.1103/PhysRevE.91.022602
- How Do Evaporating Thin Films Evolve? Unravelling Phase-Separation Mechanisms during Solvent-Based Fabrication of Polymer Blends. DOI:10.1063/1.4898136
- Dissipative Particle Dynamics Study for the Phase Separated Structures of Polymer Thin Film Caused by Solvent Evaporation. DOI:10.1678/rheology.36.93
- Variety of Ordered Patterns in Donor–Acceptor Polymer Semiconductor Films Crystallized from Solution. ACS Appl. Mater. DOI:10.1021/acsami.1c00079
- From Molecular Packing Structures to Electronic Processes: Theoretical Simulations for Organic Solar Cells. DOI:10.1002/aenm.201702743
7. Glossary of Terms
Entry List - a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
Included - the entry is represented in the Workspace, the circle in the In column is blue
Project Table - displays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data
Recent actions - This is a list of your recent actions, which you can use to reopen a panel, displayed below the Browse row. (Right-click to delete.)
Scratch Project - a temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project
Selected - (1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Working Directory - the location where files are saved
Workspace - the 3D display area in the center of the main window, where molecular structures are displayed