Diffusion
Tutorial Created with Software Release: 2026-1
Topics: Consumer Packaged Goods , Energy Capture & Storage , Pharmaceutical Formulations , Polymeric Materials
Methodology: All-Atom Molecular Dynamics
Products Used: Desmond , MS Maestro , MS Transport
|
0.5 GB |
This tutorial is written for use with a 3-button mouse with a scroll wheel.
Words found in the Glossary of Terms are shown like this: Workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Abstract:
In this tutorial, we will learn to build and equilibrate a prototypical system for a Diffusion Coefficient calculation. We will then use the Diffusion Coefficient Calculation and Viewer panels to determine the diffusion coefficient for the Li ions in the model system.
Tutorial Content
1. Introduction to Diffusion
The diffusion coefficient, also known as diffusivity, is formally the proportionality constant between the molar flux due to molecular diffusion and the gradient in the concentration of the species. The diffusion coefficient has units of area per time (typically m2/s). For molecules in the gas phase, typical diffusion coefficients are on the order of 10-6-10-5 m2/s, whereas typical molecules in liquid solution diffuse ~5 orders of magnitude slower.
There are two primary ways to measure diffusion constants from MD simulation: via Mean squared displacement and via Velocity autocorrelation functions. The mean squared displacement method measures the average displacement of particles versus time. For Fickian diffusion, we expect a plot of mean squared displacement versus time to be linear and its slope to be proportional to the diffusion constant. The velocity autocorrelation method uses a Green-Kubo relationship which relates diffusion constant to the integral of the velocity autocorrelation function. In this tutorial, you will calculate the diffusion coefficient via the Mean square displacement method.
Predicting diffusivity is useful in many application areas. For example, in pharmaceutical formulations, we may be interested in diffusion coefficients of APIs (active pharmaceutical ingredients) in various media, or in battery design, we may be interested in ion diffusion coefficients.
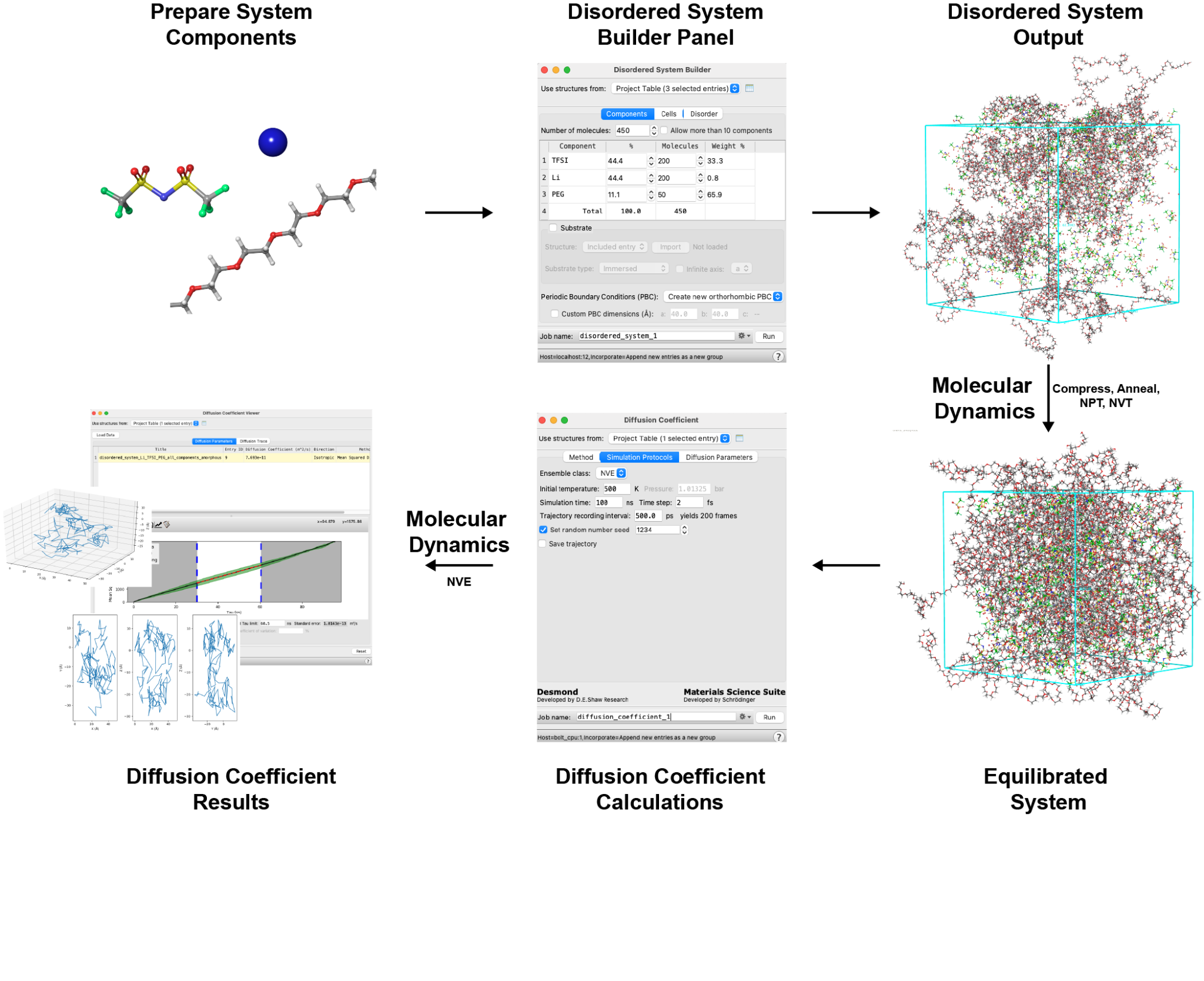
In this tutorial, we explore a prototypical system of interest with respect to battery design. Specifically, we will build a system containing lithium cations in a polyethylene glycol (PEG) electrolyte mixed with bis(trifluoromethylsulfonyl)imide (TFSI) anions. We will predict the diffusion coefficient of the Li ions at 500 K employing a workflow in Materials Science (MS) Maestro using classical molecular dynamics.
To execute this workflow, we will first use the Disordered System Builder panel to construct the system. Then, we will utilize the MD Multistage Workflow panel to execute a best practice molecular dynamics (MD) equilibration in preparation for a diffusion calculation (a compressive protocol, simulated annealing to heat the system to 500 K, an NPT simulation and finally an NVT simulation). Next, we will use the Diffusion Coefficient calculation panel to run a production NVE simulation and collect diffusion data for the Li ions. Finally, we will employ the Diffusion Coefficient Results panel to analyze the results, including extracting the diffusion coefficient and plotting traces of ion diffusion.
The overall workflow can be summarized as follows:
The system studied herein and the best practices described are based on the publications listed in the References section.
2. Creating Projects and Importing Structures
At the start of the session, change the file path to your chosen Working Directorythe location where files are saved in MS Maestro to make file navigation easier. Each session in MS Maestro begins with a default Scratch Projecta temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project, which is not saved. A MS Maestro project stores all your data and has a .prj extension. A project may contain numerous entries corresponding to imported structures, as well as the output of modeling-related tasks. Once a project is saved, the project is automatically saved each time a change is made.
Structures can be built in MS Maestro or can be imported using File > Import Structures (or drag-and-dropped), and are added to the Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data. The Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion is located to the left of the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed. The Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data can be accessed by Ctrl+T (Cmd+T) or Window > Project Table if you would like to see an expanded view of your project data.
- Double-click the Materials Science icon
- (No icon? See Starting Maestro)
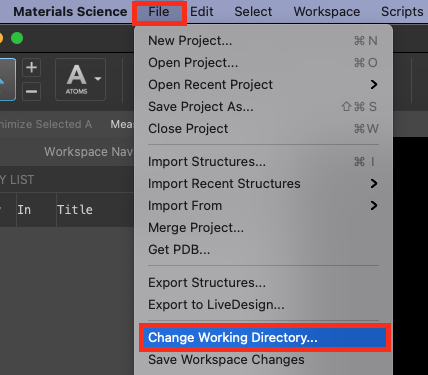
- Go to File > Change Working Directory
- Find your directory, and click Choose
- Pre-generated files are included for running jobs or examining output. Download the zip file here: schrodinger.com/sites/default/files/s3/release/current/Tutorials/zip/diffusion.zip
- After downloading the zip file, unzip the contents in your Working Directorythe location where files are saved for ease of access throughout the tutorial
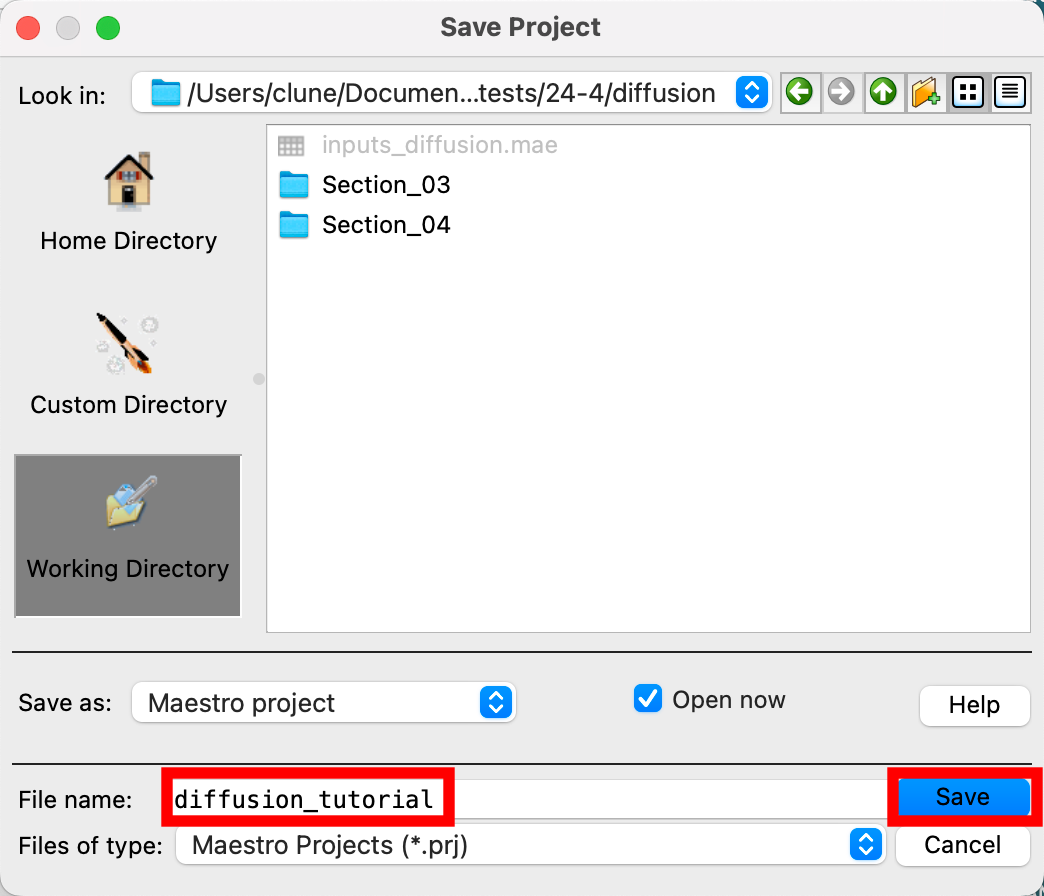
- Go to File > Save Project As
- Change the File name to diffusion_tutorial, click Save
- The project is now named
diffusion_tutorial.prj
- The project is now named
We will construct a typical system containing three components: lithium cations, TFSI anions and amorphous polyethylene glycol (PEG). These components are provided:
- Go to File > Import Structures
- Navigate to where you downloaded the provided tutorial files, choose
inputs_diffusion.maeand click Open- A new entry group is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion containing three entries
If you would prefer to practice preparing the components yourself, draw the ions in the 2D Sketcher and be sure to include charge. Generate the PEG 50-mer using the polymer builder. For background on using these tools, see the Introduction to Maestro for Materials Science and Building, Equilibrating and Analyzing Amorphous Polymers tutorials.
Note: The PEG 50-mer was prepared with the polymer builder and is linear. Because we will use the Tangled chain setting in Section 3, it is no problem to build the starting model in this way. If you are using Snapped to grid or Amorphous states, you should generate a more reasonable starting model.
3. Preparing a System for Diffusion Coefficient Calculations
Systems for diffusion calculations should be well-equilibrated. In this section, we will learn to construct the system of interest using the Disordered System Builder panel, and then we will use the MD Multistage Workflow panel to employ a best practice molecular dynamics (MD) protocol for preparing a system for diffusion calculations. Before building the system, we will assign custom ESP (electrostatic potential) charges to the ionic components of the system.
While we can proceed directly to construct a disordered system, it is worth noting that the current atomic charges (+1 for the lithium ion and -1 for the TFSI ion) may have a disproportionate effect on the system’s dynamics. To account for this, it is often practical to first scale the atomic charges on the ions to dampen their impact on the simulation and better match experiment.
To determine the quantities to assign as the partial charges, it is typical to perform a quantum mechanical calculation to calculate ESP charges for a small aperiodic system composed of one or more ion pairs in a relevant environment.
To learn how to perform an atomic charge calculation, visit the Computing Atomic Charges or the Introduction to Multistage Quantum Mechanical Workflows tutorial.
In this tutorial, we will provide the scaled atomic charges and learn how to designate them before performing any molecular dynamics simulations.
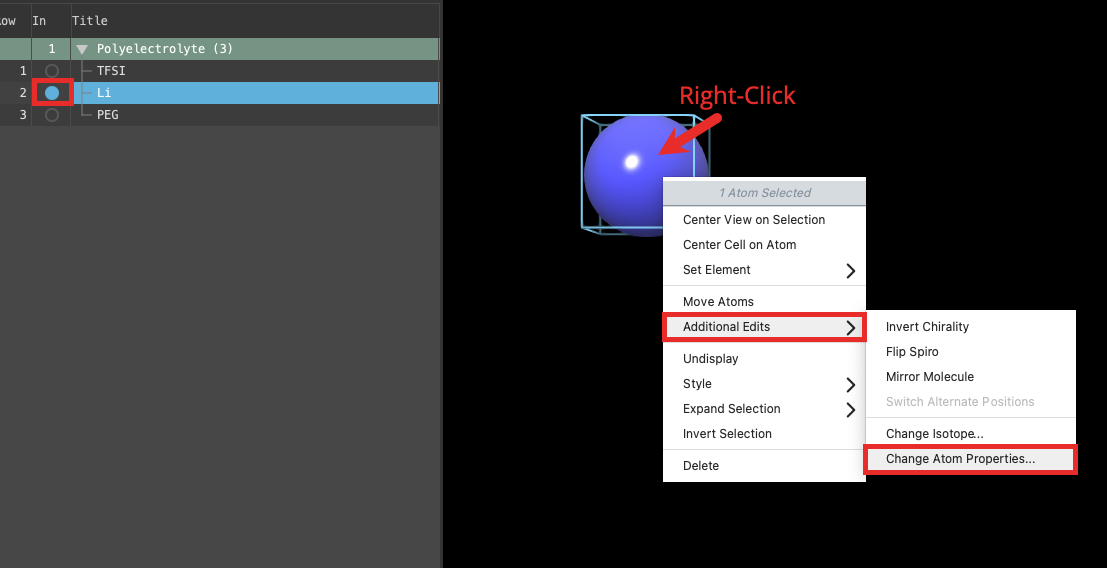
- Includethe entry is represented in the Workspace, the circle in the In column is blue the lithium ion in the workspace
- Right-click on the atom and choose Additional Edits > Change Atom Properties
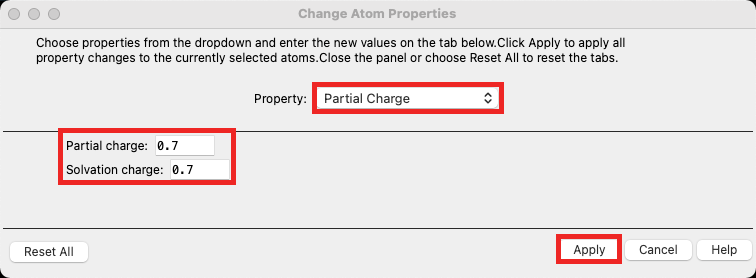
- The Change Atom Properties window opens
- Choose Partial Charge from the dropdown
- Set the Partial charge to 0.7 and the Solvation charge to 0.7
- Click Apply
- Click Close
We can quickly confirm that the charge is set by viewing the labels in the workspace.
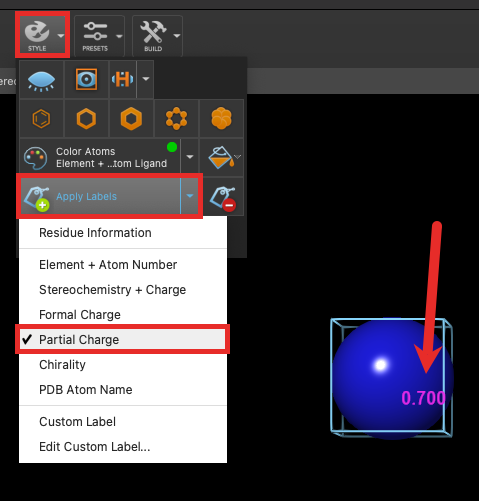
- From the Style palette, click the dropdown next to Apply Labels and choose Partial Charge
- The ion is labeled with 0.700 in the workspace
Now we need to assign the atomic charges in TFSI.
- Includethe entry is represented in the Workspace, the circle in the In column is blue the TFSI entry in the workspace
-
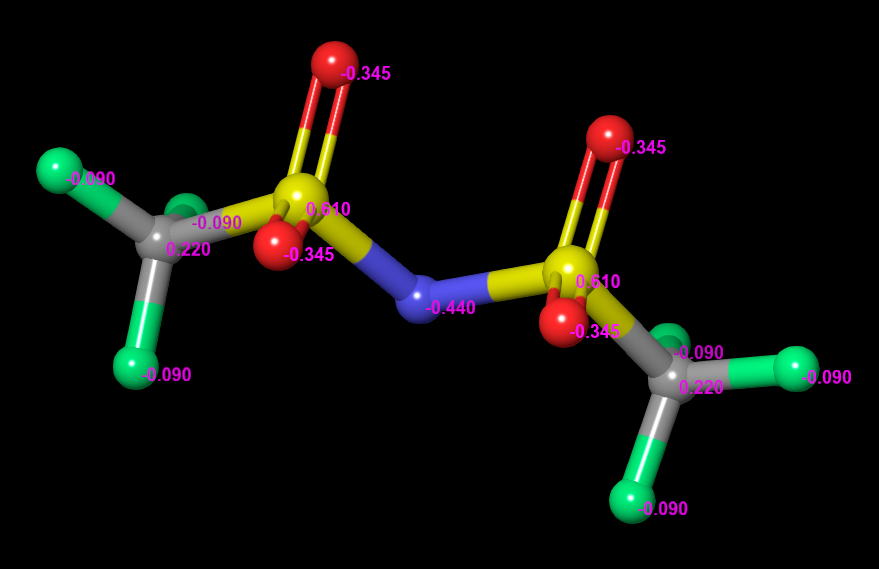
Proceed to repeat the above steps for the 11 atoms in TFSI. Note that you can select, for example, all six fluorine atoms at the same time, to set their charge simultaneously. Set the charges as follows:
- N: -0.44
- S: 0.61
- O: -0.345
- F: -0.09
- C: 0.22
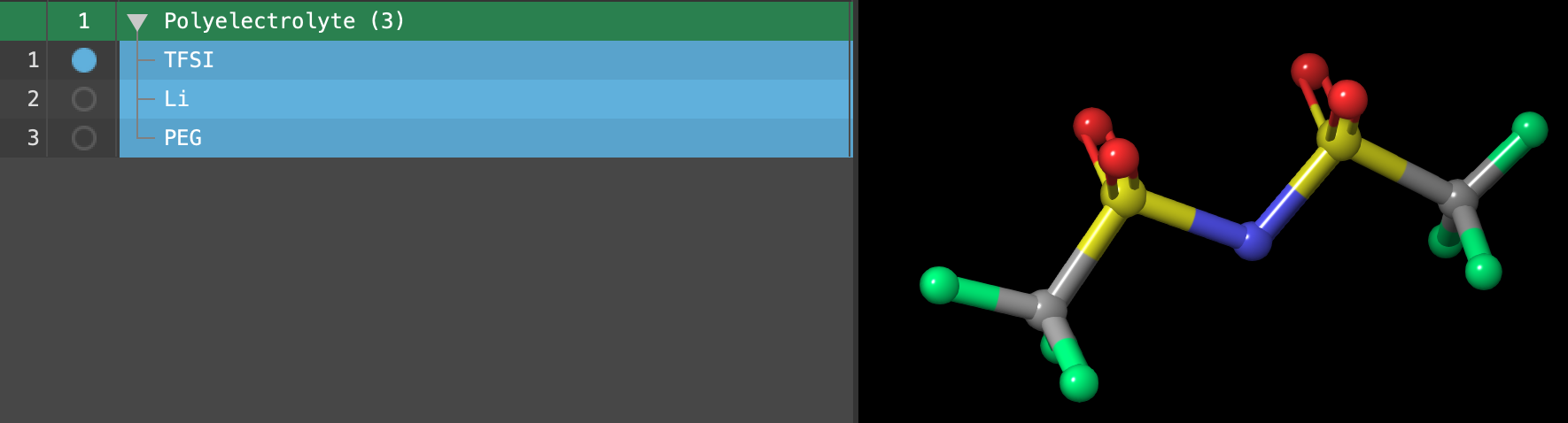
- After updating the charges, view the labels in the workspace to confirm that your entry matches the Figure.
We have now scaled the ion charges to +/- 0.7.
These quantities are based on a quantum mechanical calculation on a Li / TFSI cluster with a short PEG polymer chain (see References). There are many strategies for determining these charges, and exploring the effect of different charges on the subsequent simulations may be necessary in a study. See the Computing Atomic Charges tutorial for more details on how to obtain these charge values.
Note: An MD simulation is recommended to be performed on a neutral system. So, it is important to be sure that your overall system will be neutral once fully constructed.
Now we can proceed to build our disordered system:
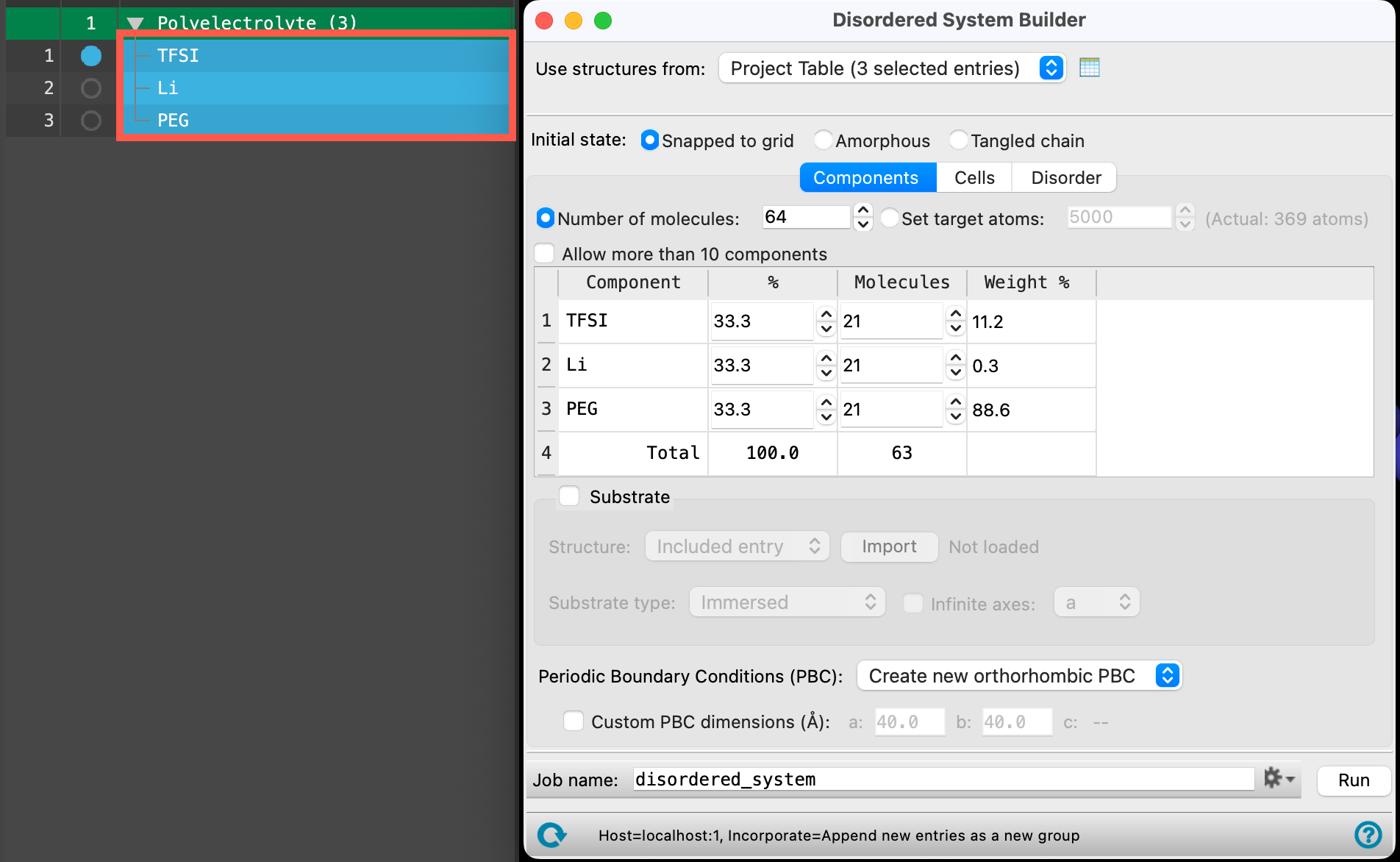
- Select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries the entire Polyelectrolyte (3) entry group
- Recall that select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries means to highlight the entries in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Structure Builders > Disordered System
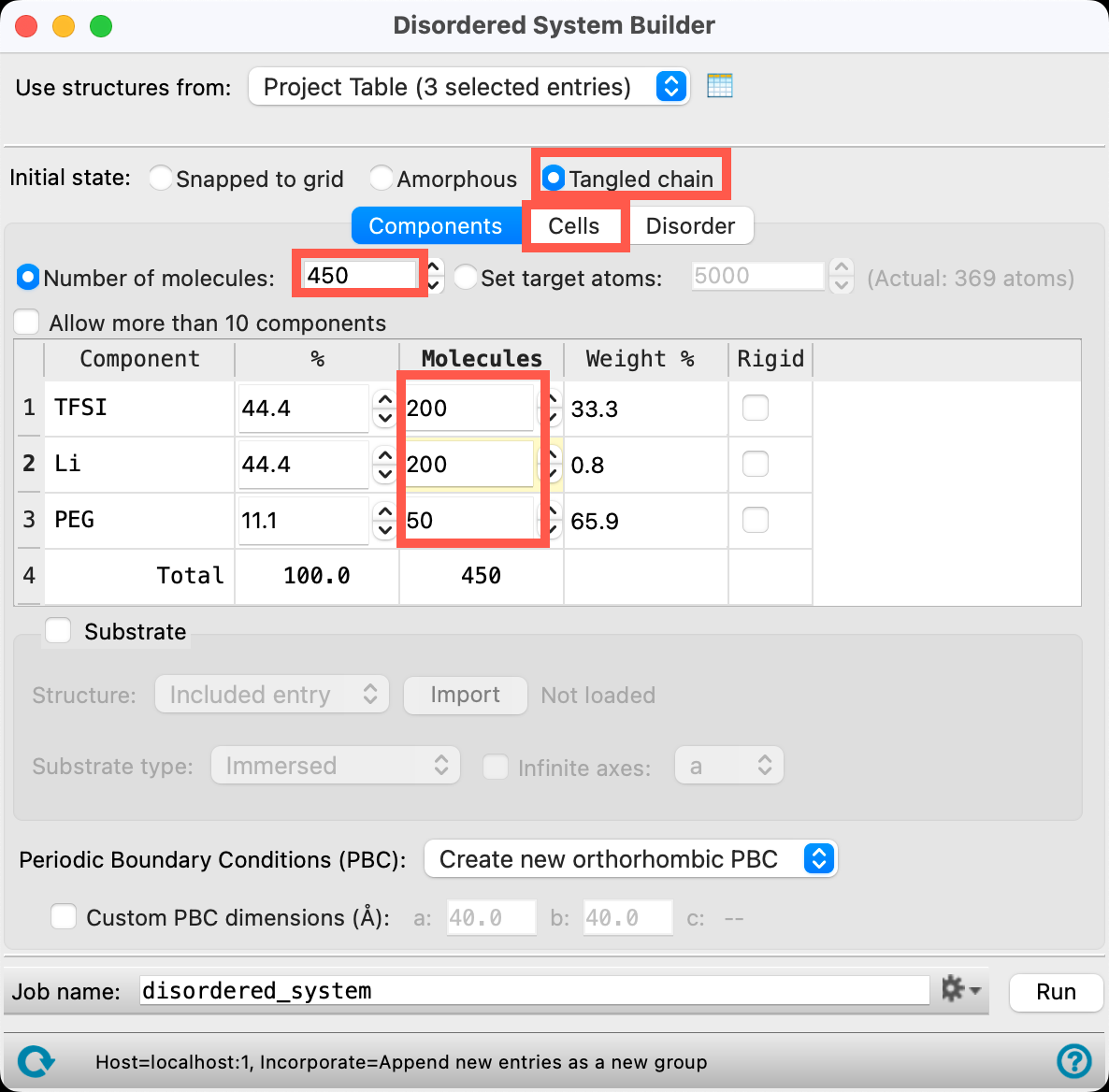
- The Disordered System Builder panel opens
- Change the Initial state to Tangled chain
- Change the Number of molecules to 450
- Change the number of TFSI and Li molecules to 200
- The number of PEG molecules should update automatically to 50
- Go to the Cells tab
Because we are using custom charges, we need to update the force field:
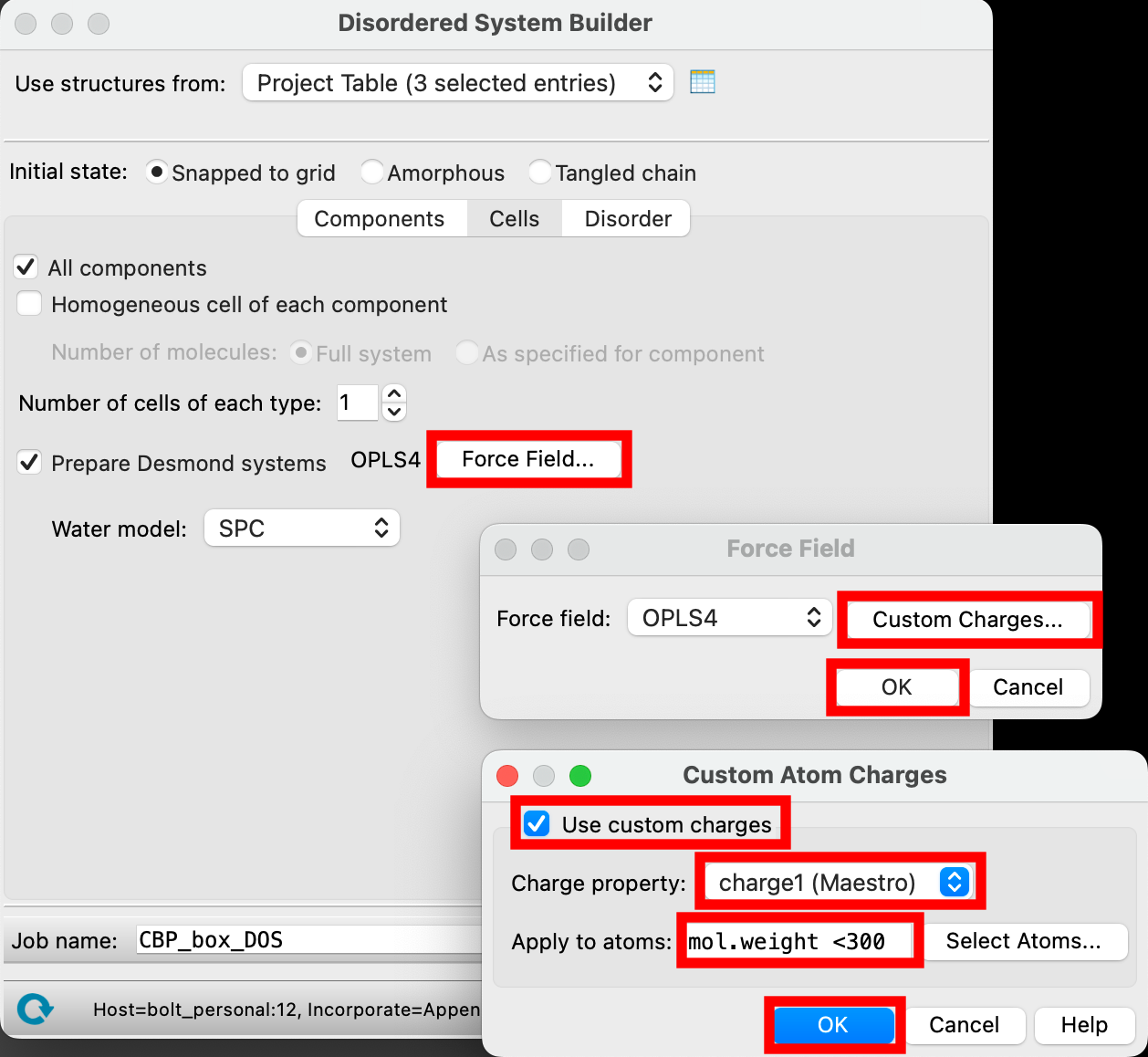
- Click Force Field
- Click Custom Charges
- For Charge property choose charge1 (Maestro)
- These refer to the charges we just assigned
- For Apply to atoms, input
mol.weight <300- This will indicate that the custom charges should only be used for molecules under 300 amu, basically excluding the polymer chains but including the ions
- This input utilizes the Atom Specification Language (ASL), and there are many ways to specific components
- Click OK to close the Custom Atom Charges Window and OK again to close the Force Field window
An option for a machine learning force field (MLFF) is available as an alternative. See the Disordered System Builder panel documentation to learn more.
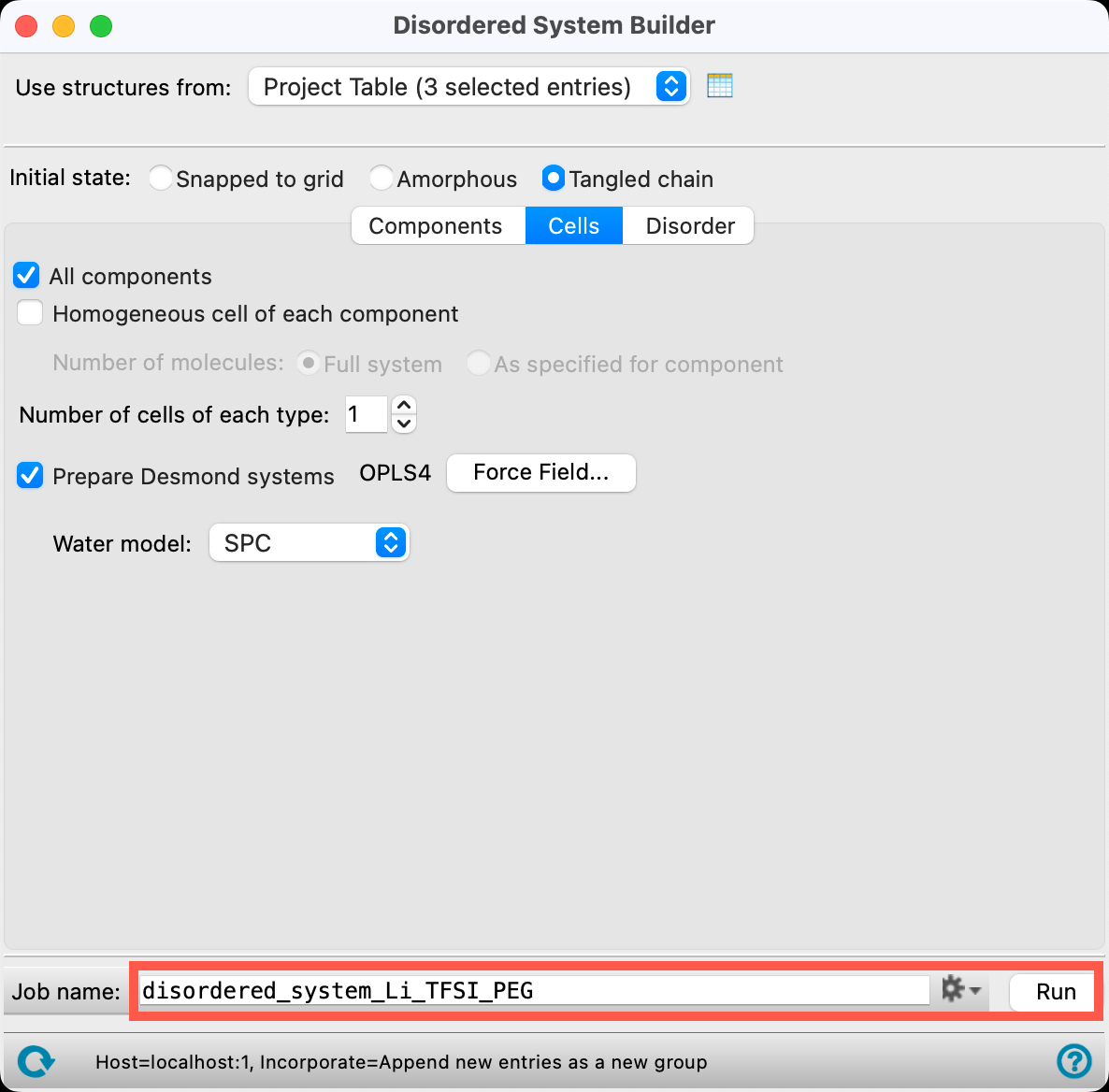
- Change the Job name to disordered_system_Li_TFSI_PEG
- Adjust the job settings (
 ) as needed
) as needed
- This job requires a CPU host. The job can be completed in about 5 minutes
- If you would prefer not to run the job, import
Section_03 > disordered_system_Li_TFSI_PEG > disordered_system_Li_TFSI_PEG_system-out.cmsfrom the provided tutorial files via File > Import Structures. Otherwise, click Run - Close the Disordered System Builder panel
Note: If the labels are still turned on and are cluttering the workspace, turn them off with the  button in the bottom right corner of the workspace.
button in the bottom right corner of the workspace.
Note: For general practice with the Disordered System Builder and more information about the various parameters, see the Disordered System Building and MD Multistage Workflows tutorial
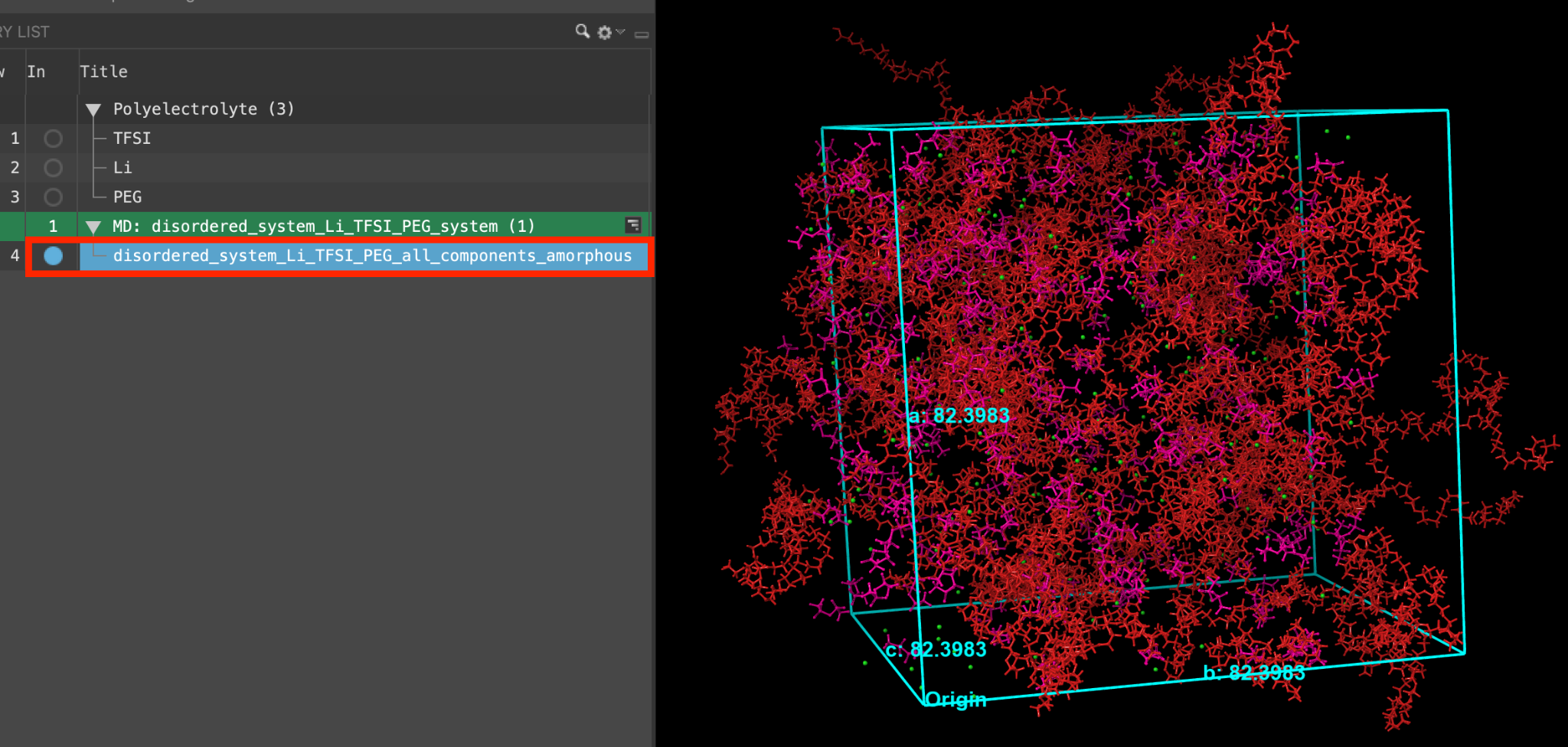
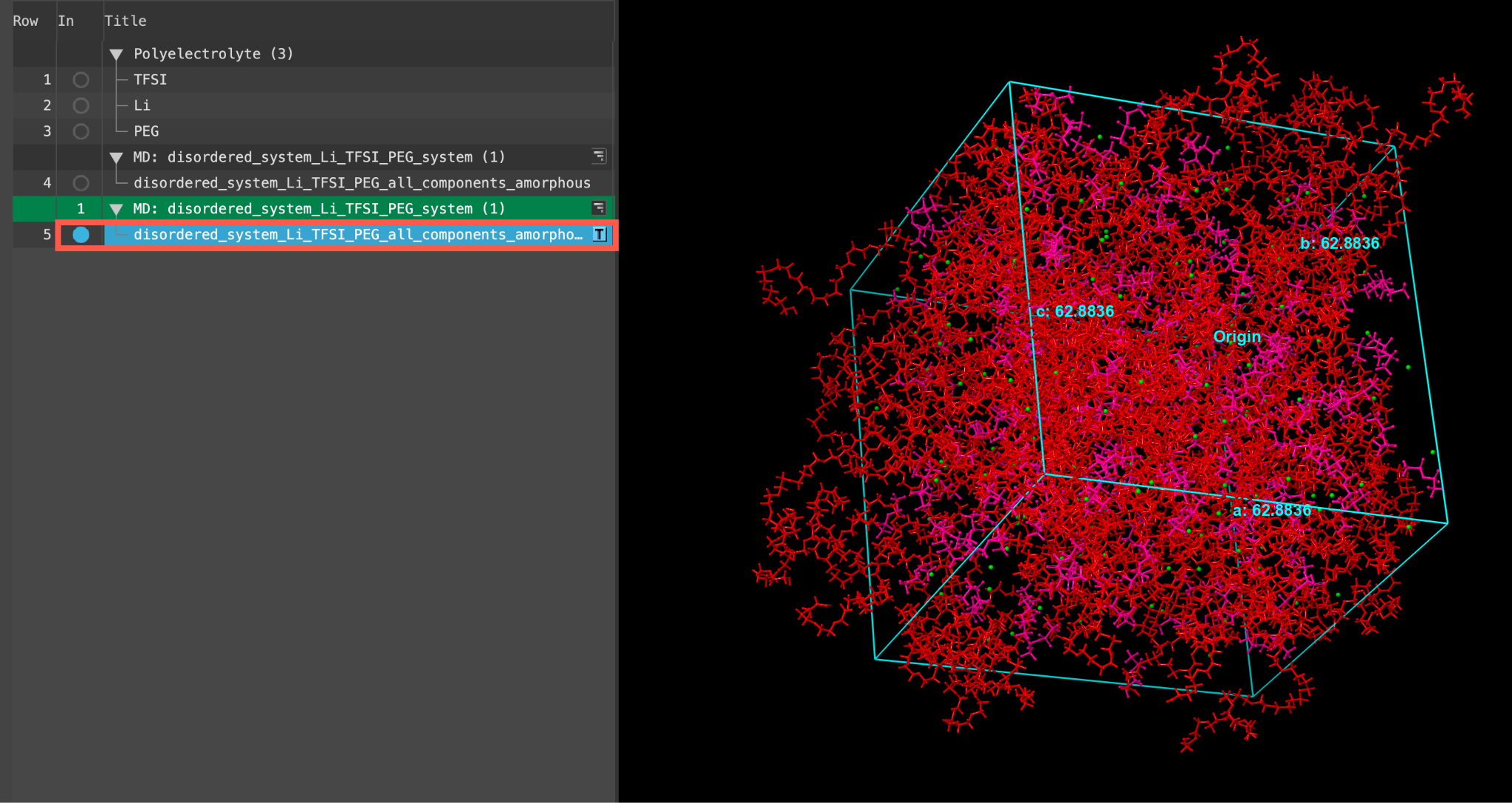
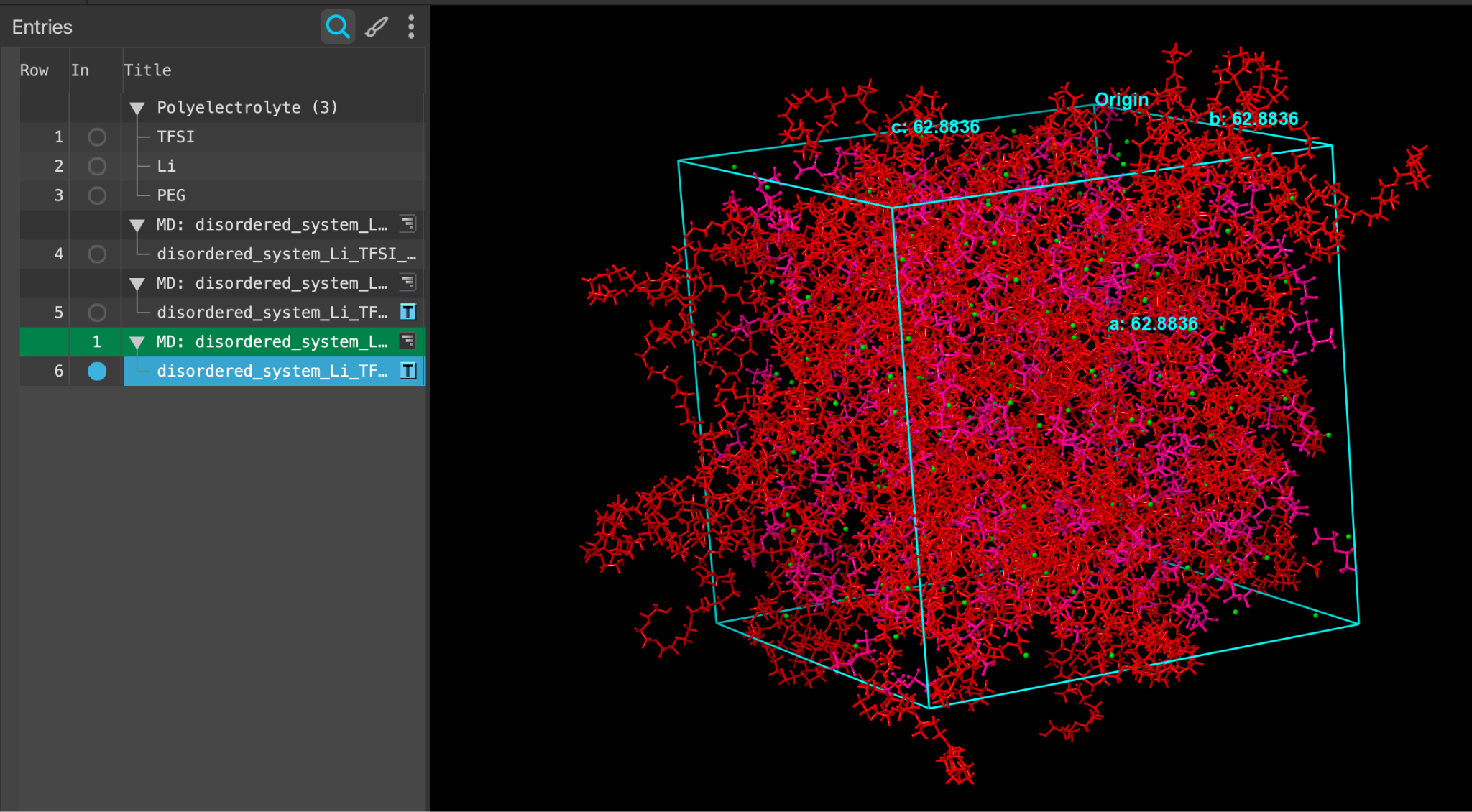
- When the job is finished or after importing, select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue disordered_system_Li_TFSI_PEG_all_components_amorphous
It is recommended to use the following best practices for preparing a system for diffusion calculations:
- Equilibrate the system in the NPT ensemble to ensure that equilibrium density is obtained
- Equilibrate the system in the NVT ensemble, using the average equilibrated box vectors from the NPT simulation to equilibrate the temperature for a structure at average density
- Do a production run in the NVE ensemble using the output of the NVT simulation to focus the simulation only on movement of the molecules
In our case, we also want to study the system at 500 K, which is an experimentally relevant condition. As a result, before the first NPT equilibration, we will compress the cell and then use an annealing procedure to heat the system to 500 K.
Note that the NVE stage can be included in the diffusion calculation setup to follow, and so in this section of the tutorial, we will finish the MD simulation after the NVT stage. In Section 4, we will do the production run in the NVE ensemble. Alternatively, you can run all of the MD simulation with the MD Multistage Workflow and use the diffusion setup to only perform the diffusion coefficient calculations.
For more information about best practices for diffusion calculations, see the References.
- Go to Tasks > Materials > Classical Mechanics > MD Simulations > MD Multistage Workflow
- The MD Multistage Workflow panel opens
- Alternatively, use the WAM (
 ) button in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion to access the panel
) button in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion to access the panel
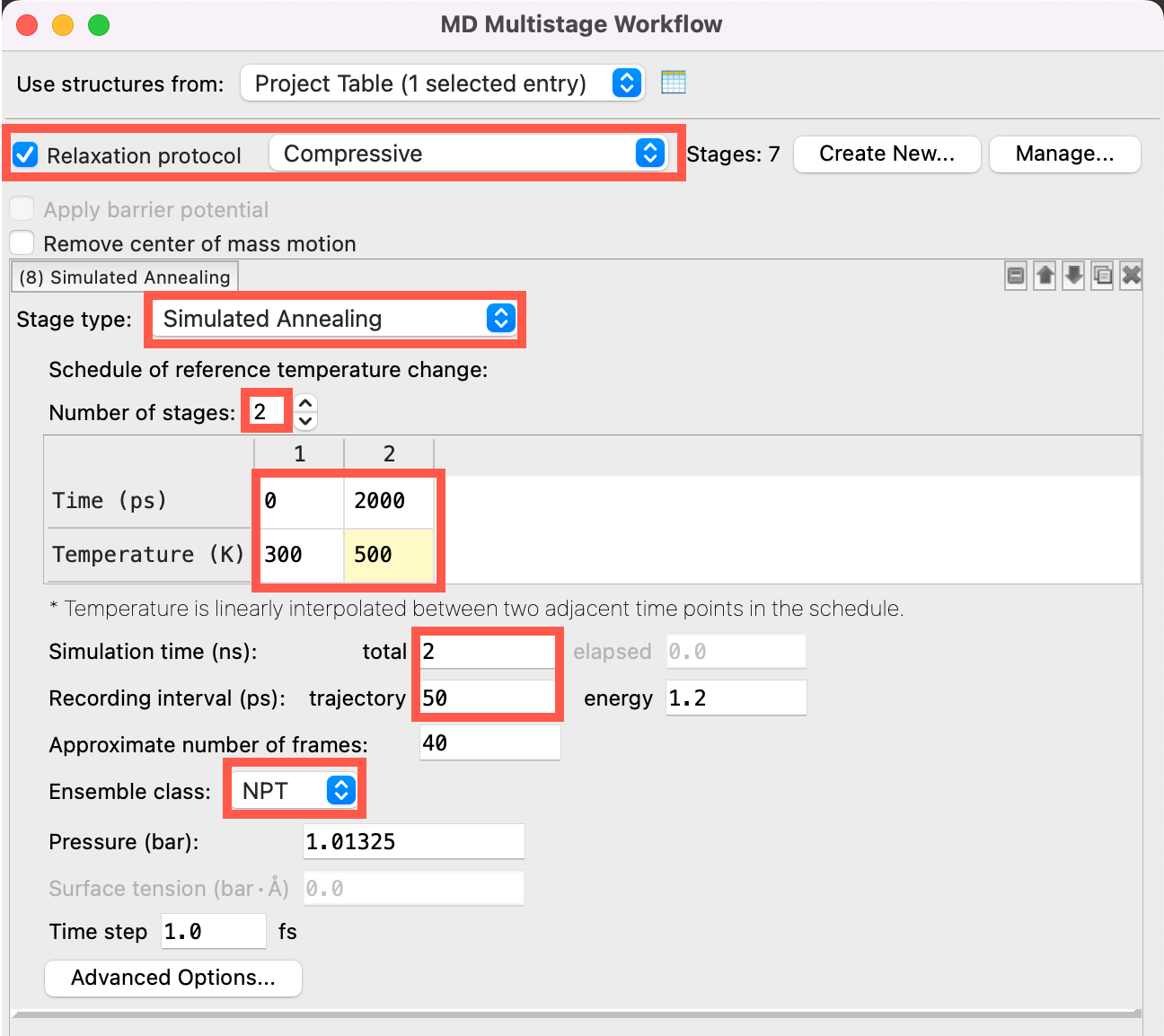
- Check Relaxation protocol and choose Compressive from the dropdown list
- For Stage (8), change to Simulated Annealing
- Change the Number of stages to 2
- Set the Time to 0, 2000 and the Temperature to 300, 500
- The system temperature will be linearly ramped from 300 to 500 K over the course of 2 ns
- Change the Simulation time to 2 ns and the Trajectory Recording interval to 50 ps
- Change the Ensemble class to NPT
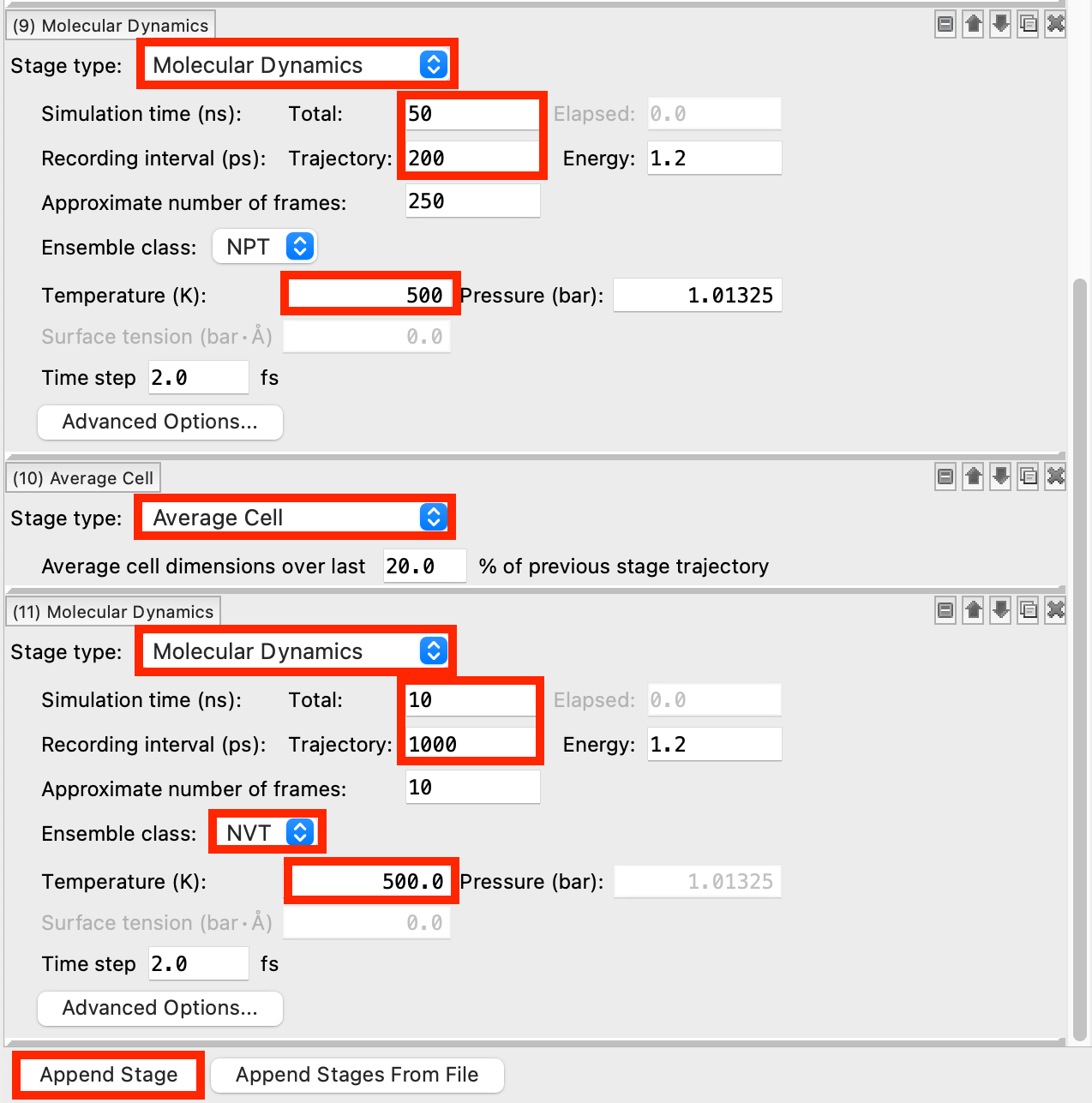
- Click Append Stage
- For Stage (9), change to Molecular Dynamics
- Change the Simulation time to 50 ns and the Trajectory Recording interval to 200 ps
- This recording interval was chosen to give a sufficient number of samples to the Average Cell stage that follows
- Change the Temperature to 500 K
- Click Append Stage
- For Stage (10), change to Average Cell
- Click Append Stage
- For Stage (11), change to Molecular Dynamics
- Change the Simulation time to 10 ns and the Trajectory Recording interval to 1000 ps
- Change the Ensemble class to NVT
- Change the Temperature to 500 K
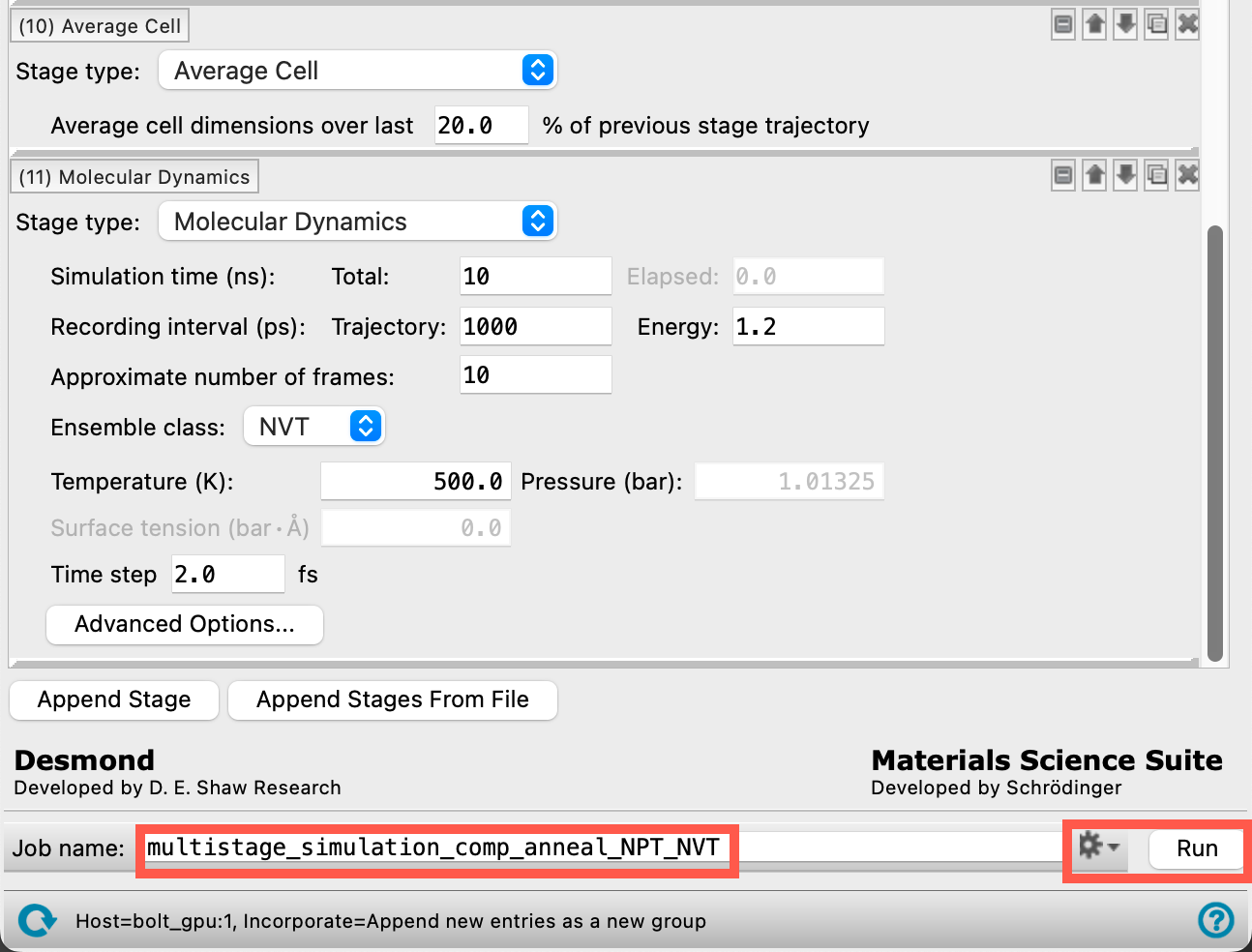
- Change the Job name to multistage_simulation_comp_anneal_NPT_NVT
- Adjust the job settings (
 ) as needed
) as needed
- This job requires a GPU host. The job can be completed in about 12 hours
- If you would prefer not to run the job, import
Section_03 > multistage_simulation_comp_anneal_NPT_NVT > multistage_simulation_comp_anneal_NPT_NVT-out.cmsfrom the provided tutorial files via File > Import Structures. Otherwise, click Run - Close the MD Multistage Workflow panel
Note: If the labels are still turned on and are cluttering the workspace, turn them off with the  button in the bottom right corner of the workspace.
button in the bottom right corner of the workspace.
Note: For general practice with the MD Multistage Workflow panel, see the Disordered System Building and MD Multistage Workflows tutorial
- When the job is finished or after importing, select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue disordered_system_Li_TFSI_PEG_all_components_amorphous
Feel free to visualize this well-equilibrated output.
Note: Of course, depending on your own system of interest, it will be important to verify that your simulation equilibrates. See the Disordered System Building and MD Multistage Workflows tutorial for more discussion and best practices.
4. Performing and Analyzing Diffusion Coefficient Calculations
Now that we have equilibrated the system with our prescribed protocol, we are ready to move towards the final NVE simulation to be used for the Diffusion Coefficient calculation. In this section, we will use the Diffusion Coefficient panel to run the NVE simulation and collect diffusion data for the Li ions, and then use the Diffusion Coefficient Results panel to analyze the results.
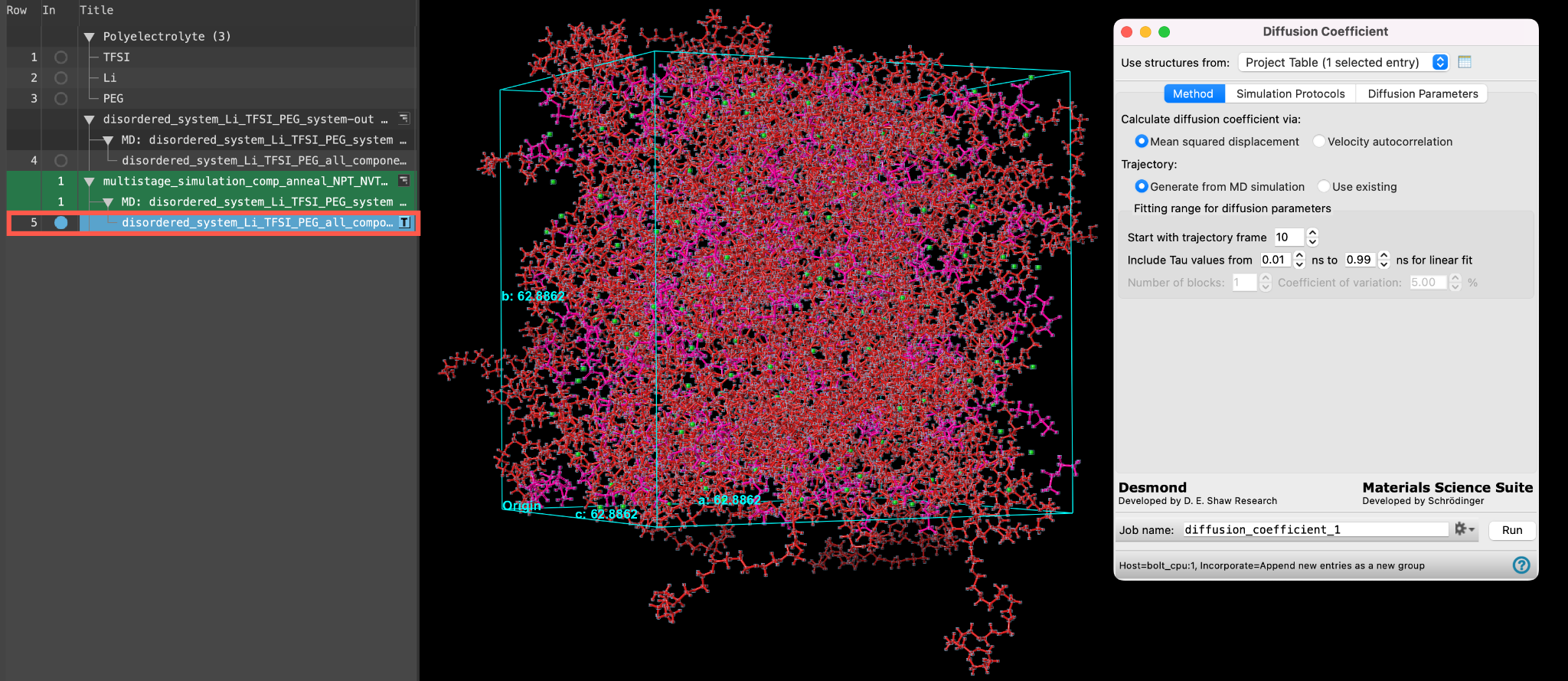
- Ensure that disordered_1system_Li_TFSI_PEG_all_components_amorphous is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and includedthe entry is represented in the Workspace, the circle in the In column is blue in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
- Go to Tasks > Materials > Classical Mechanics > Diffusion Coefficient > Diffusion Coefficient Calculations
- The Diffusion Coefficient panel opens
Before we prepare this job submission, let’s learn a bit more about the general capabilities of the Diffusion Coefficient calculations panel.
As mentioned earlier, there are two primary ways to measure diffusion constants from MD simulation: via Mean squared displacement (MSD) and via Velocity autocorrelation functions. The mean squared displacement method measures the average displacement of particles versus time. For Fickian diffusion, we expect a plot of mean squared displacement versus time to be linear and its slope to be proportional to the diffusion constant. The velocity autocorrelation method uses a Green-Kubo relationship which relates diffusion constant to the integral of the velocity autocorrelation function. Use the Calculation diffusion coefficient via radio buttons to choose a preferred method. The Velocity autocorrelation is usually less practical because velocities need to be recorded very frequently. Generally, mean squared displacement converges to a similar result with much coarser output frequency and no need to save velocities. Velocity autocorrelation can be used to confirm results of MSD.
With respect to the trajectory to analyze, you can either use the panel to set up the diffusion simulation (Generate from MD simulation), or you can use the panel on a previously simulated trajectory by choosing Use existing.
You must set an initial Fitting range, but note that this range can be changed in the analysis stage with the viewer panel.
On the Simulation Protocols tab, set up the MD if you are running a new simulation at this stage.
In the Diffusion Parameters tab, you can narrow the calculation to only the species of interest. It is most convenient to use the Atom Specification Language (ASL) to make a selection, though other selection methods are available.
Finally, anisotropic diffusion coefficients can be calculated using the parameters available.
Visit the help documentation to read more about using the panel.
As mentioned earlier, we will use the Mean squared displacement method here, and we will run a production NVE simulation at this stage.
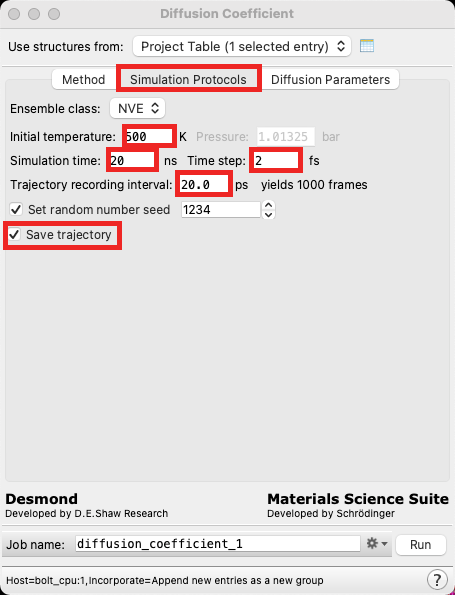
- Maintain the defaults on the Method tab and proceed to the Simulation Protocols tab
- Change the Initial temperature to 500 K, the Simulation time to 20 ns, the Time step to 2 fs and the Trajectory recording interval to 20 ps
- Note that it is important to yield a significant amount of frames for calculating the diffusion constant
- Click Save trajectory
- This will take some extra memory, but can be useful if we want to visualize the NVE simulation or perform subsequent analysis beyond the diffusion calculation (for example, Polymer Electrolyte Analysis)
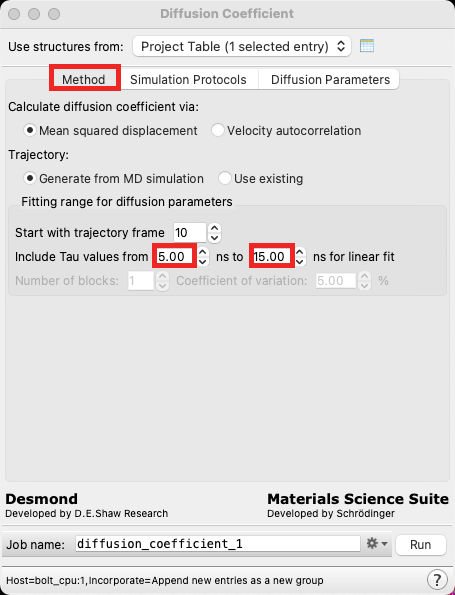
- Return to the Method tab
- In the Fitting range section, change the Include Tau values from range from 5 to 15 ns
Note: During the analysis stage, the Tau values can be adjusted again, and so there is no need to take care when selecting this range at this moment. However, if you were running a batch job (say, hundreds of systems at many temperatures) without the intention of using the viewer panel to analyze the diffusivity, you may want to use care when deciding this range initially.
Now we can specify which ions or molecules for which to calculate diffusion parameters.
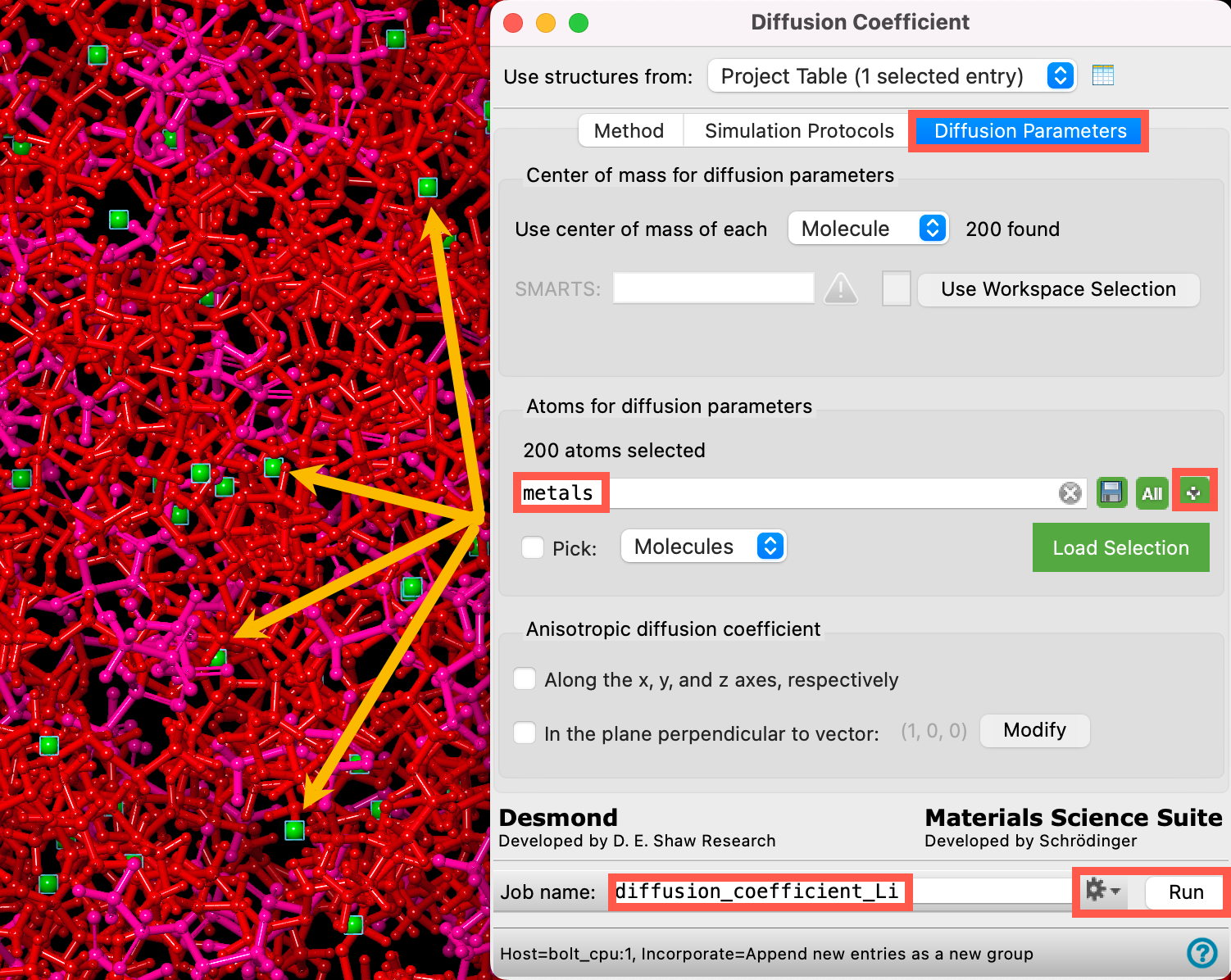
- Go to the Diffusion Parameters tab
- For Atoms for diffusion parameters, we must define the lithium ions. Clear the default (
 ) and use the
) and use the  button to choose Metal Atoms
button to choose Metal Atoms
- The panel updates to show metals and 200 atoms selected
- Confirm that in the workspace, the lithium ions are all selected
- Other methods for selecting the lithium atoms are available including Pick or the other dropdowns in the panel
- Read more about ASL
- Change the Job name to diffusion_coefficient_Li
Adjust the job settings ( ) as needed. This job requires a CPU and GPU host (for the analysis and MD, respectively). The job can be completed in about 2 hours.
) as needed. This job requires a CPU and GPU host (for the analysis and MD, respectively). The job can be completed in about 2 hours.
- If you would prefer not to run the job, import
Section_04 > diffusion_coefficient_Li > diffusion_coefficient_Li-out.cmsfrom the provided tutorial files via File > Import Structures in the diffusion_coefficient_Li directory. Otherwise, click Run - Close the Diffusion Coefficient panel
When the job completes or after importing, a new entry is added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion entitled disordered_system_Li_TFSI_PEG_all_components_amorphous
- Select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue the output in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and workspacethe 3D display area in the center of the main window, where molecular structures are displayed, respectively.
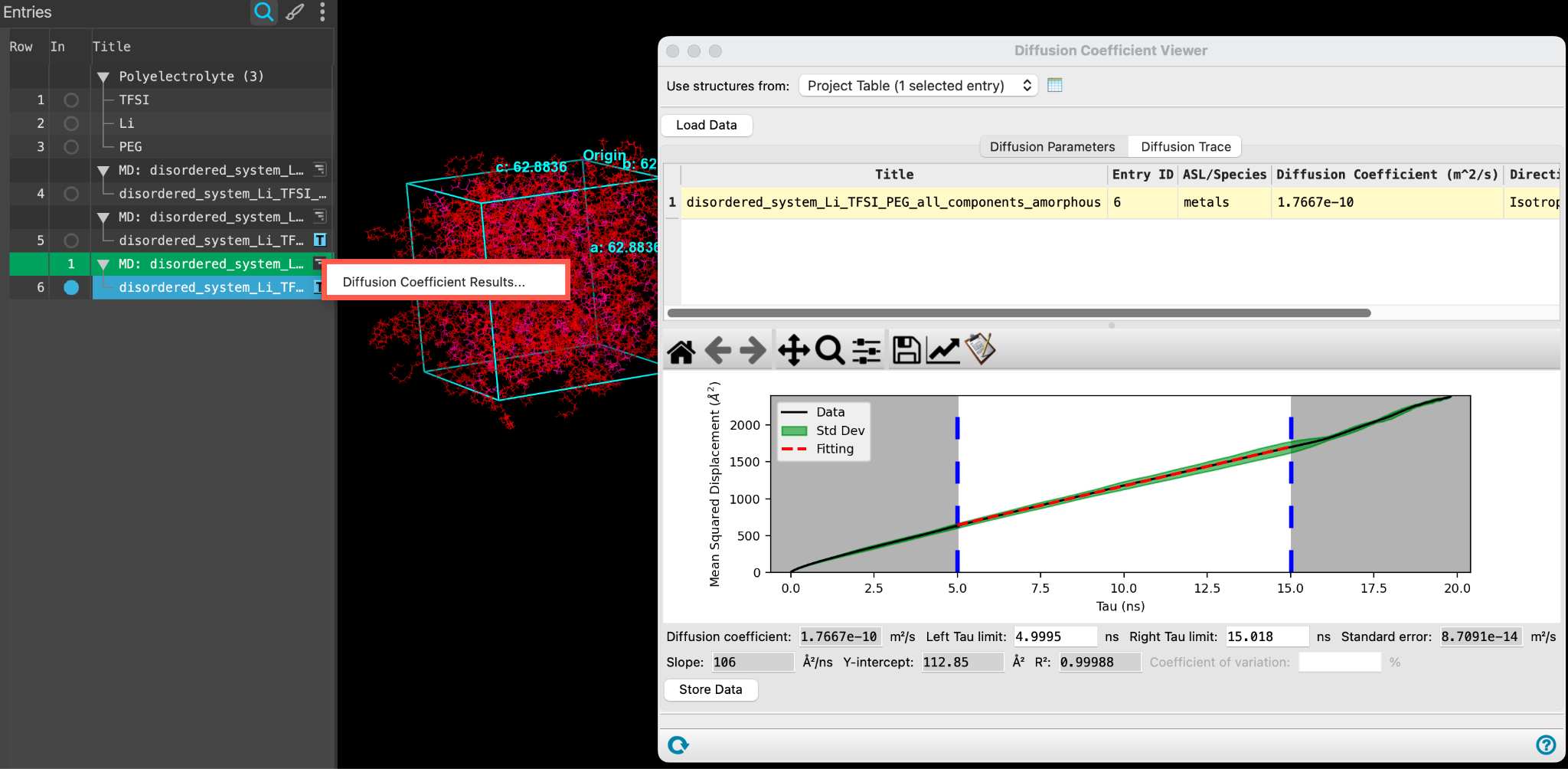
- Use the WAM (workflow action menu) button (
 ) to open the Diffusion Coefficient Viewer
) to open the Diffusion Coefficient Viewer
- Alternatively, access the panel via Tasks > Materials > Classical Mechanics > Diffusion Coefficient > Diffusion Coefficient Results
- The Diffusion Coefficient Viewer panel opens
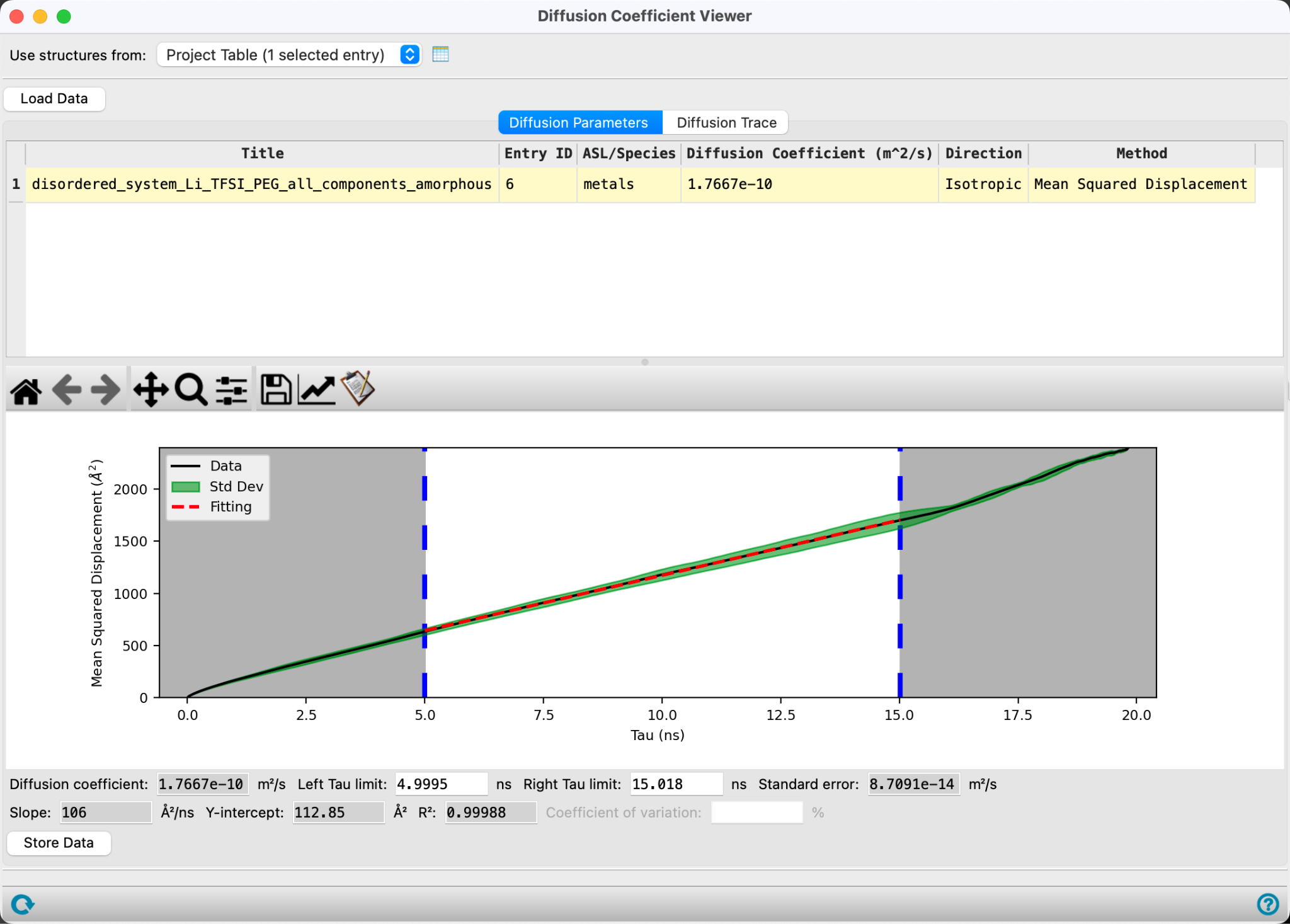
The Diffusion Parameters tab displays values of diffusion coefficients and includes a plot of the mean square distance (MSD) against simulation time difference (Tau).
You can manually adjust the bounds of the linear fit using the interactive sliders in the plot. The diffusion coefficient based on these modified bounds is computed in real-time below. In general, you should fit into the linear region of the curve. There is no consensus method for choosing the bounds of the fit, therefore it’s important to report these bounds for the sake of reproducibility. As a very loose recommendation, one should exclude the short timescale behavior, where the curve follows a ballistic trajectory and the atoms velocities have not been greatly impacted by interaction with surrounding atoms, as well as long times where the uncertainty is higher.
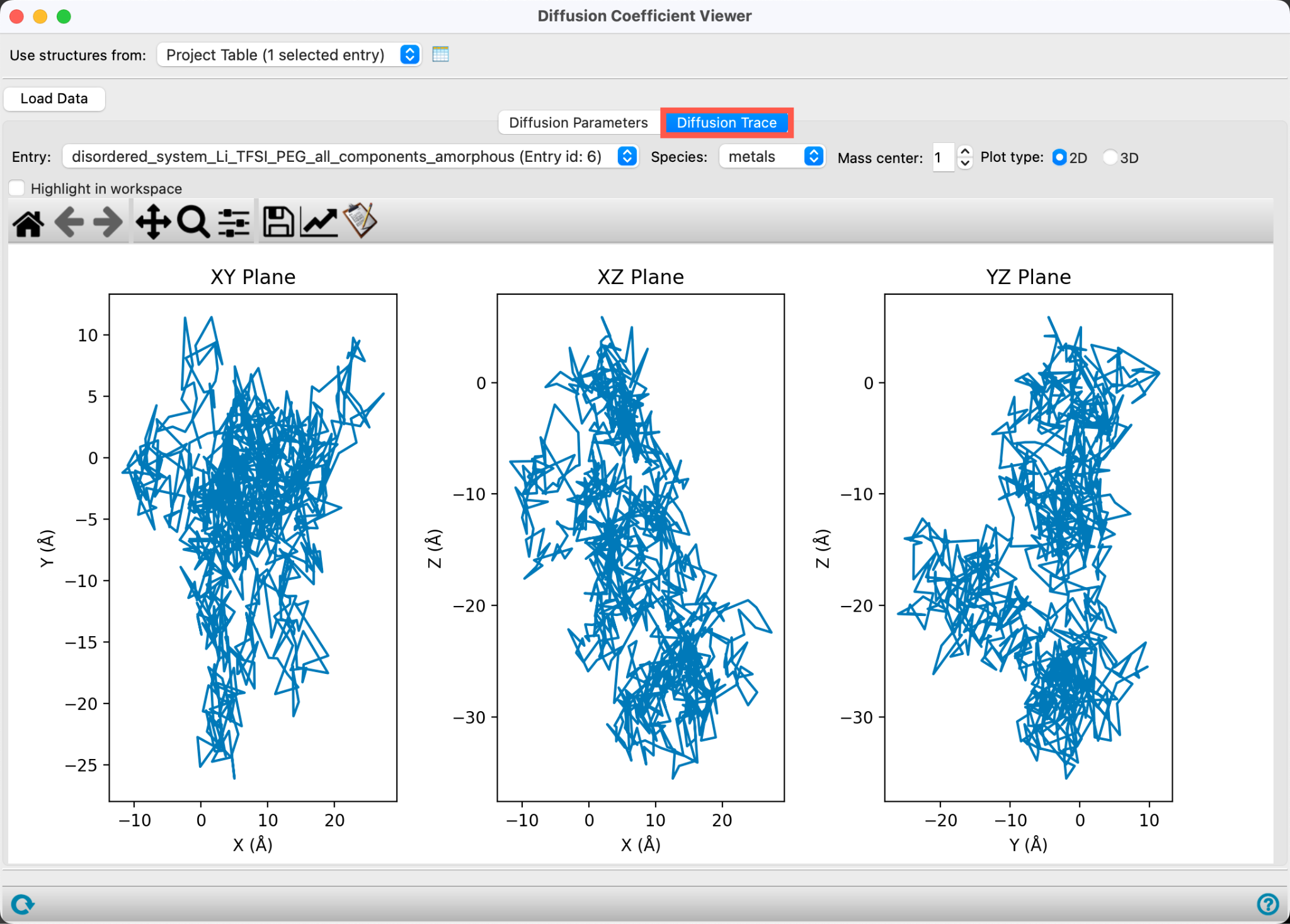
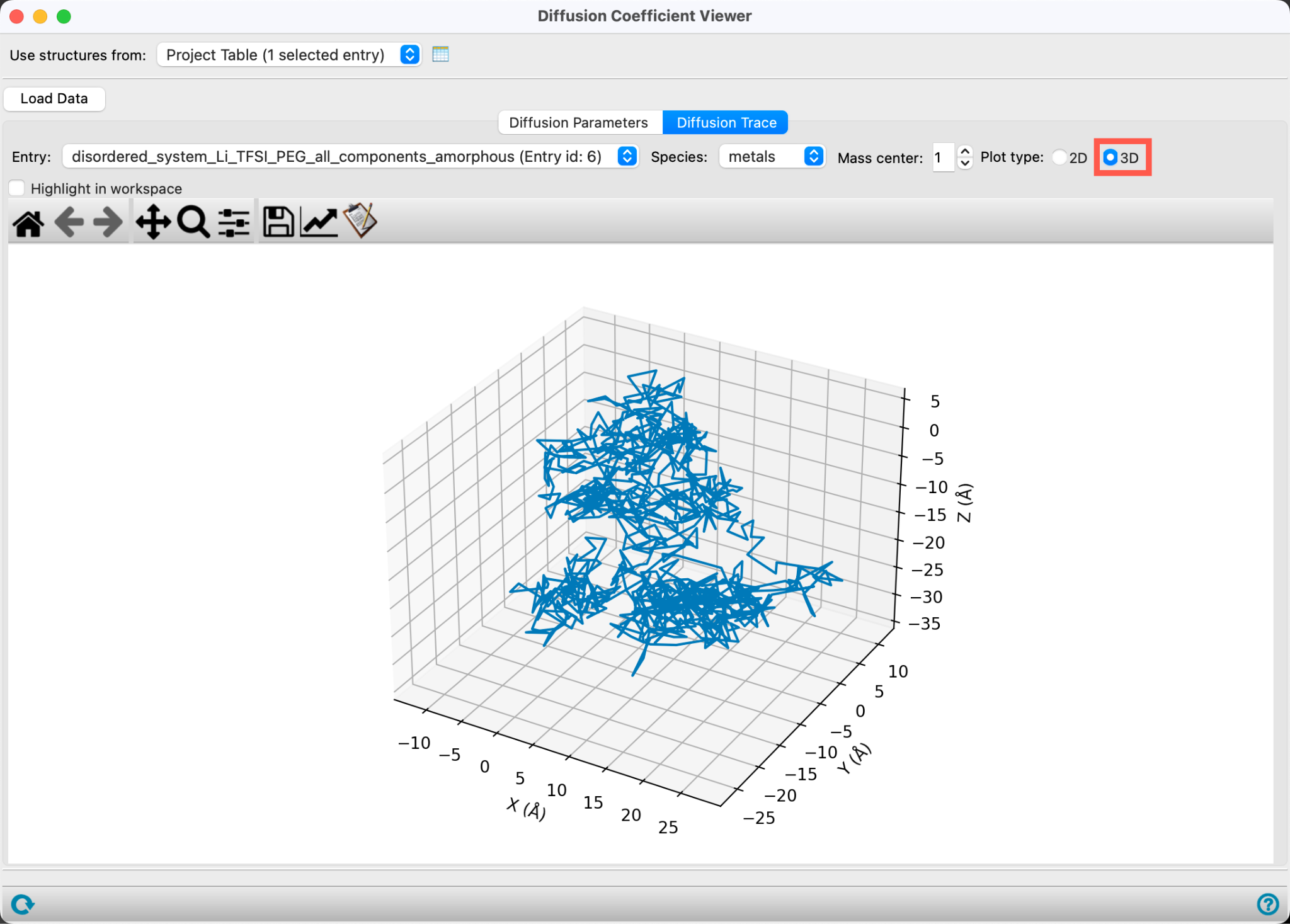
- Go to the Diffusion Trace tab
The trace for a specified Mass center (by default 1, the first lithium ion) is shown in three 2D plots. These plots trace the movement of a single species over the course of the trajectory. They are a visual tool that can help identify anisotropic diffusive behavior as well as deviations from Brownian motion. The 2D plots show projections of the coordinates in each of the three planes: XY, XZ and YZ.
- Change the Plot type to 3D
The trace for a specified Mass center (by default 1, the first lithium ion) is shown in a 3D plot
Feel free to explore the various traces for additional Li ions by changing the Mass center input.
Note: The 3D plot can be rotated (left-click and drag) or zoomed (right-click and drag) similar to typical mouse actions in the MS Maestro workspacethe 3D display area in the center of the main window, where molecular structures are displayed
5. Conclusion and References
In this tutorial, we learned how to best prepare a prototypical Li/TFSI and polymer system for a Diffusion Coefficient calculation. We then used the Diffusion Coefficient Calculation and Viewer panels to determine the diffusion coefficient for the Li cations. Studying the diffusion coefficients for the Li cations can help study electrolyte performance. See below for other related tutorials.
For further learning:
For introductory content, focused on navigating the Schrödinger Materials Science interface, an Introduction to Materials Science Maestro tutorial is available. Please visit the materials science training website for access to 70+ tutorials. For scientific inquiries or technical troubleshooting, submit a ticket to our Technical Support Scientists at help@schrodinger.com.
For self-paced, asynchronous, online courses in Materials Science modeling, including access to Schrödinger software, please visit the Schrödinger Online Learning portal on our website.
For some related practice, proceed to explore other relevant tutorials:
- Disordered System Building and Molecular Dynamics Multistage Workflows
- Building, Equilibrating and Analyzing Amorphous Polymers
- Building Solvated Systems
- Crosslinking Polymers
- Building a Semicrystalline Polymer
- Thermal Conductivity
- Polymer Property Prediction
- Penetrant Loading
- Molecular Dynamics Simulations for Active Pharmaceutical Ingredient (API) Miscibility
- Cluster Analysis
- Computing Atomic Charges
- Calculating Surfactant Tilt and Electrostatic Potential of a Bilayer System
- Building a Polymer-Polymer Interface
- Building a Carbohydrate Polymer
- Applying Barrier Potentials for Molecular Dynamics Simulations
- Viscosity
- Evaporation
- Droplet Contact Analysis
- Liquid Electrolyte Properties: Part 1 and Part 2
- Solid Electrolyte Interphase Calculations
- Building a Coarse-Grained Surfactant Model with Martini Force Field
- Ibuprofen Cyclodextrin Inclusion Complexes with the Martini Coarse-Grained Force Field
- Machine Learning Force Field
For further reading:
- See the help documentation for the Diffusion Coefficient and the Diffusion Coefficient Viewer
- Atomistic Description of Ionic Diffusion in PEO-LiTFSI: Effect of Temperature, Molecular Weight, and Ionic Concentration. DOI:10.1021/acs.macromol.8b01753
- Effects of Temperature Control Algorithms on Transport Properties and Kinetics in Molecular Dynamics Simulations. DOI:10.1021/ct400109a
- Best Practices for Computing Transport Properties 1. Self-Diffusivity and Viscosity from Equilibrium Molecular Dynamics. DOI:10.33011/livecoms.1.1.6324
- Prediction of self-diffusion coefficients of chemically diverse pure liquids by all-atom molecular dynamics simulations. DOI:10.1002/jcc.26975
6. Glossary of Terms
Entry List - a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
Included - the entry is represented in the Workspace, the circle in the In column is blue
Project Table - displays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data
Recent actions - This is a list of your recent actions, which you can use to reopen a panel, displayed below the Browse row. (Right-click to delete.)
Scratch Project - a temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project
Selected - (1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Working Directory - the location where files are saved
Workspace - the 3D display area in the center of the main window, where molecular structures are displayed