Electroporation
Tutorial Created with Software Release: 2026-1
Topics: Consumer Packaged Goods , Polymeric Materials
Methodology: All-Atom Molecular Dynamics
Products Used: Desmond , MS Maestro
|
1.5 GB |
This tutorial is written for use with a 3-button mouse with a scroll wheel.
Words found in the Glossary of Terms are shown like this: Workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Abstract:
In this tutorial, we will learn to use the electroporation calculations panel to apply an electric field to bacterial membranes to calculate pore formation using molecular dynamics simulations.
Tutorial Content
1. Introduction to Electroporation
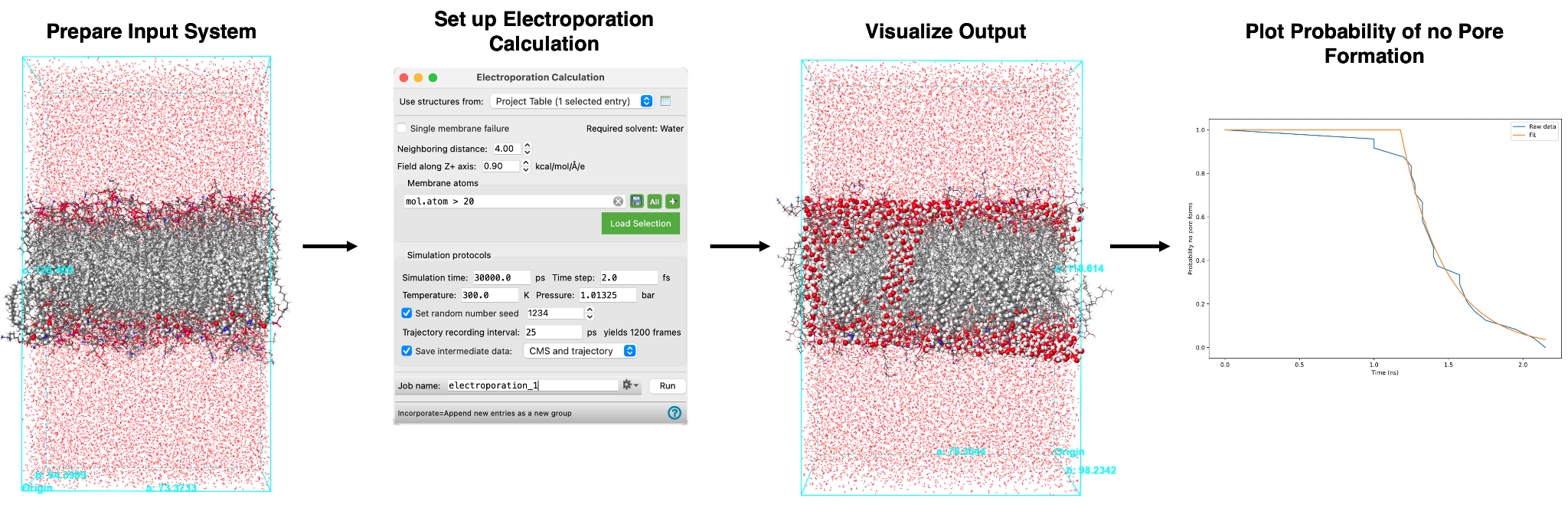
The Electroporation Calculation panel allows you to investigate membrane stability by applying electric fields to a membrane system to simulate pore formation. In Schrödinger’s Materials Science (MS) Maestro suite, membrane stability is characterized by a cumulative probability as a function of time. It can be calculated following a straightforward workflow as summarized here:
The calculation involves two components; first, an electric field, as defined by the user, is applied to the system. Next, the system is relaxed using molecular dynamics (MD) simulation. The electric field will cause a pore to form in the membrane at some point during the simulation, which is detected by the distance between water clusters decreasing past a user-defined threshold. The simulation is performed for a series of input structures to generate multiple sets of data for pore formation. After the calculation is complete, a plot of probability of no pore formation is produced and is utilized to analyze the simulation results.
Electroporation calculations are useful for a variety of applications, for example, studying membrane stability as a function of changing membrane compositions which is significant in the consumer packaged goods space. The impact of changes in membrane structures on their properties can be better understood using this tool.
In this tutorial, we will study two examples of electroporation calculations; (1) in Section 3 we will calculate the probability of pore formation for a Staphylococcus aureus (S. aureus) membrane and (2) in Section 4 we will calculate the probability of pore formation for an S. aureus membrane which has been asymmetrically loaded with cationic C16-diethyl ester dimethyl ammonium (BQT). We will then compare the pore formation results for both structures. This exercise is adapted from the following paper, Bacterial Membranes Are More Perturbed by the Asymmetric Versus Symmetric Loading of Amphiphilic Molecules [DOI:110.3390/membranes12040350].
For a more in depth explanation of building and equilibrating structured liquid systems such as those used here, see the Calculating Surfactant Tilt and Electrostatic Potential of a Bilayer System tutorial.
2. Creating Projects and Importing Structures
At the start of the session, change the file path to your chosen Working Directorythe location where files are saved in MS Maestro to make file navigation easier. Each session in MS Maestro begins with a default Scratch Projecta temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project, which is not saved. A MS Maestro project stores all your data and has a .prj extension. A project may contain numerous entries corresponding to imported structures, as well as the output of modeling-related tasks. Once a project is saved, the project is automatically saved each time a change is made.
Structures can be built in MS Maestro or can be imported using File > Import Structures (or drag-and-dropped), and are added to the Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data. The Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion is located to the left of the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed. The Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data can be accessed by Ctrl+T (Cmd+T) or Window > Project Table if you would like to see an expanded view of your project data.
- Double-click the Materials Science icon
- (No icon? See Starting Maestro)
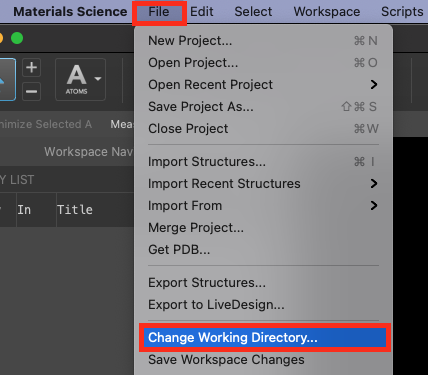
- Go to File > Change Working Directory
- Find your directory, and click Choose
- Pre-generated files are included for running jobs or examining output. Download the zip file here: schrodinger.com/sites/default/files/s3/release/current/Tutorials/zip/electroporation.zip
- After downloading the zip file, unzip the contents in your Working Directorythe location where files are saved for ease of access throughout the tutorial
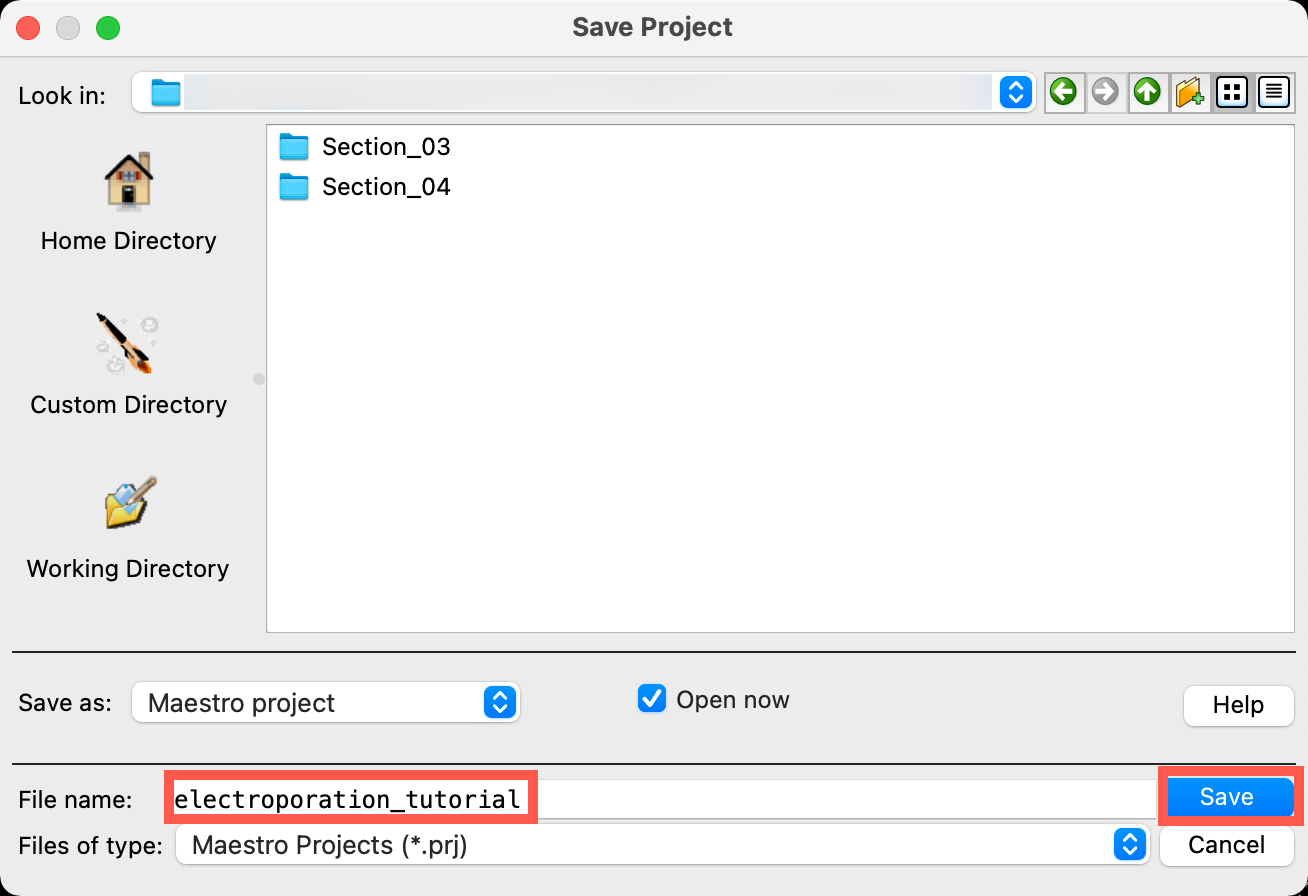
- Go to File > Save Project As
- Change the File name to electroporation_tutorial, click Save
- The project is now named
electroporation_tutorial.prj
- The project is now named
3. Electroporation of the S. aureus Membrane
In this section, we will use the Electroporation panel to study the stability of the S. aureus membrane. The input systems should be well-relaxed MD outputs, as we have provided here. We will then visualize the output of the electroporation calculation.
For our first example, our structure of interest has been built in advance and is available for importing in the next step. We will import 24 structure files (.cms) of an equilibrated cell of the S. aureus membrane surrounded by water. These 24 structure files were generated by building four independent replicates of the membrane system, equilibrating the four systems, and then taking six equilibrated frames from each of the four resulting trajectories. We use 24 structures here according to what was done in the literature.
If you are interested in building and equilibrating the system yourself, feel free to do so. For practice building and equilibrating systems containing membranes, see the Calculating Surfactant Tilt and Electrostatic Potential of a Bilayer System tutorial to review the relevant tools.
- Go to File > Import Structures
- Navigate to where you downloaded the tutorial files (presumably your working directory) and choose
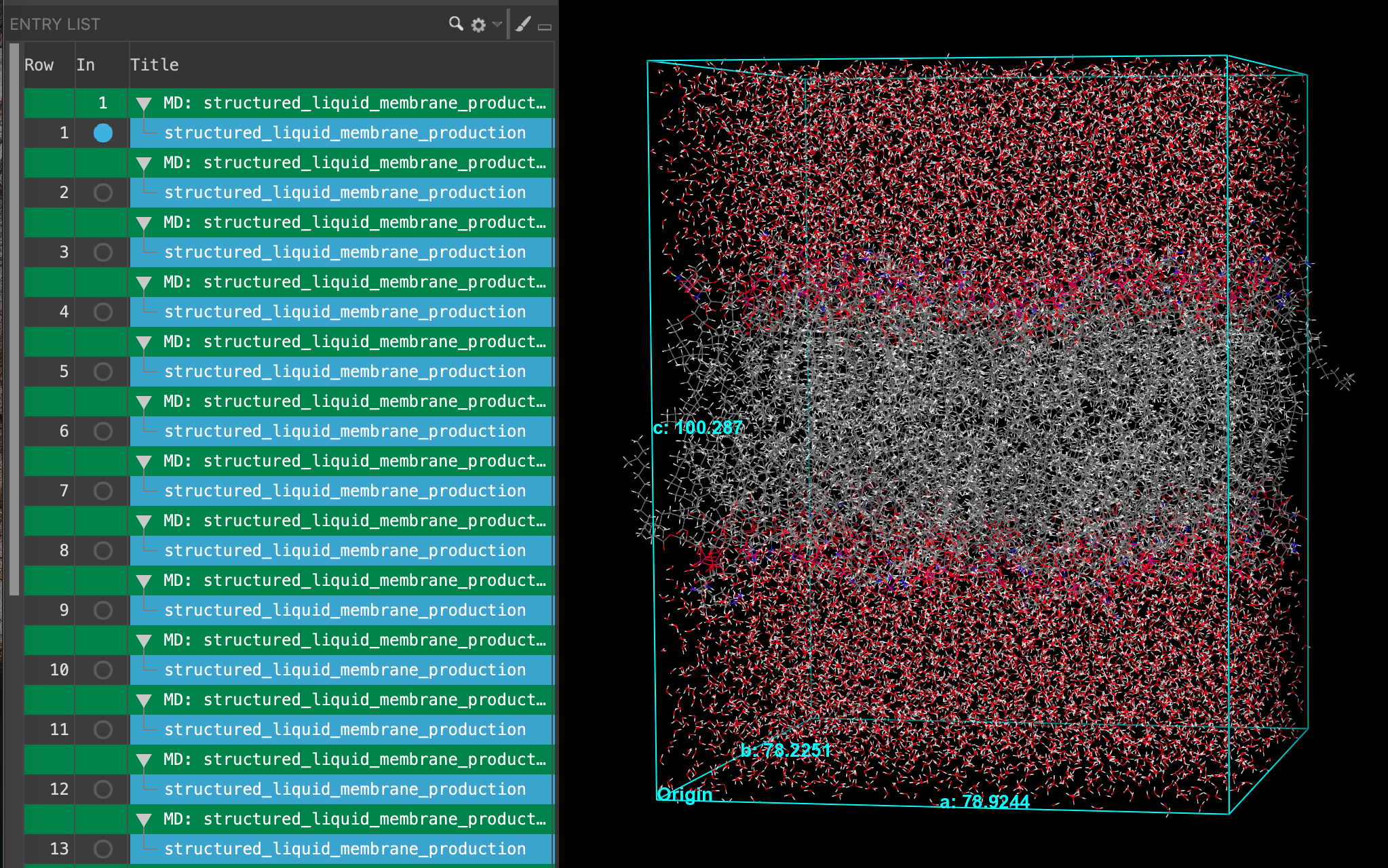
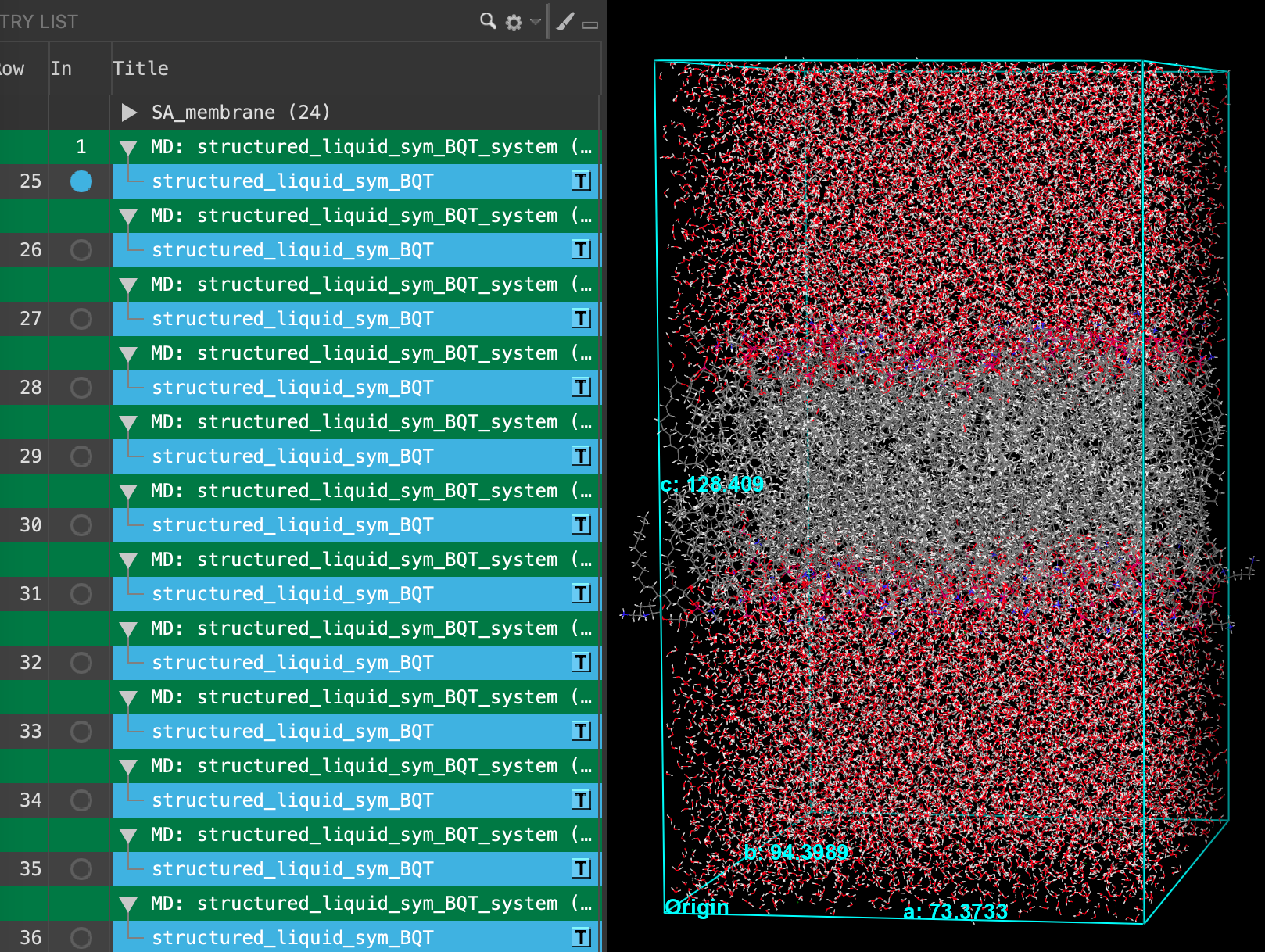
Section_03 > SA_membrane_input > structured_liquid_membrane_production-n_system-out.cms. Where n = 1 - 24. Click Open- 24 new entries are added to the entry list
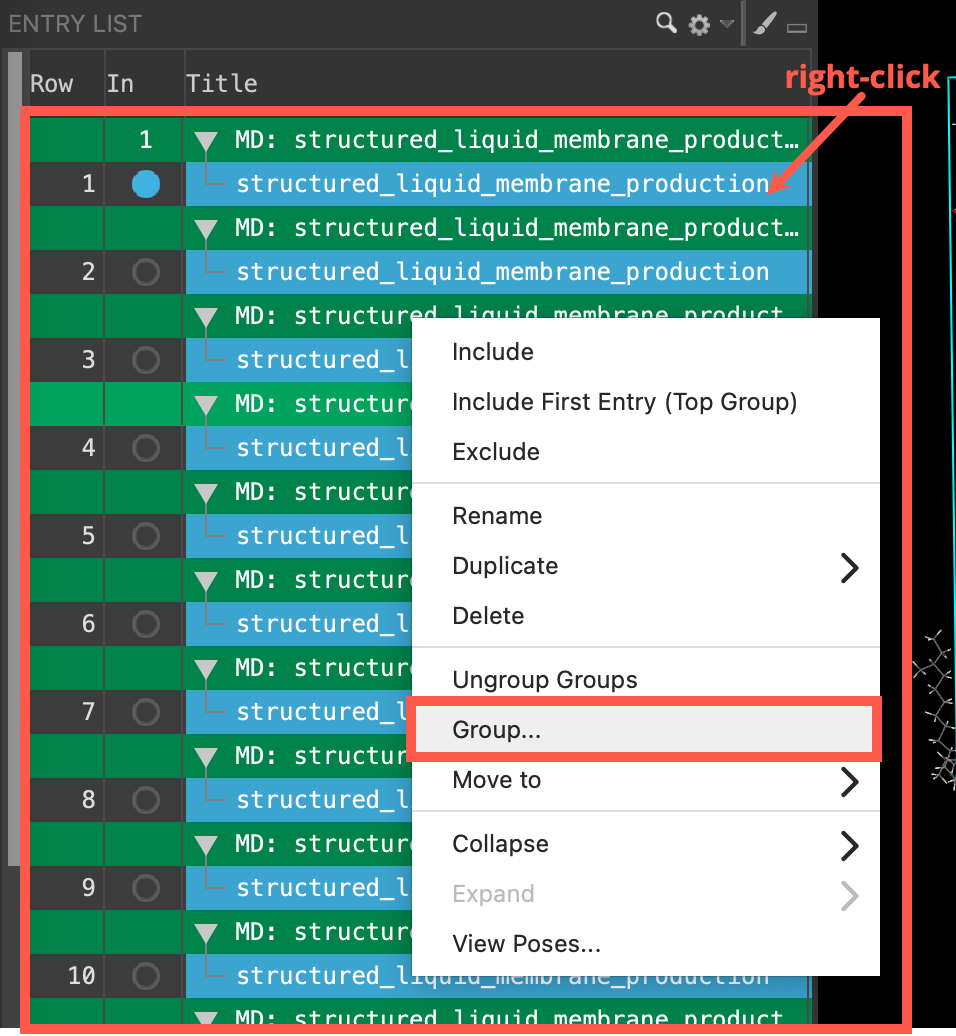
- Select all 24 entries (Shift+Click)
- Right-click on the entries and click Group
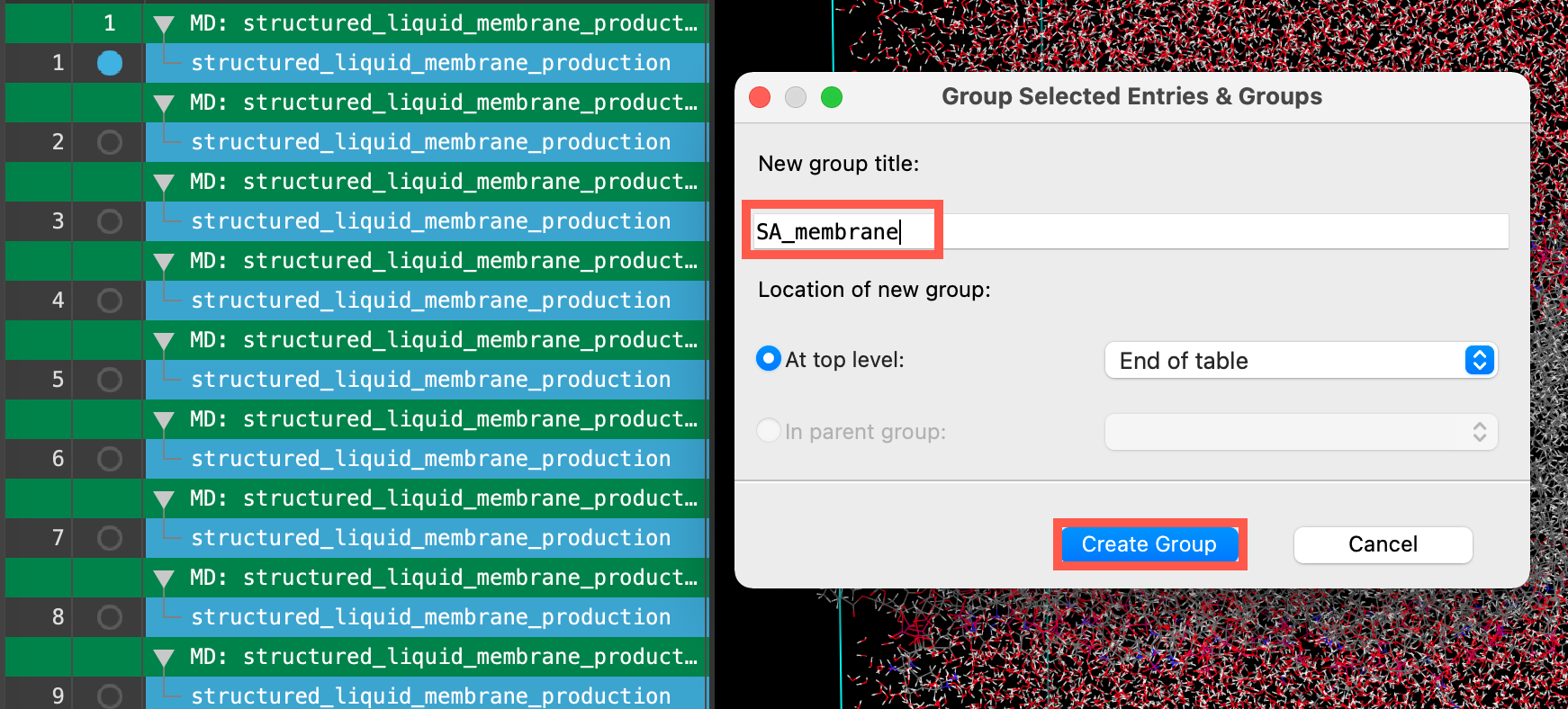
- Enter SA_membrane for the New group title
- Click Create Group
- Ensure that the SA_membrane entry group is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Classical Mechanics > Electroporation
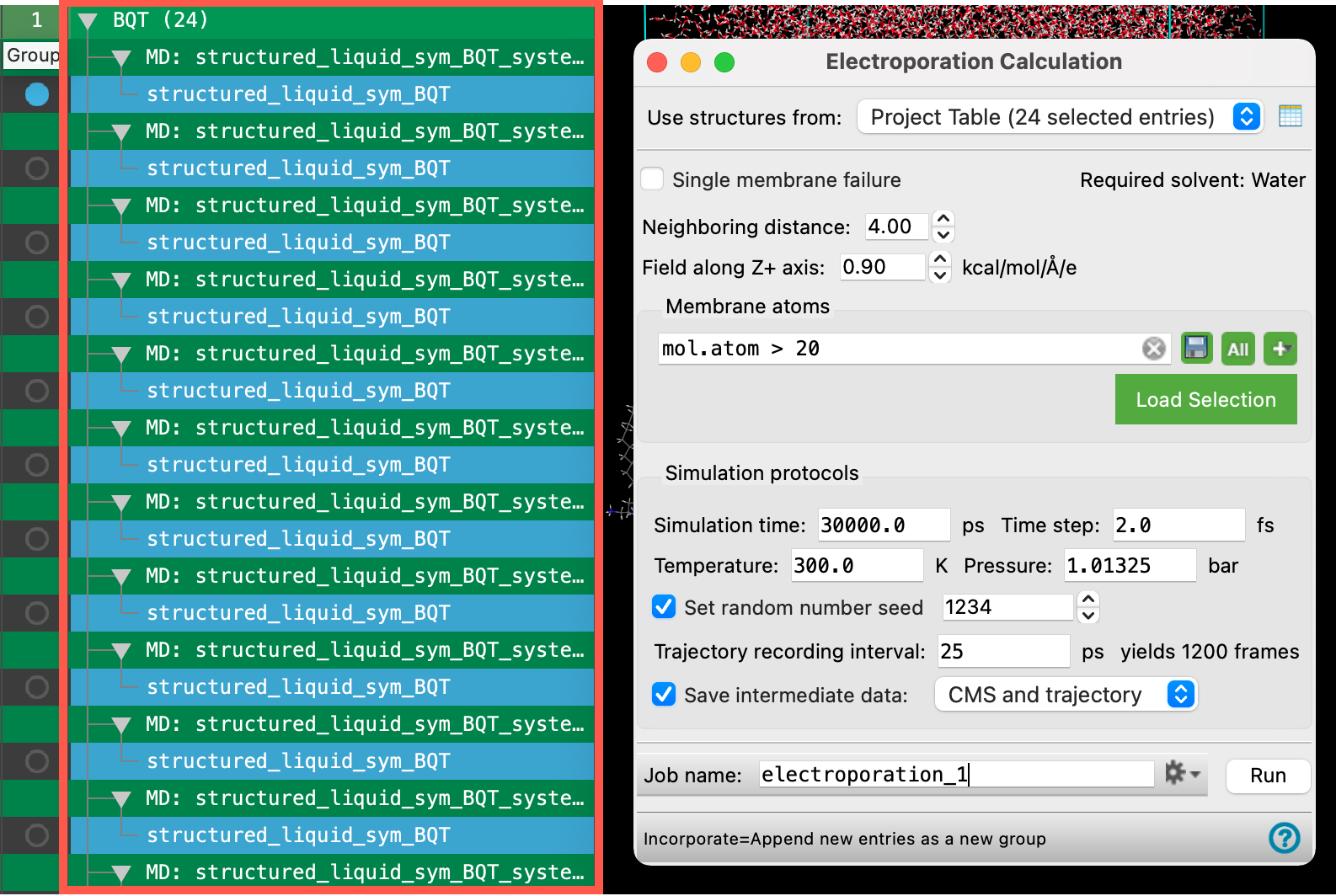
- The Electroporation panel opens
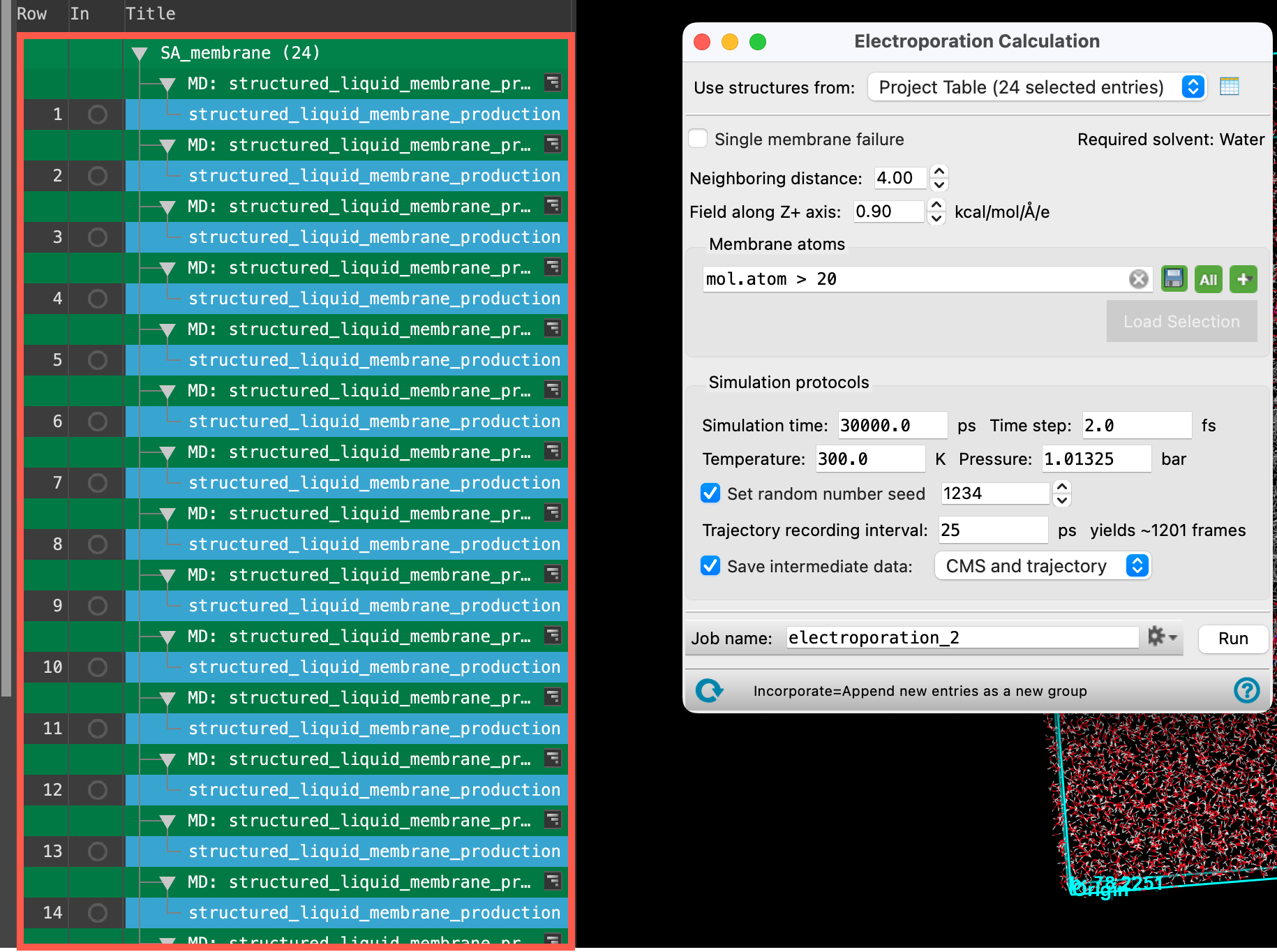
Let’s learn about the settings and capabilities of the Electroporation panel a bit more:
- The input structures for the electroporation panel, typically 20-25 replicates of a well-equilibrated system, contain a membrane and water as the solvent. The Structured Liquid Builder can be used to generate such a system.
- A pore is detected when water clusters come within a certain distance of each other, the Neighboring distance. This parameter allows you to enforce strict or loose criterion for pore formation.
- Pore formation probability can be calculated for the rupturing of a single membrane, Single membrane failure, or when all membranes present in the structure are ruptured.
- The Field along Z-axis parameter allows you to apply an electric field (of positive or negative value) along the Z-axis.
- Membrane atoms can be defined using ASL selection.
- The Simulation Protocol section allows us to specify parameters for the NPT MD simulations to be run on our input structures.
- The Simulation time should be sufficient to allow pores to begin forming at the specified electric field.
- Trajectory data is generated for each of the structures as a part of the electroporation calculation. Be cautious when choosing to Save intermediate data for the CMS and trajectory or CMS files only as the trajectories are disk space intensive.
Visit the help documentation for a complete summary of the parameters.
MD Simulations have a number of files associated with the job, for a full description of each file type see the help documentation on Desmond Files.
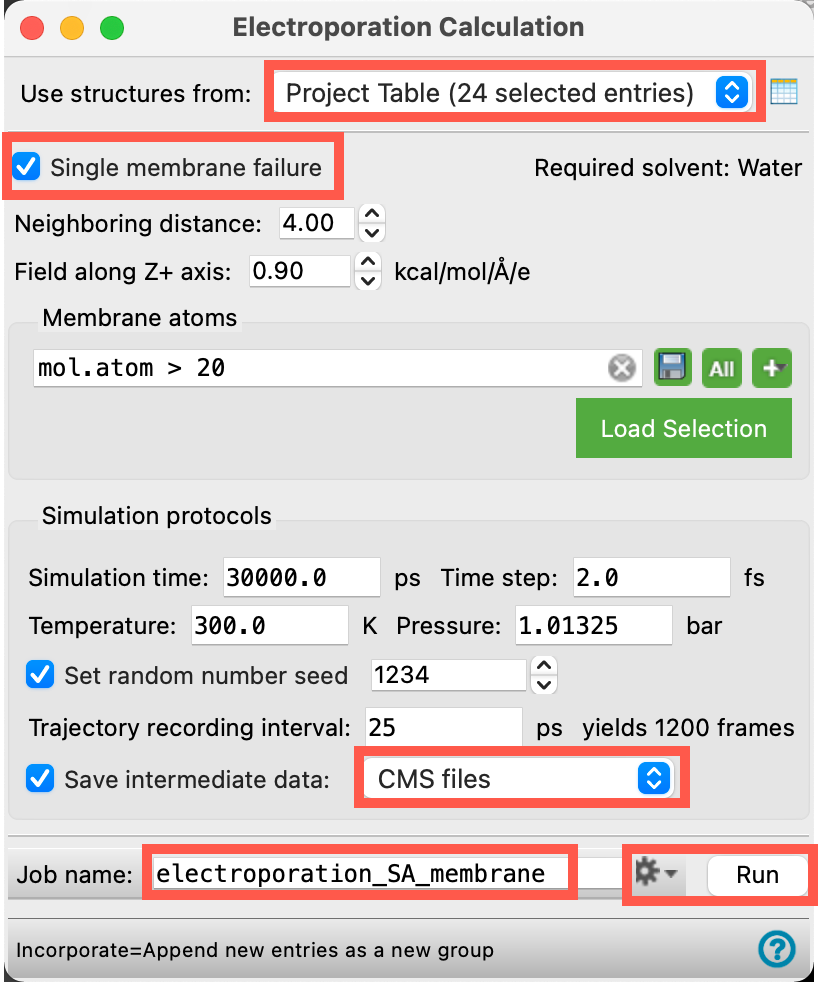
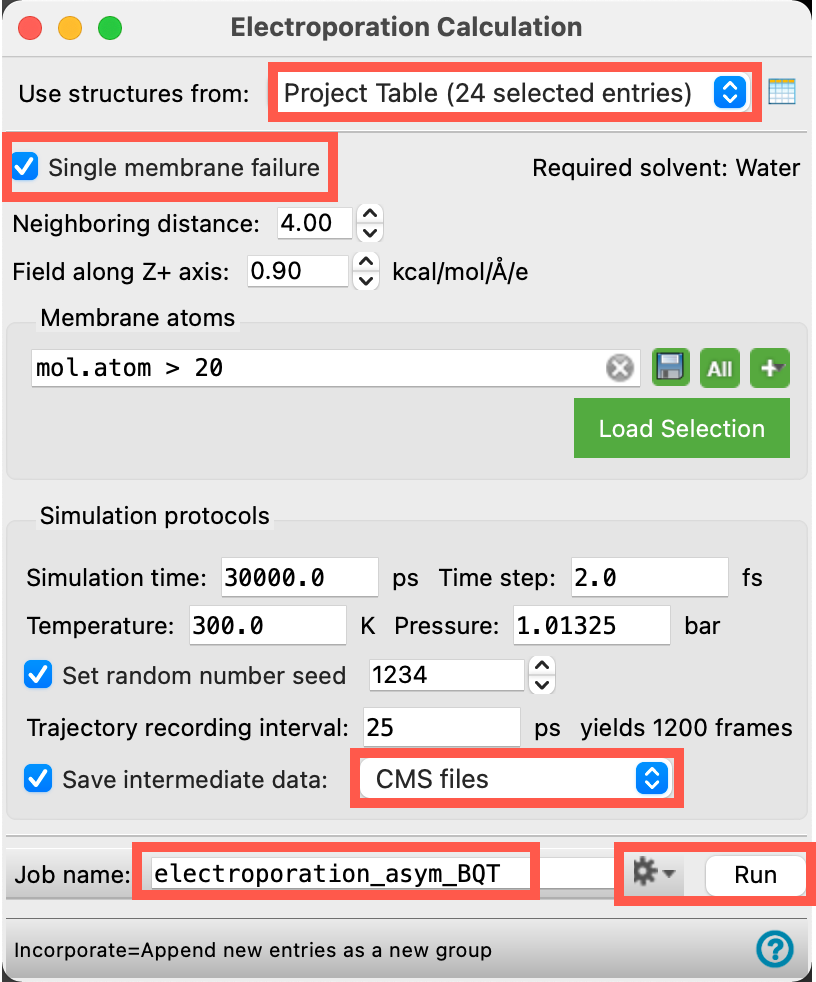
We will retain most of the defaults in the panel. Change the necessary parameters as specified below:
- Ensure Use structures from shows Project Table (24 selected entries)
- We will run our electroporation calculation on 24 replicates of the same system
- Check Single membrane failure
- We will calculate pore formation time and probability for a single membrane
- For Save intermediate data, select CMS files from the drop-down menu
- If you are interested in visualizing the trajectories for each replicate, feel free to select CMS and trajectory instead, but do so with caution as the trajectories are disk space intensive
- Change the Job name to electroporation_SA_membrane
- Adjust the job settings (
 ) as needed
) as needed
- This job requires a GPU host. The job can be completed in a day on a GPU host
- It is suggested to distribute subjobs across GPU subhosts. We will be running 24 independent MD simulations as a part of this calculation which can be sped up if running on multiple GPUs
- If you would like to run the job yourself, click Run. Otherwise, go to File > Import Structures, navigate to the provided tutorial files and Open
Section_03 > electroporation_SA_membrane > electroporation_SA_membrane-out.maegz - Close the Electroporation panel
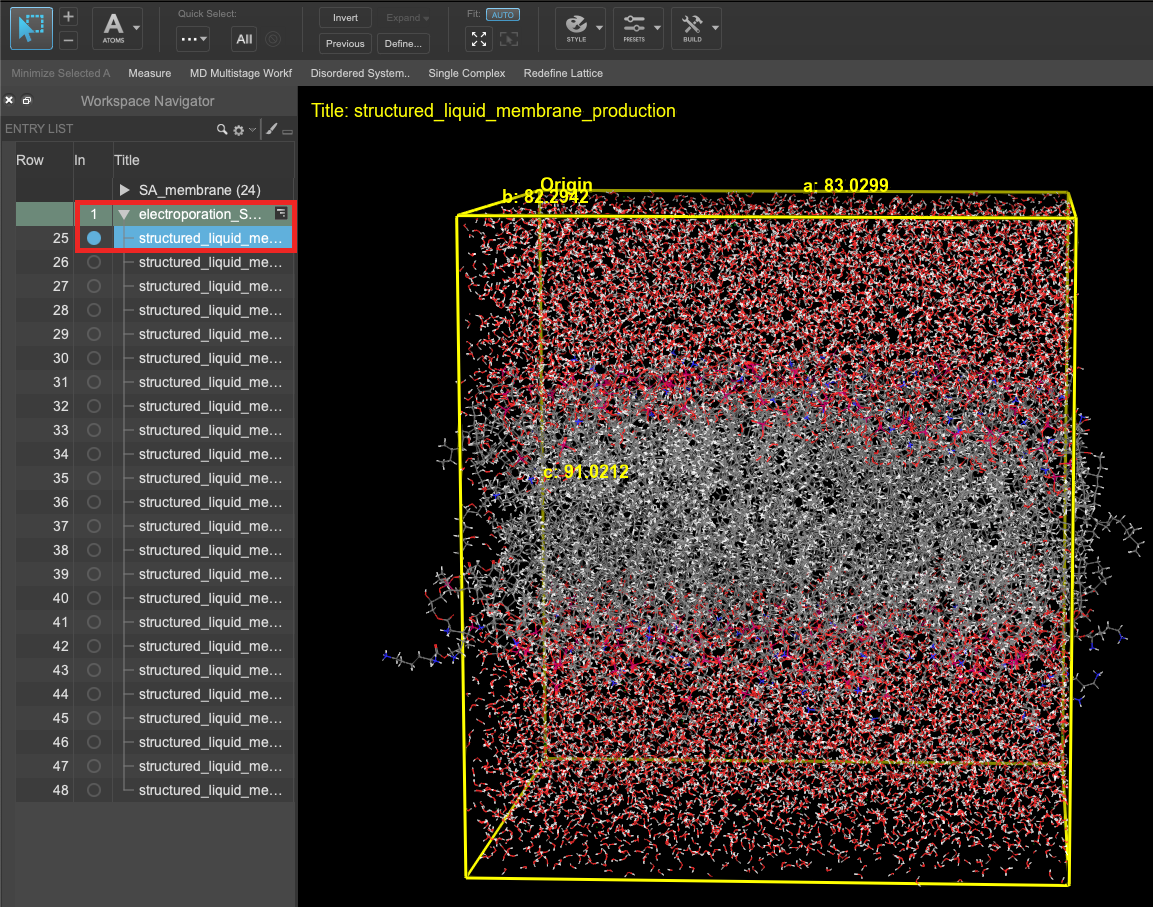
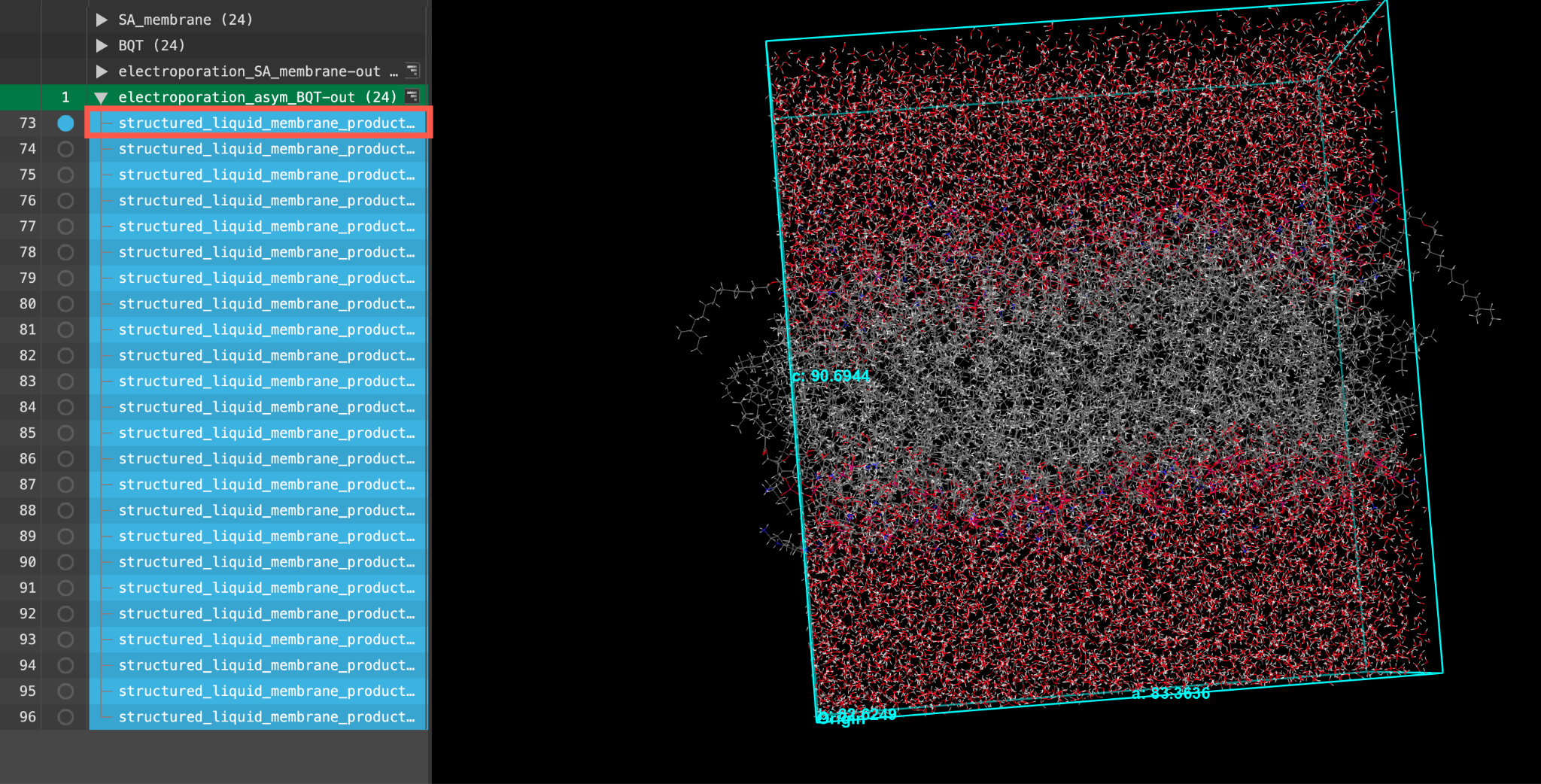
- When the job is finished or after importing, select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue the first structured_liquid_membrane_production entry in the entry list.
- Feel free to stylize and visualize the output system in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
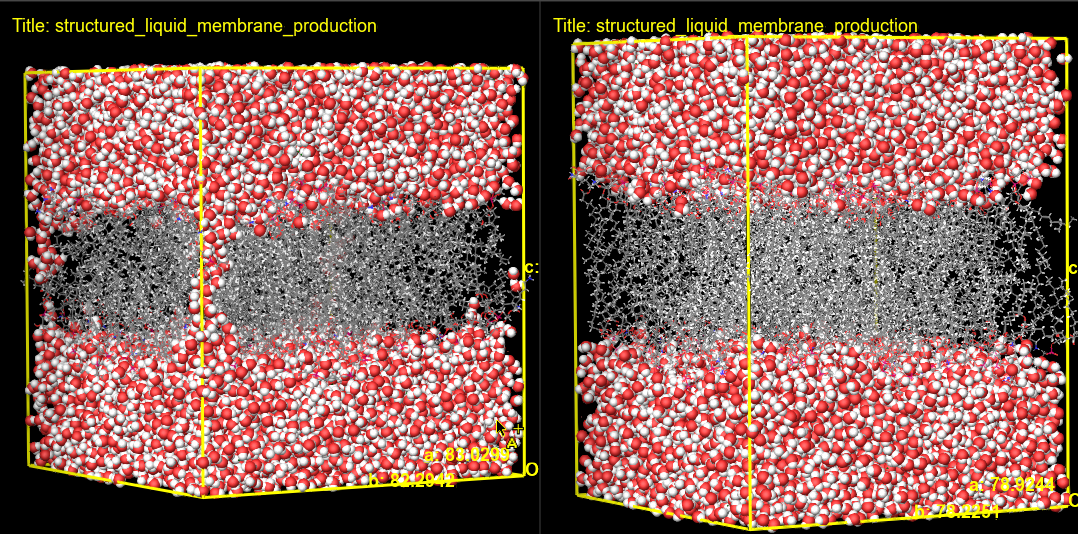
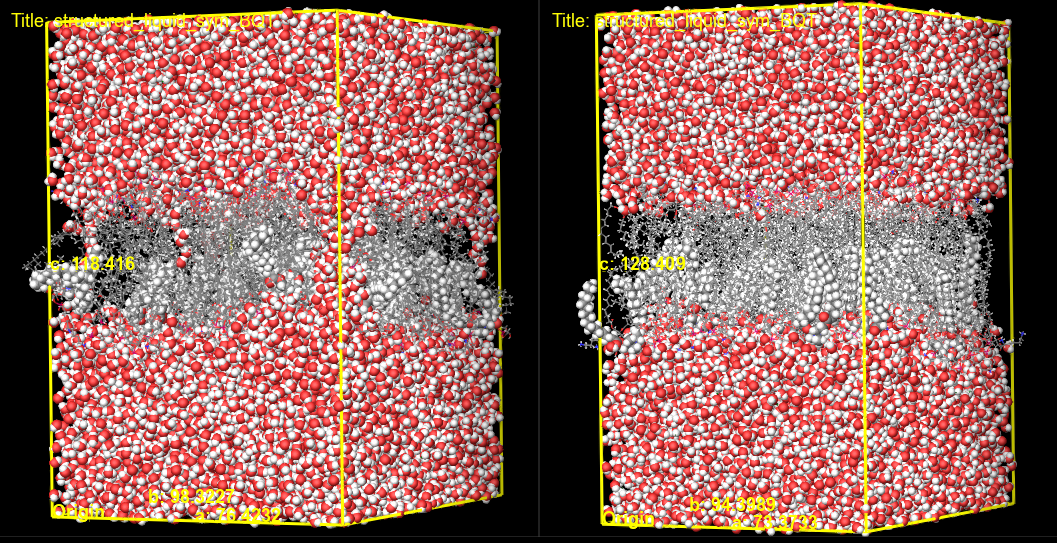
Figure 3-7. Visualizing a S. aureus membrane structure before (left) and after (right) the electroporation calculation.
- Maintain the inclusion of the output, structured_liquid_membrane_production, as instructed above and also include (Ctrl or Cmd click) the corresponding input structure from the SA_membrane entry group from the beginning of this section
- We will visualize pore formation in a single input and output structure of the electroporation calculation
- Open the Workspace Configuration Panel

- Click on Tile (
 )
)
- The two membrane structures are tiled in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Here, we can visualize the pore formation resulting from the electroporation calculation. We can most easily visualize the pores formed by changing the representation of the water molecules to CPK. In this example, we see the initial structure with water surrounding the membrane before the electroporation calculation and water molecules passing through the membrane after. If interested, feel free to visualize other replicates.
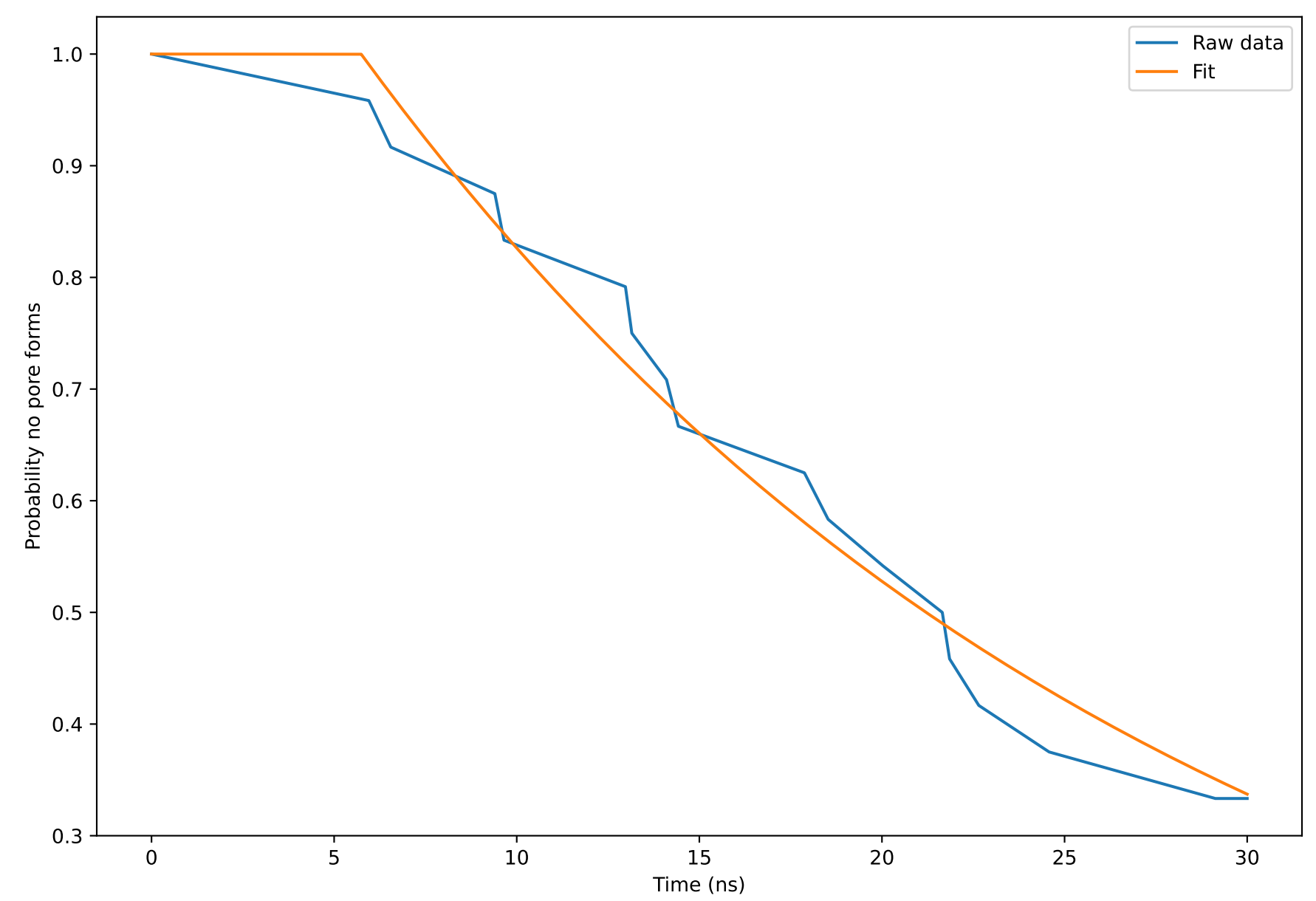
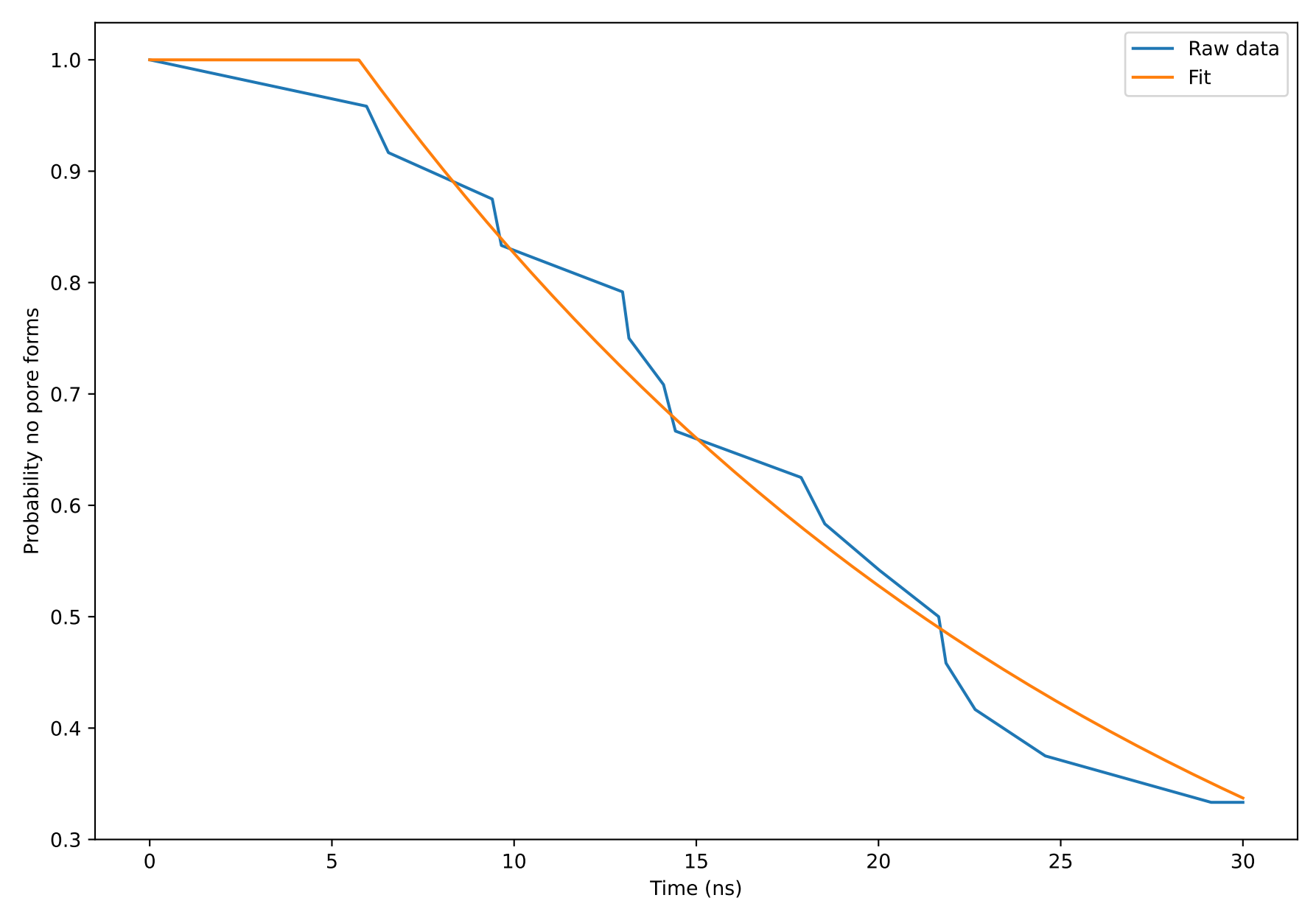
Now we will open the pore formation probability plot generated from the simulation to extract some quantitative metrics. The plot is generated by combining the pore formation data across the 24 replicates.
The probability that no pore is formed reduces with time for the S. aureus membrane. At the maximum simulation time of 30 ns, there is a 40% chance that no pore has yet formed in the system. In other words, there is a 60% chance that a pore has formed at this time. In the next section, we will compare these results to those of the asymmetrically loaded BQT membrane.
4. Electroporation of the Asymmetric BQT Membrane
In this section, we will use the Electroporation panel to study the stability of the S. aureus membrane asymmetrically loaded with BQT. The input systems should be well-relaxed MD outputs, as we have provided here. We will then visualize the output of the electroporation calculation.
For this example, our structure of interest has been built in advance and is available for importing in the next step. We will import 24 structure files (.cms) of an equilibrated cell of the S. aureus membrane with 20 BQT molecules inserted in one leaflet. These 24 structure files were generated by building four independent replicates of the membrane system, equilibrating the four systems, and then taking six equilibrated frames from each of the four resulting trajectories.
If you are interested in building and equilibrating the system yourself, feel free to do so. For practice building and equilibrating systems containing membranes, see the Calculating Surfactant Tilt and Electrostatic Potential of a Bilayer System tutorial to review the relevant tools.
- Go to File > Import Structures
- Navigate to where you downloaded the tutorial files (presumably your working directory) and choose
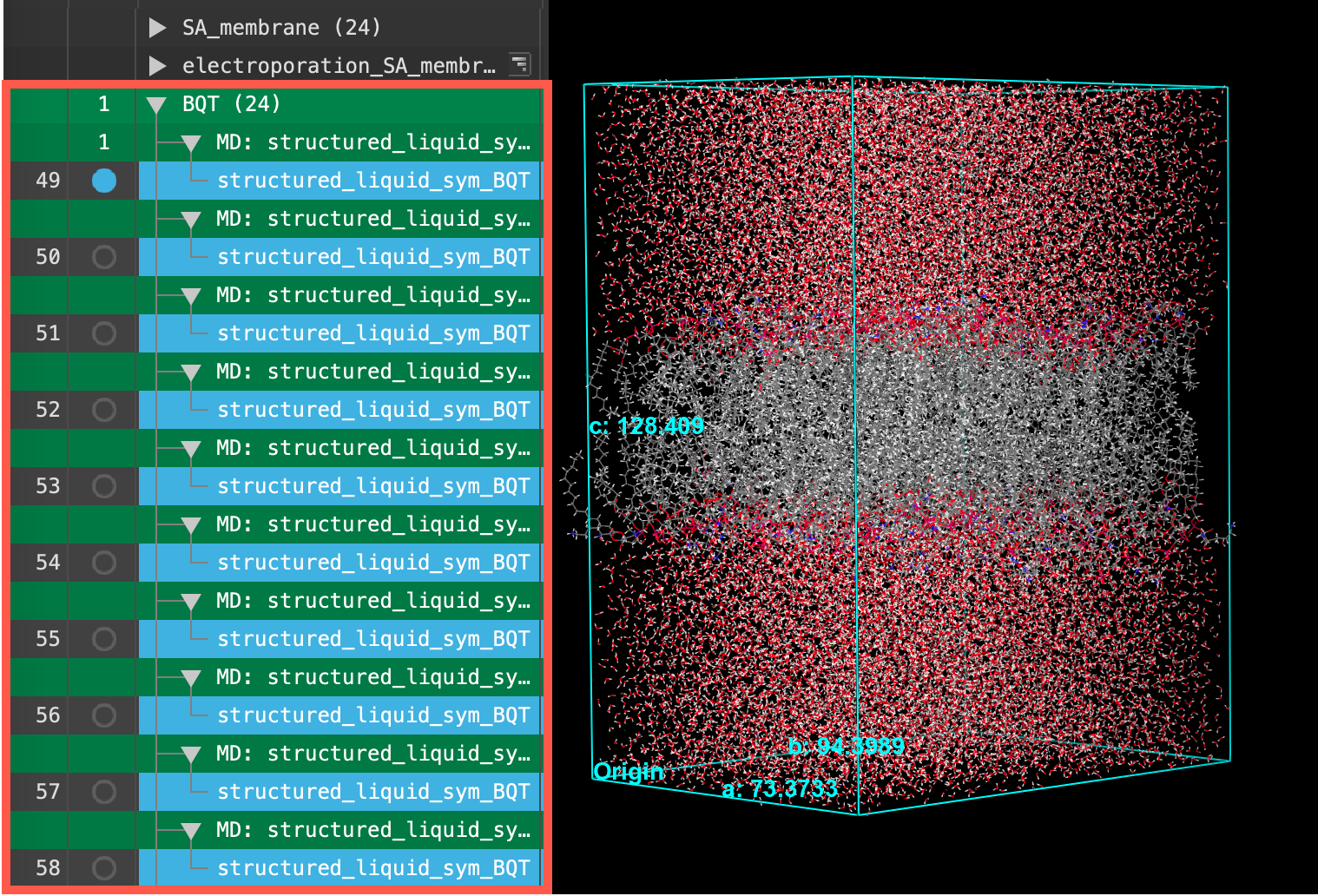
Section_04 > asym_BQT_input > structured_liquis_membrane_production_n.cms. Where n = 1 - 24. Click Open- 24 new entries are added to the entry list
- Repeat Steps 3-6 in Section 3 to create an entry group titled BQT for the 24 structures imported in the previous step
- Ensure that the BQT entry group is selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries in the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
- Go to Tasks > Materials > Classical Mechanics > Electroporation
- The Electroporation panel opens
We will retain most of the defaults in the panel. Change the necessary parameters as specified below:
- Ensure Use structures from shows Project Table (24 selected entries)
- We will run our electroporation calculation on 24 replicates of the same system
- Check Single membrane failure
- We will calculate pore formation time and probability for a single membrane
- For Save intermediate data, select CMS files from the drop-down menu
- If you are interested in visualizing the trajectories for each replicate, feel free to modify your choice here, but do so with caution as the trajectories are disk space intensive.
- Change the Job name to electroporation_asym_BQT
- Adjust the job settings (
 ) as needed
) as needed
- This job requires a GPU host. The job can be completed in ~4 hours on a GPU host
- It is suggested to distribute subjobs across GPU subhosts. We will be running 24 independent MD simulations as a part of this calculation which can be sped up if running on multiple GPUs
- If you would like to run the job yourself, click Run. Otherwise, go to File > Import Structures, navigate to the provided tutorial files and Open
Section_04 > electroporation_asym_BQT > electroporation_asym_BQT-out.maegz - Close the Electroporation panel
- When the job is finished or after importing, select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includethe entry is represented in the Workspace, the circle in the In column is blue the first structured_liquid_sym_BQT entry in the entry list.
- Feel free to stylize and visualize the output system in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Figure 4-6. Visualizing the asymmetric BQT membrane structure before (left) and after (right) the electroporation calculation.
- Maintain the inclusion of the output, structured_liquid_sym_BQT, as instructed above and also include (Ctrl or Cmd click) the corresponding input structure from the BQT entry group from the beginning of this section
- We will visualize pore formation in a single input and output structure of the electroporation calculation
- Open the Workspace Configuration Panel

- Click on Tile (
 )
)
- The two membrane structures are tiled in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Here, we can visualize the pore formation resulting from the electroporation calculation. We can most easily visualize the pores formed by changing the representation of the water molecules to CPK. Additionally in this rendering, we have set the BQT molecules to have a CPK representation. In this example, we see the initial structure with water surrounding the membrane before the electroporation calculation and water molecules passing through the membrane after. If interested, feel free to visualize other replicates.
Now we will open the pore formation probability plot generated from the simulation to extract some quantitative metrics. The plot is generated by combining the pore formation data across the 24 replicates.
The probability that no pore is formed reduces with time for the asymmetric BQT membrane. At a simulation time of 2.0 ns, there is a 0% chance that no pore has yet formed in the system (100% chance of pore formation). Recalling the results from the end of Section 3, we can draw the conclusion that the asymmetric BQT membrane ruptures significantly faster than the symmetric S. aureus membrane. This tells us that asymmetric BQT insertion makes the S. aureus membrane much less stable. This qualitatively matches what was found in the literature.
5. Conclusion and References
In this tutorial, we learned how to use the Electroporation panel to simulate pore formation in bacterial membranes by applying an electric field.
For further learning:
For introductory content, focused on navigating the Schrödinger Materials Science interface, an Introduction to Materials Science Maestro tutorial is available. Please visit the materials science training website for access to 100+ tutorials. For scientific inquiries or technical troubleshooting, submit a ticket to our Technical Support Scientists at help@schrodinger.com.
For self-paced, asynchronous, online courses in Materials Science modeling, including access to Schrödinger software, please visit the Schrödinger Online Learning portal on our website.
For some related practice, proceed to explore other relevant tutorials:
- Disordered System Building and Molecular Dynamics Multistage Workflows
- Applying Barrier Potentials for Molecular Dynamics Simulations
- Building, Equilibrating and Analyzing Amorphous Polymers
- Crosslinking Polymers
- Polymer Property Prediction
- Penetrant Loading
- Diffusion
- Evaporation
- Cluster Analysis
- Building a Semicrystalline Polymer
- Thermal Conductivity
- Building a Polymer-Polymer Interface
- Building a Carbohydrate Polymer
- Surface Tension
- Droplet Contact Analysis
- Viscosity
- Building a Coarse-Grained Surfactant Model with Martini Force Field
For further reading:
- See the help documentation on the Electroporation Calculation panel
- Bacterial Membranes Are More Perturbed by the Asymmetric Versus Symmetric Loading of Amphiphilic Molecules. DOI:110.3390/membranes12040350
- Membrane pore formation in atomistic and coarse-grained simulations. DOI:10.1016/j.bbamem.2015.12.031
- Reversibility of membrane permeabilization upon pulsed electric field treatment in Lactobacillus plantarum WCFS1. DOI:10.1038/s41598-019-56299-w
6. Glossary of Terms
Entry List - a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
Included - the entry is represented in the Workspace, the circle in the In column is blue
Project Table - displays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data
Recent actions - This is a list of your recent actions, which you can use to reopen a panel, displayed below the Browse row. (Right-click to delete.)
Scratch Project - a temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project
Selected - (1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Working Directory - the location where files are saved
Workspace - the 3D display area in the center of the main window, where molecular structures are displayed