Viscosity
Tutorial Created with Software Release: 2026-1
Topics: Consumer Packaged Goods , Organic Electronics , Pharmaceutical Formulations , Polymeric Materials
Methodology: All-Atom Molecular Dynamics
Products Used: Desmond , MS Maestro , MS Transport
|
15 GB |
This tutorial is written for use with a 3-button mouse with a scroll wheel.
Words found in the Glossary of Terms are shown like this: Workspacethe 3D display area in the center of the main window, where molecular structures are displayed
Abstract:
In this tutorial, we will calculate the shear viscosities of a series of alkanes using Materials Science (MS) Maestro. Computationally determined viscosities and densities from the workflow will be compared to known experimental values.
Tutorial Content
1. Introduction to Viscosity
The Green–Kubo integral of the pressure tensor autocorrelation function can be used to calculate the shear viscosity of a system that has been equilibrated with molecular dynamics. To use the Green–Kubo relationship for viscosity, the auto-correlation function of the off-diagonal elements of the pressure tensor (Pɑβ) is calculated and integrated to give shear viscosity as a function of simulation time differences (τ). The final viscosity value is obtained by curve fitting or averaging over a specified range of the integral.
Green–Kubo relation
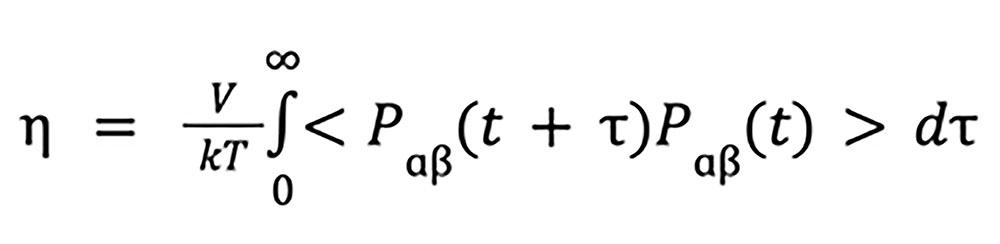
where
 is the shear viscosity, V is the system volume, k is the Boltzmann constant, T is temperature, Pɑβ is the element ɑβ of the pressure tensor where the brackets indicate the ensemble average over xy, yz and xz.
is the shear viscosity, V is the system volume, k is the Boltzmann constant, T is temperature, Pɑβ is the element ɑβ of the pressure tensor where the brackets indicate the ensemble average over xy, yz and xz.
The integral of <P (t + τ) P(t)> over τ from 0 to infinity converges to a constant. This presents a practical issue, as we cannot integrate to infinity due to cost and accuracy. Thus, here we fit using an analytical function, which converges to a value at infinite τ. The form of the analytical function used here is a double exponential (see References), which is used to obtain the viscosity as a function of τ, which in the limit of τ=infinity gives a value for the viscosity at infinite τ.
For calculation of accurate viscosity values, sampling is critical. Thus, the workflow available for calculating shear viscosity in Materials Science (MS) Maestro is optimized to maximize sampling with efficient use of GPU.
For a complete overview of viscosity calculations in Materials Science Maestro, visit the help documentation. Note that the workflow for calculating viscosity can be applied to mixed solvent systems in addition to homogeneous systems.
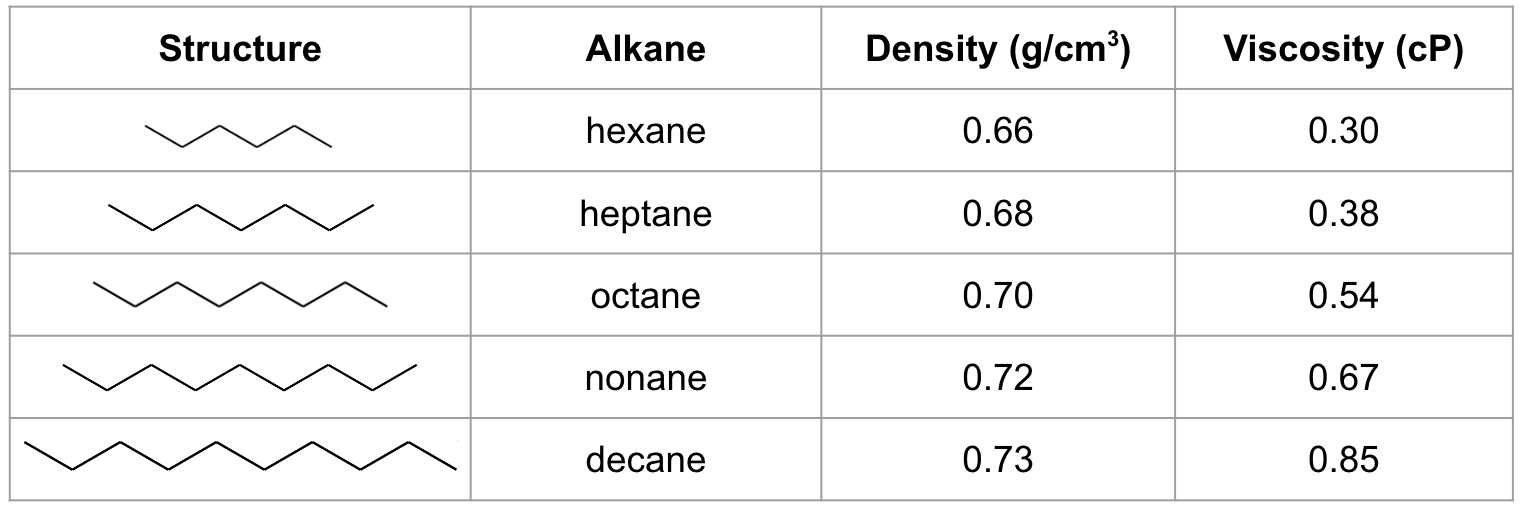
In this tutorial, we will consider the homologous series of alkanes from hexane to decane, which are all liquid hydrocarbons under standard conditions. We will determine their viscosities using a workflow in Materials Science Maestro and compare to experimental values.
Experimental densities and viscosities of the alkanes of interest
2. Creating Projects and Importing Structures
At the start of the session, change the file path to your chosen Working Directorythe location where files are saved in MS Maestro to make file navigation easier. Each session in MS Maestro begins with a default Scratch Projecta temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project, which is not saved. A MS Maestro project stores all your data and has a .prj extension. A project may contain numerous entries corresponding to imported structures, as well as the output of modeling-related tasks. Once a project is saved, the project is automatically saved each time a change is made.
Structures can be built in MS Maestro or can be imported using File > Import Structures (or drag-and-dropped), and are added to the Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data. The Entry Lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion is located to the left of the Workspacethe 3D display area in the center of the main window, where molecular structures are displayed. The Project Tabledisplays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data can be accessed by Ctrl+T (Cmd+T) or Window > Project Table if you would like to see an expanded view of your project data.
- Double-click the Materials Science icon
- (No icon? See Starting Maestro)
- Go to File > Change Working Directory
- Find your directory, and click Choose
- Pre-generated input and results files are included for running jobs or examining output. Download the zip file here: schrodinger.com/sites/default/files/s3/release/current/Tutorials/zip/viscosity.zip
- After downloading the zip file, unzip the contents in your Working Directorythe location where files are saved for ease of access throughout the tutorial
- Go to File > Save Project As
- Change the File name to viscosity_tutorial, click Save
- The project is now named
viscosity_tutorial.prj
- The project is now named
3. Sketching the Initial Alkanes
We will begin by drawing the alkane inputs using the 2D Sketcher. To skip manually drawing, simply import alkanemolecules.mae and proceed to Step 13.
- From the main menu, go to Edit > 2D Sketcher
- The 2D Sketcher panel opens
For a complete overview of the sketcher panel, see the 2D Sketcher Panel documentation.
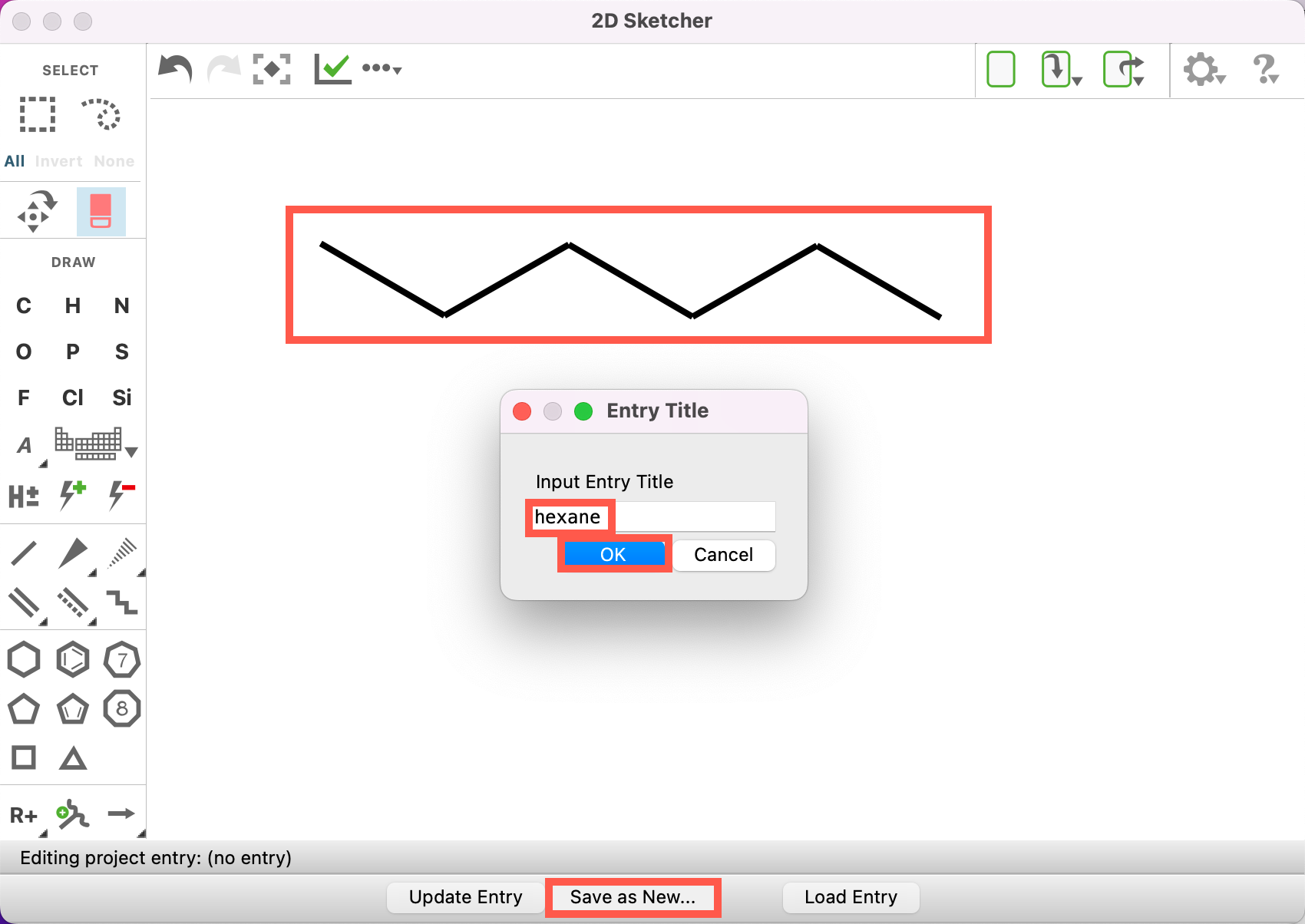
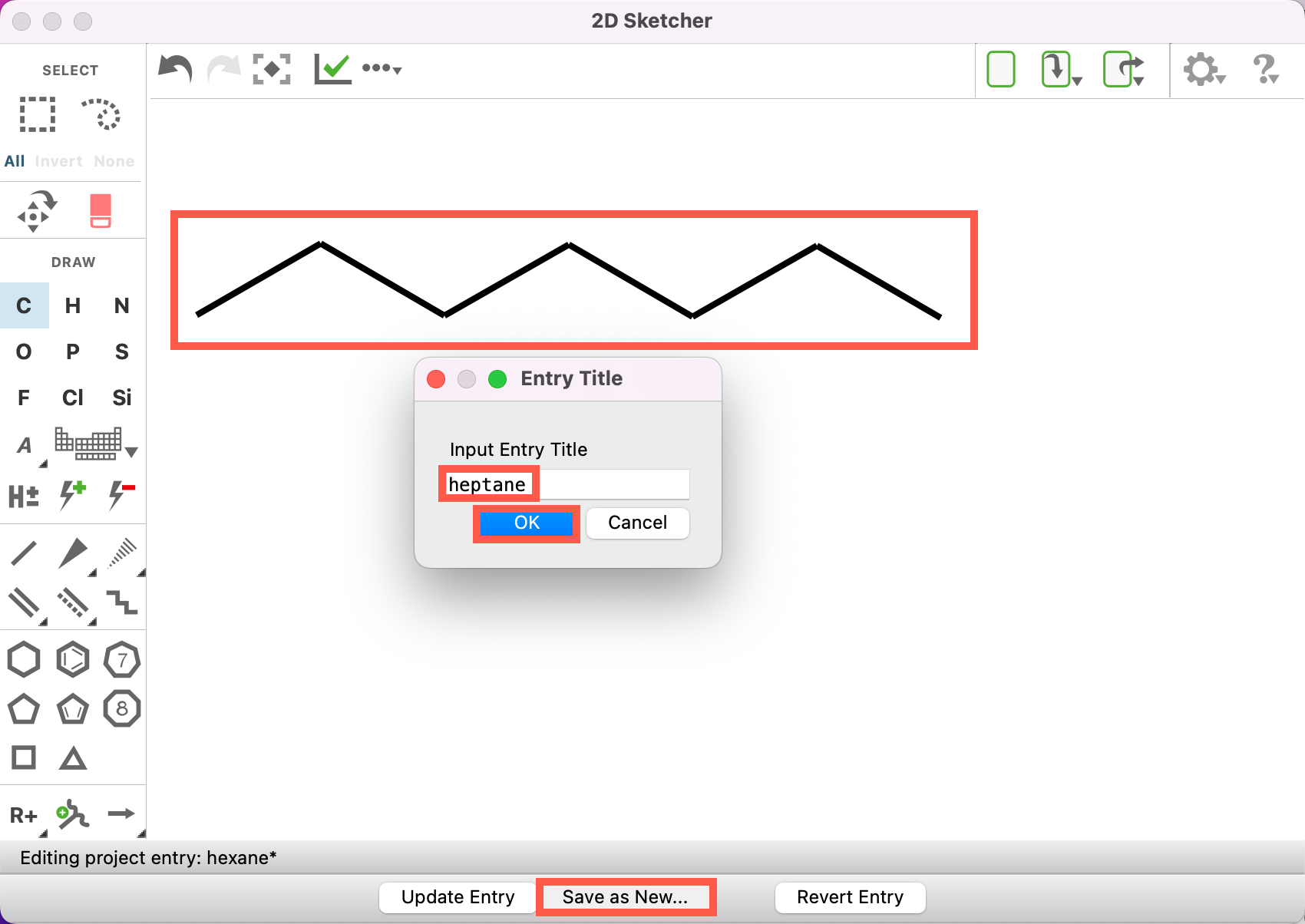
- Draw a hexane molecule (C6H14)
- Click Save as New
- For Input Entry Title write hexane
- Click OK
- Behind the 2D Sketcher in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed and entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion, the hexane molecule has been added
- Keep the 2D Sketcher open and add one carbon to the chain to draw heptane
- Click Save as New
- Be sure to not click Update Entry, as doing so will overwrite the hexane entry
- For Input Entry Title write heptane
- Click OK
- Behind the 2D Sketcher in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed and entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion, the heptane molecule has been added
- Repeat Steps 6-9 for octane, nonane and decane, such that you have created 5 new entries
- After clicking OK for decane, close the 2D Sketcher
- If necessary, reorder your entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion by clicking and dragging the entries
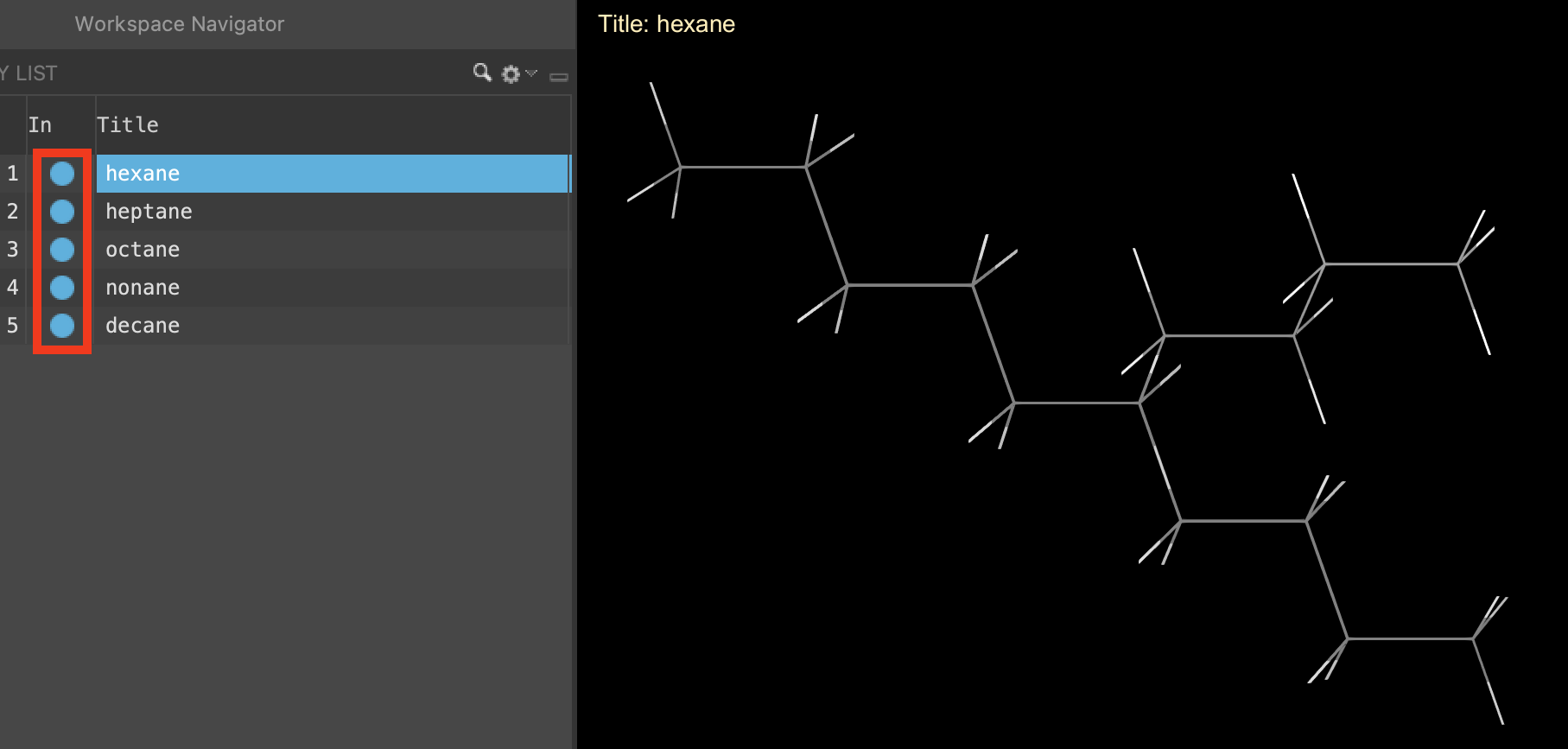
- To visualize all of the molecules in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed simultaneously, includethe entry is represented in the Workspace, the circle in the In column is blue all 5 entries (hold Shift + fill the Includethe entry is represented in the Workspace, the circle in the In column is blue circle for all five molecules)
- The molecules appear stacked in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed
4. Building the Periodic Amorphous Cell
In preparation for running molecular dynamics (MD) simulations, we will now prepare periodic amorphous cells for each molecule using the Disordered System Builder panel. For a more in depth explanation of building disordered systems, see the Disordered System Building and Molecular Dynamics Multistage Workflows tutorial.
- Select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries all five entries from the entry list a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion(Shift + Click on all 5 rows)
- Go to Tasks > Materials > Structure Builders > Disordered System
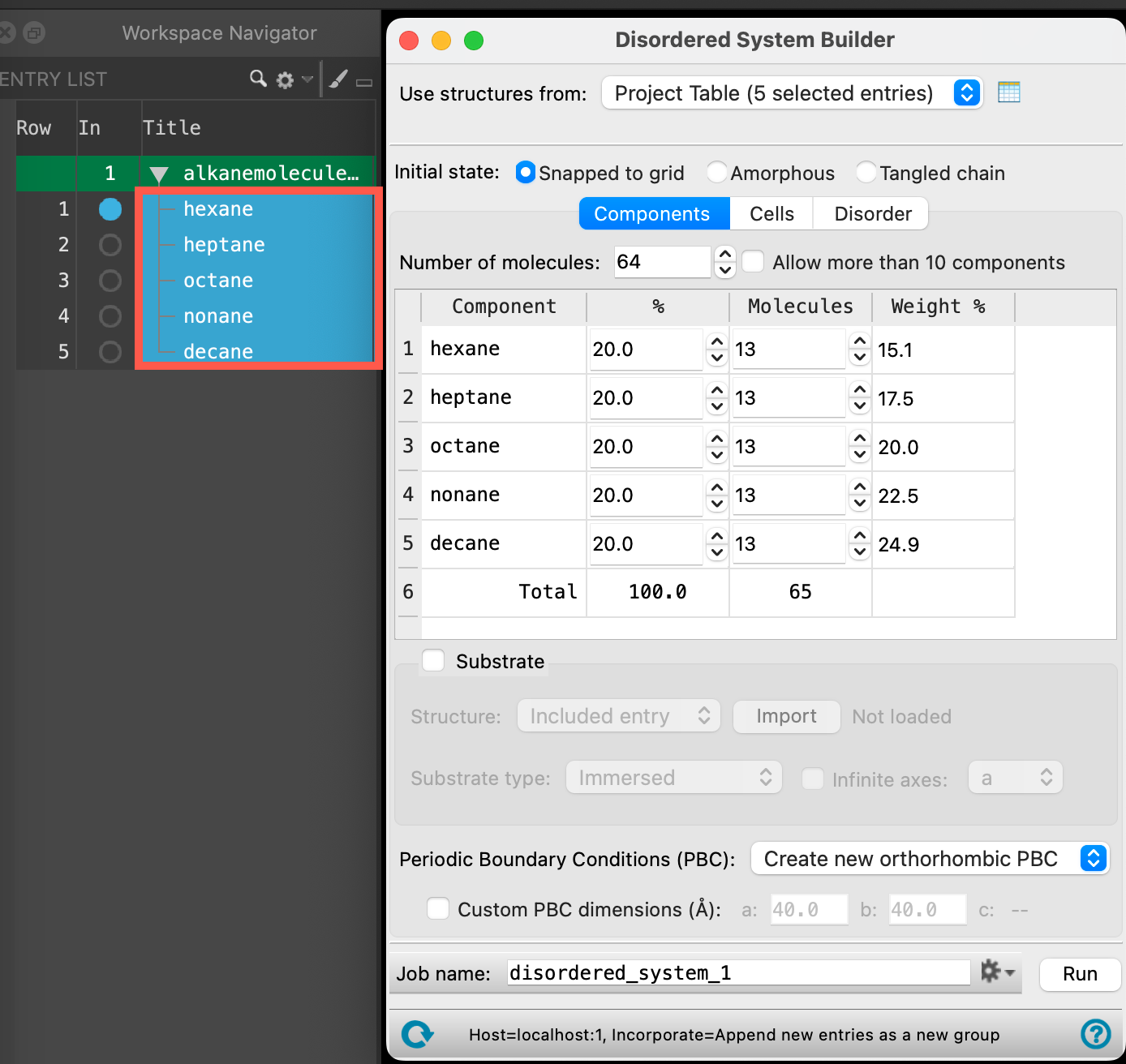
- The Disordered System Builder panel opens
The Disordered System Builder can be used to create a heterogeneous cell containing many components. Here, we wish to prepare five homogeneous cells, one for each component. To do so:
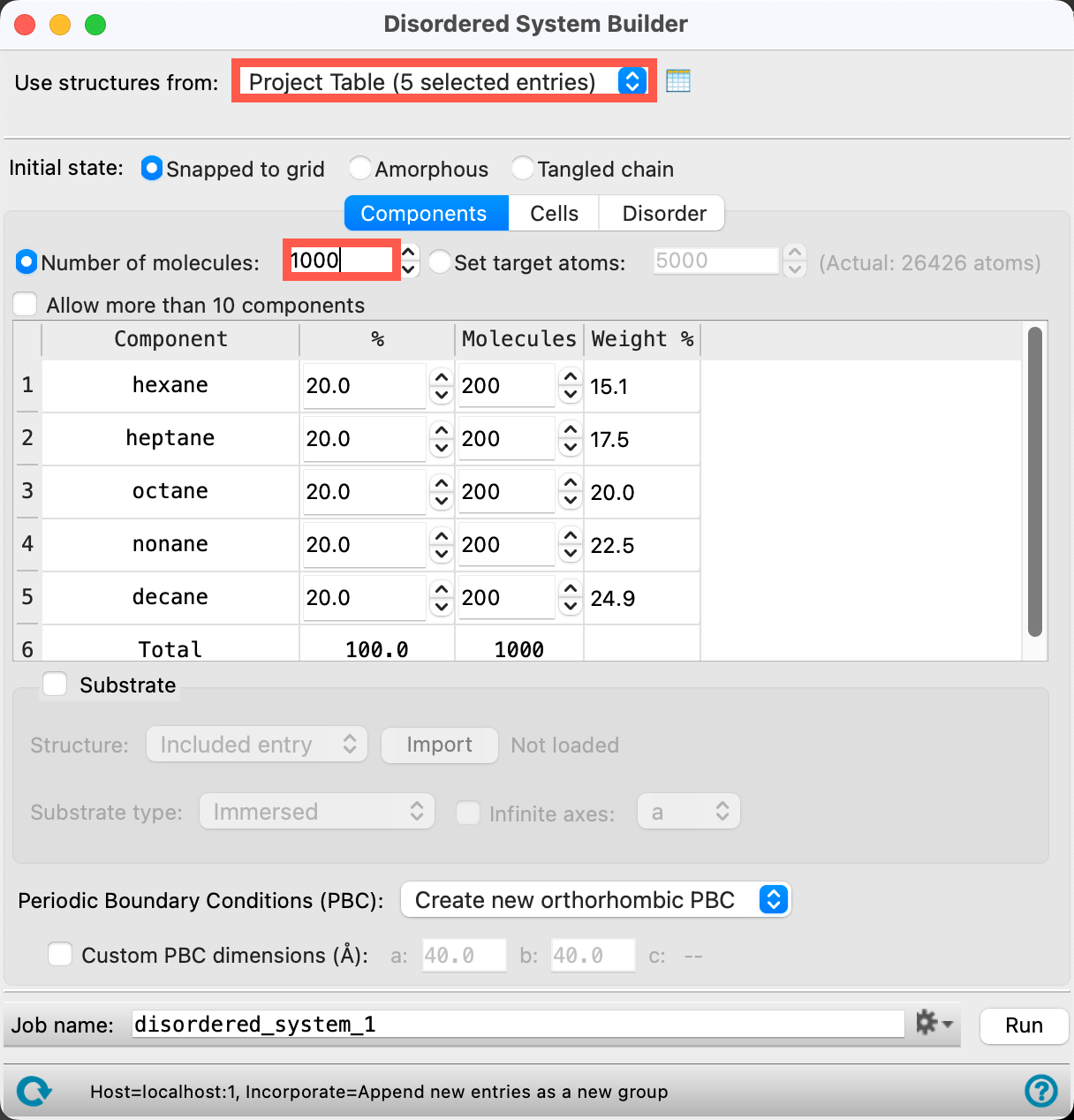
- Ensure that for Use structures from, Project Table (5 selected entries) is chosen from the dropdown
- You should see all five molecules listed as components in the panel
- For Number of Molecules, input 1000
- This will be the total number of molecules in each cell
- You can ignore the mixture in the table below, we are going to obviate that specification momentarily
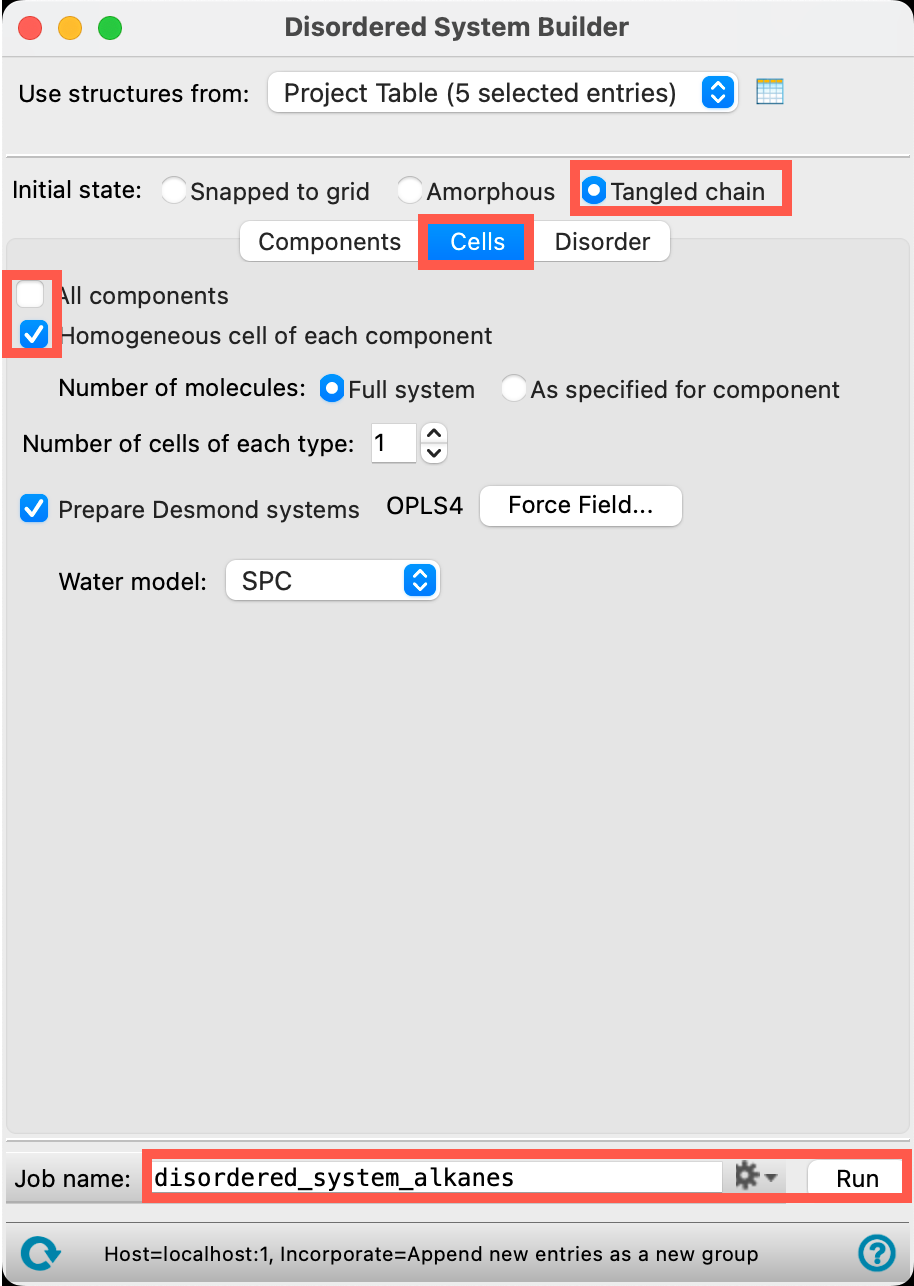
- Go to the Cells tab
- Uncheck All components
- This avoids the construction of the mixed cell
- Check Homogeneous cell of each component
- This instructs the builder to prepare the homogeneous cells instead
- For Initial state, select Tangled chain
- Change the Job name to disordered_system_alkanes
- This job takes ~10-20 minutes on a CPU Host. If you would like to run the job yourself, adjust your Job settings as needed and click Run.
If you would prefer to proceed with pre-generated structures, close the Disordered System Builder and import the amorphous cells via File > Import Structures. Navigate to where you downloaded the tutorial files and choose the Section_04 > disordered_system_alkanes_componentN_system-out.cms file where N=1-5.
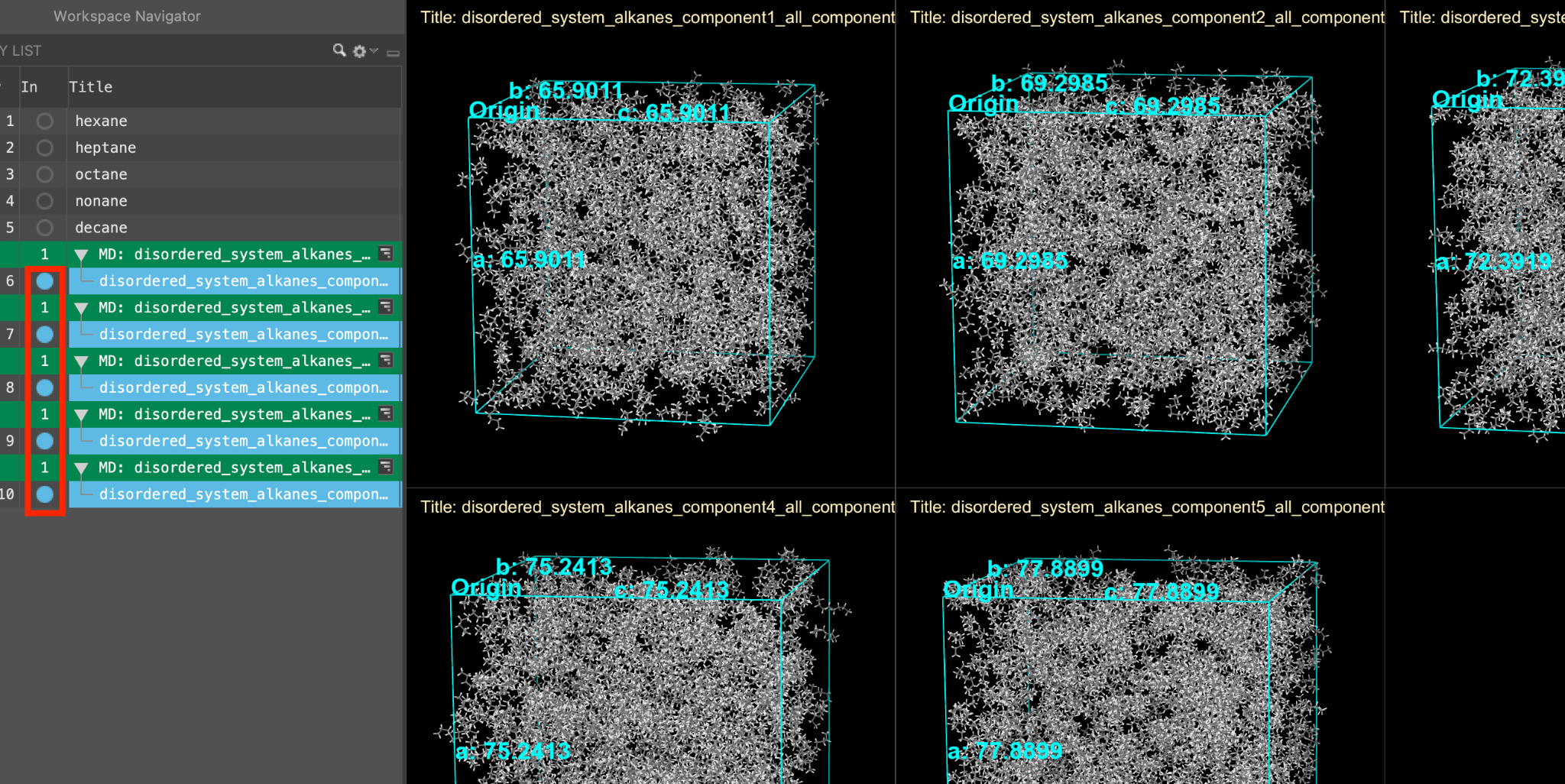
When the job is complete (or after importing), five new entry groups are added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion.
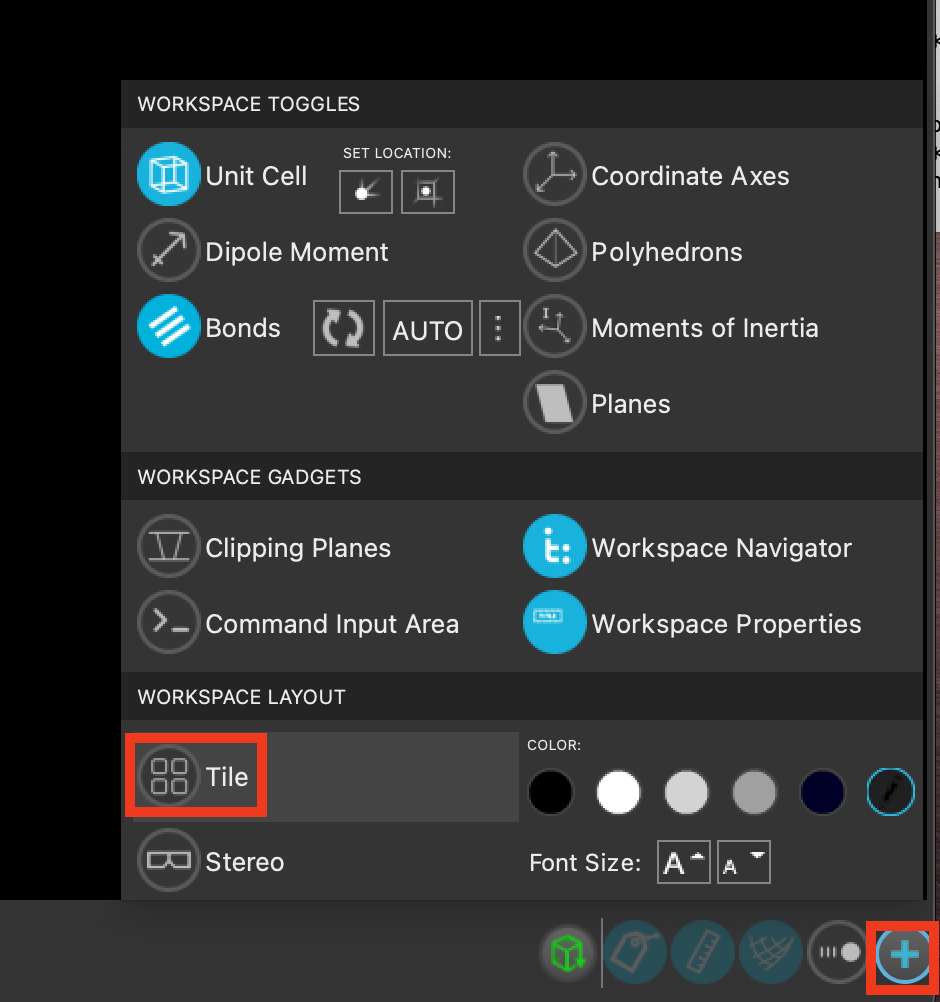
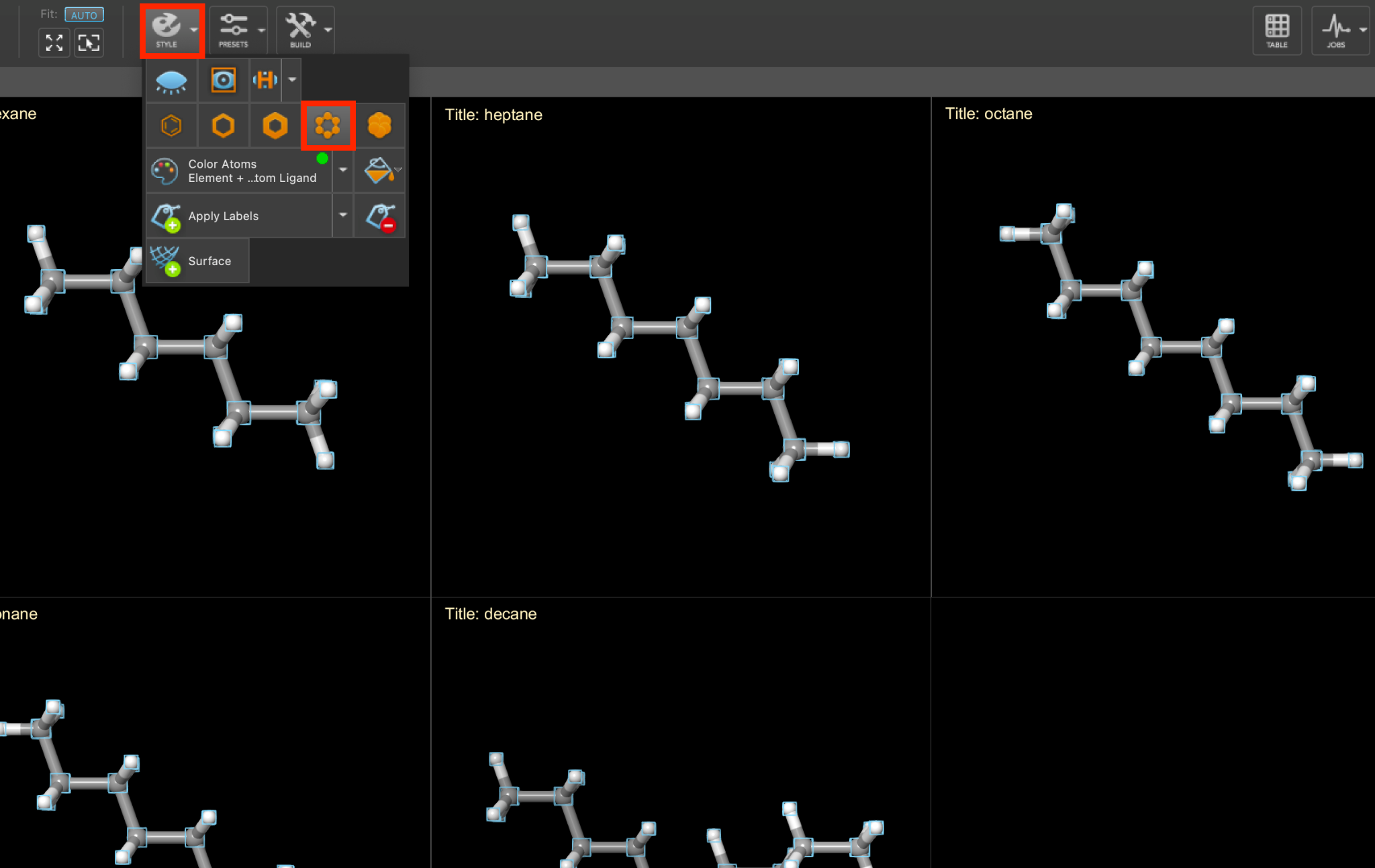
Optionally, as before, we can view all of the systems simultaneously by including the five outputs and tiling them in the workspacethe 3D display area in the center of the main window, where molecular structures are displayed.
Feel free to stylize as well. Herein, we will use ball-and-stick representation and element coloring.
Note: In this system, OPLS force fields were used in the system creation but machine Learning Force Fields (MLFF) are available as an alternative to OPLS force fields in the preparation process. Additional information regarding MLFF can be found in the help documentation or on our website.
5. Equilibrating the Amorphous Cells
The initially built periodic structures need to be brought to an experimental density at equilibrium before we can proceed with the viscosity calculations. We will bring the structures to equilibrium at 300 K and 1 atm using the MD Multistage Workflow panel. Note that the viscosity calculations do not include any equilibration steps, so this step is absolutely necessary in the workflow. For a more in depth explanation of building disordered systems, see the Disordered System Building and Molecular Dynamics Multistage Workflows tutorial.
- Select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries all five new entries from the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion (Shift + Click on all 5 rows)
- Go to Tasks > Materials > Classical Mechanics > MD Simulations > MD Multistage Workflow
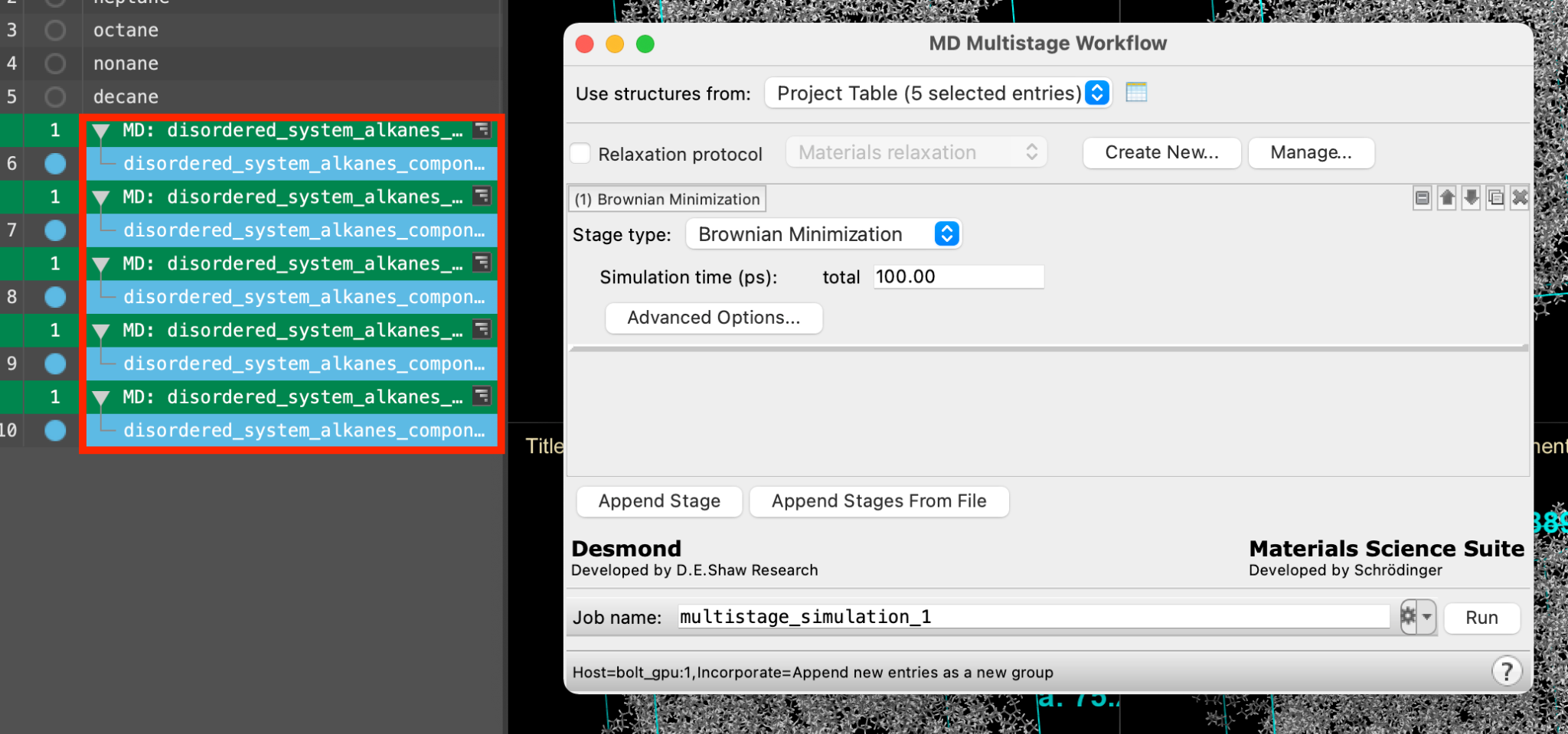
- The MD Multistage Workflow panel opens
Note: Alternatively, you can directly access the MD Multistage Workflow using the Workflow Action Menu (WAM) button ( ) next to each output from the Disordered System Builder
) next to each output from the Disordered System Builder
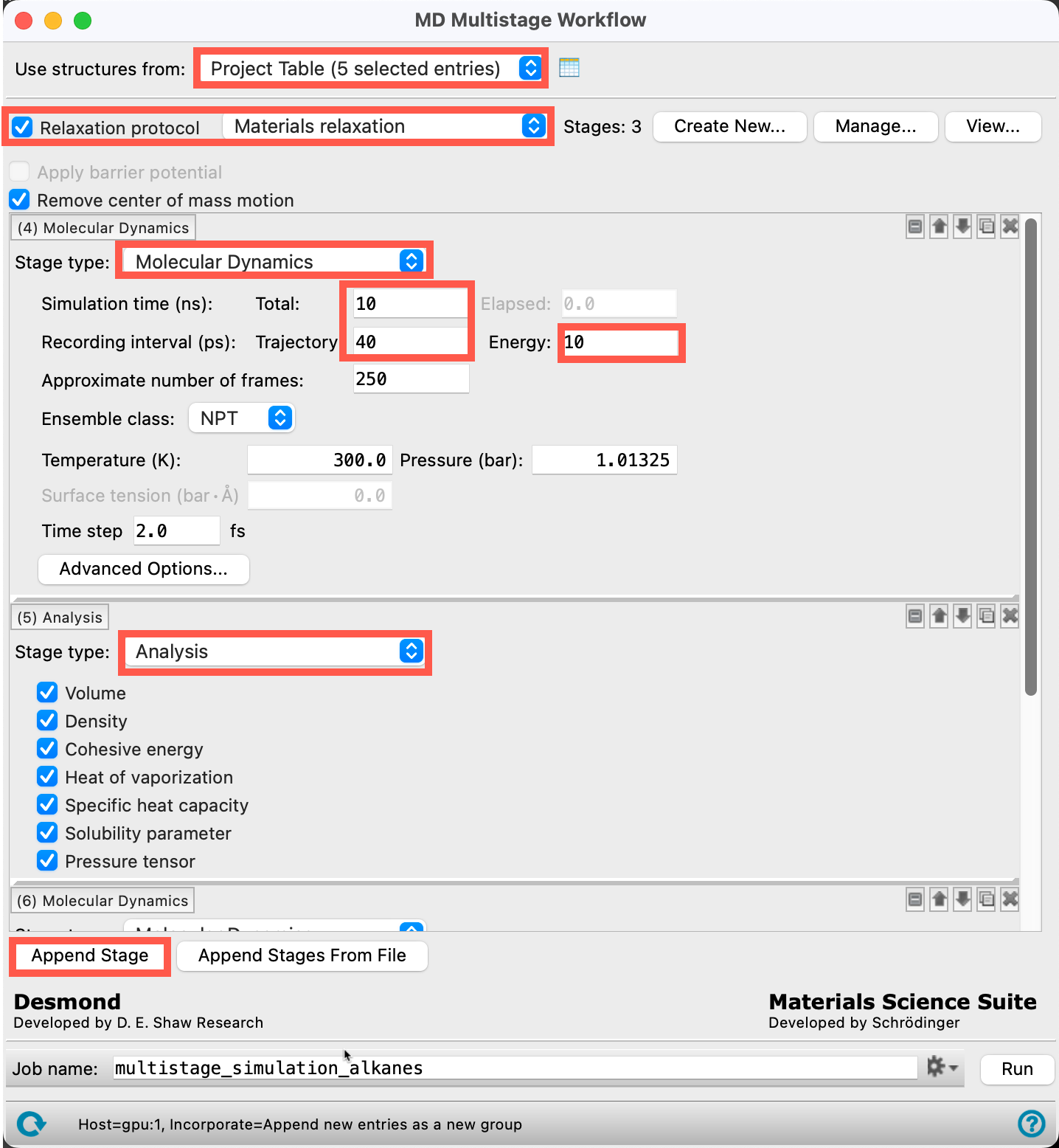
- Ensure that Use structures from directs to the 5 entries from Section 4
- Check the box for Relaxation protocol, with Materials Relaxation selected from the dropdown menu
- For the 4th stage, change the Stage type to Molecular Dynamics with the following parameters:
- Simulation time Total: 10 ns
- Recording interval Trajectory: 40 ps
- Recording interval Energy: 10 ps
- Add a 5th stage by clicking Append Stage, and change this stage to Analysis
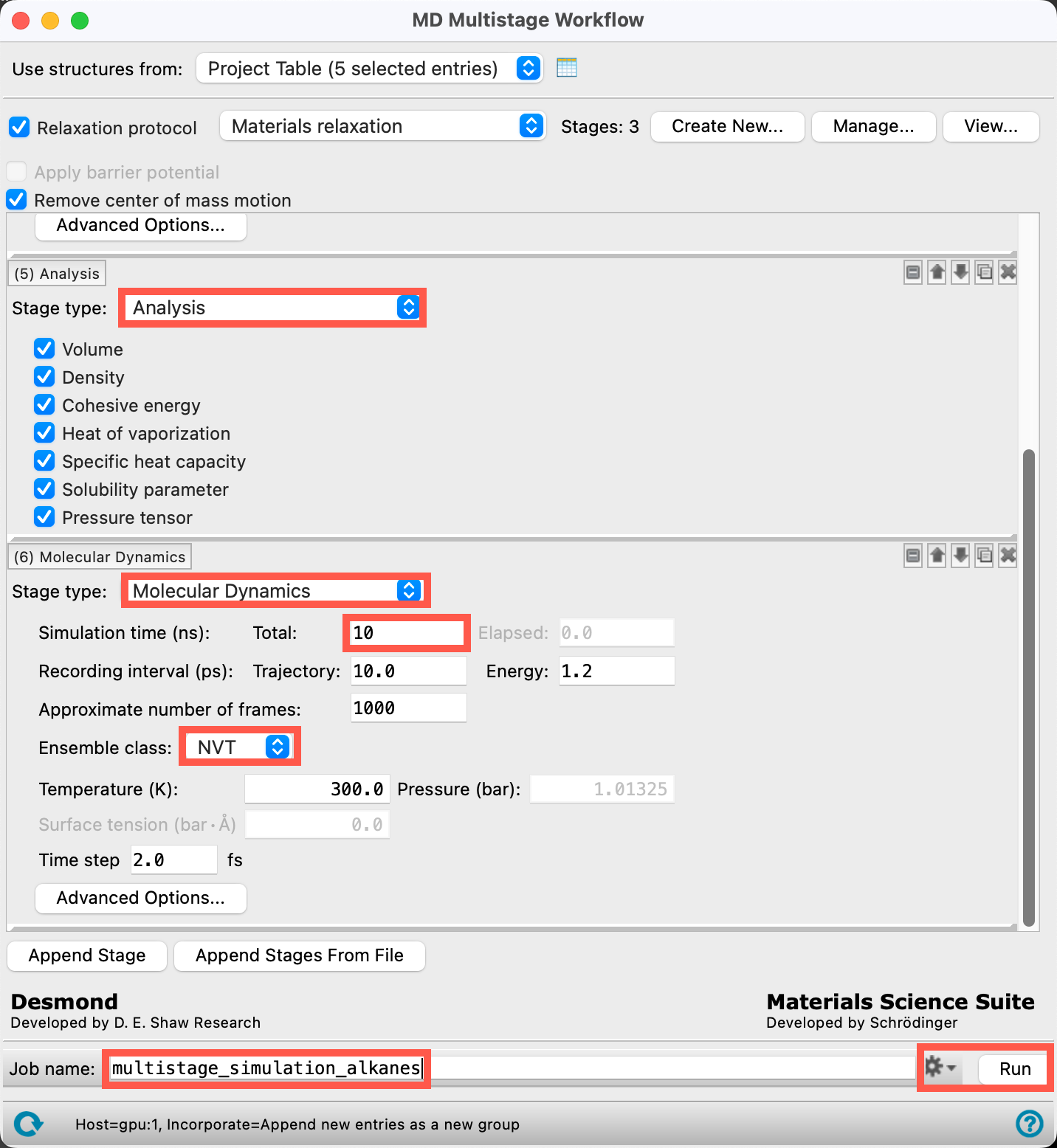
- Add a 6th stage by clicking Append Stage, and change this stage to Average Cell
- Add a 7th stage by clicking Append Stage, and change this stage to Molecular Dynamics with the following parameters:
- Simulation time Total: 10 ns
- Ensemble class: NVT
- This additional MD step is in line with best practices for equilibrating a system
- Change the Job name to multistage_simulation_alkanes
This job takes several hours on a GPU Host. If you would like to run the job yourself, adjust your Job settings as needed and click Run
If you would prefer to proceed with pre-generated structures, you can import the amorphous cells via File > Import Structures. Navigate to where you downloaded the tutorial files and choose the Section_05 > multistage_simulation_alkanes_00N > multistage_simulation_alkanes_00N-out.cms files where N=1-5.
MD Simulations have a number of files associated with the job, for a full description of each file type see the help documentation on Desmond Files
- Close the MD Multistage Workflow panel
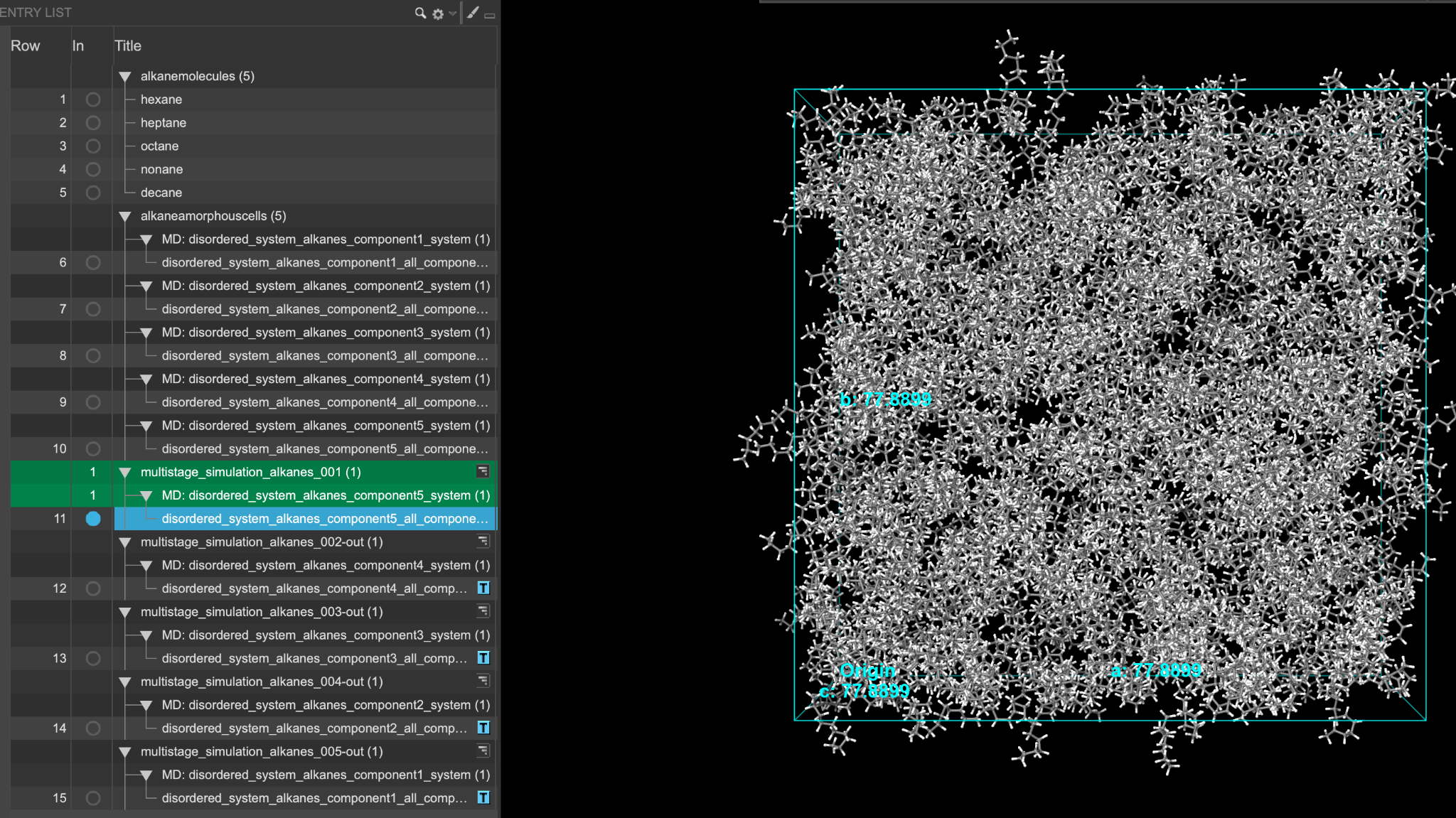
When the job is complete (or after importing), five new entry groups are added to the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion. If you ran the job, you can view the trajectories using the trajectory viewer ( ). If you imported the results, the trajectories are not provided.
). If you imported the results, the trajectories are not provided.
As before, feel free to stylize as you wish
We want to check that each system is at equilibrium, which we can assess by analyzing the densities
- With the output for hexane (the disordered_system_alkanes_component1_all_components_amorphous entry in the new MD: disordered_system_alkanes_component1_system entry group) selected(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries and includedthe entry is represented in the Workspace, the circle in the In column is blue, go to Tasks > Materials > Classical Mechanics > Trajectory Analysis > MS MD Trajectory Analysis
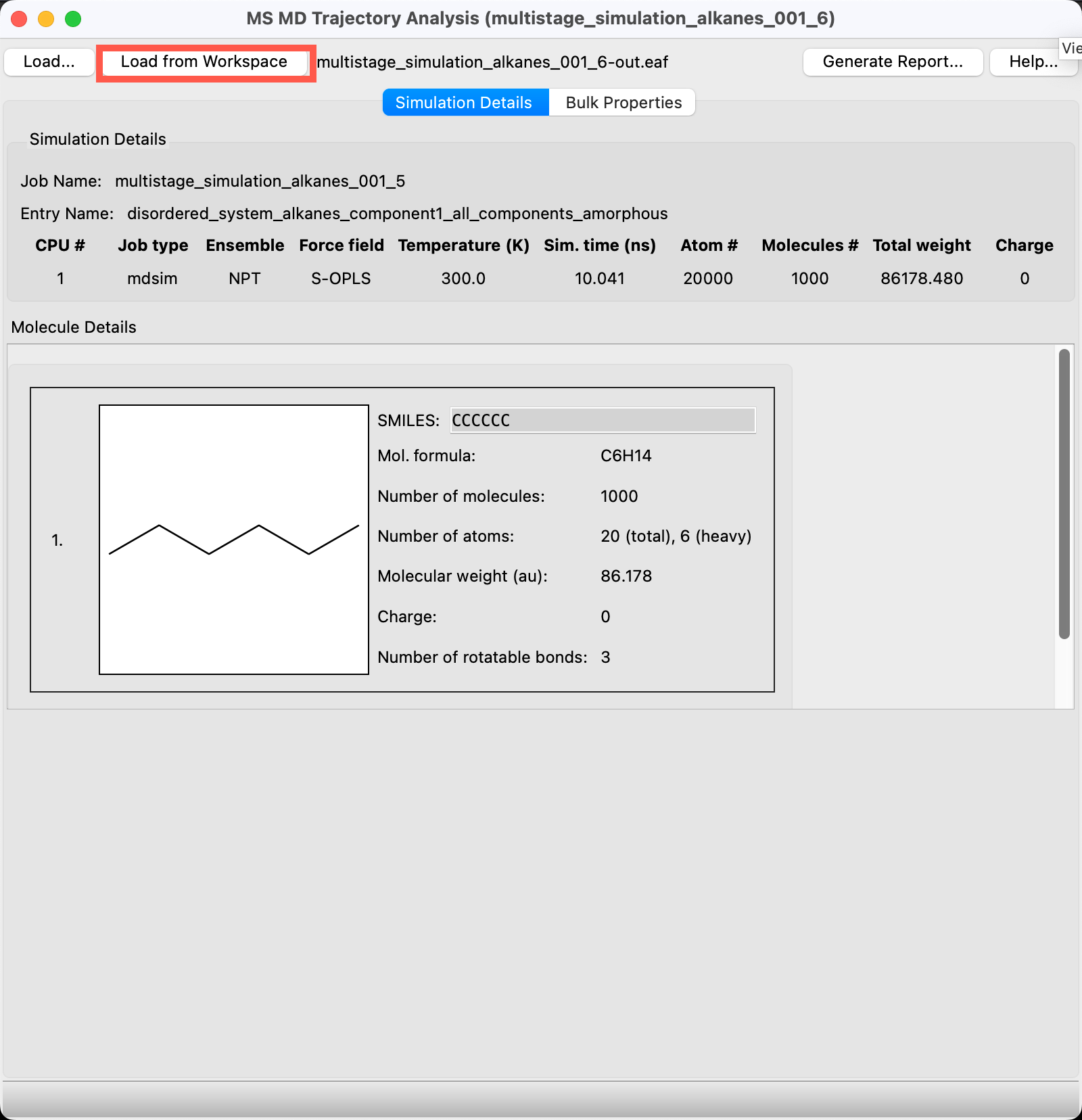
- The MS MD Trajectory Analysis panel opens
- If you ran the job, click Load from Workspace
- If you imported the results, you will be prompted to Select Trajectory File. You can use the provided
multistage_simulation_alkanes_00N-out.cmsfiles which are associated with the different alkanes - The Simulation Details are populated with information about the NPT stage MD run
- If you imported the results, you will be prompted to Select Trajectory File. You can use the provided
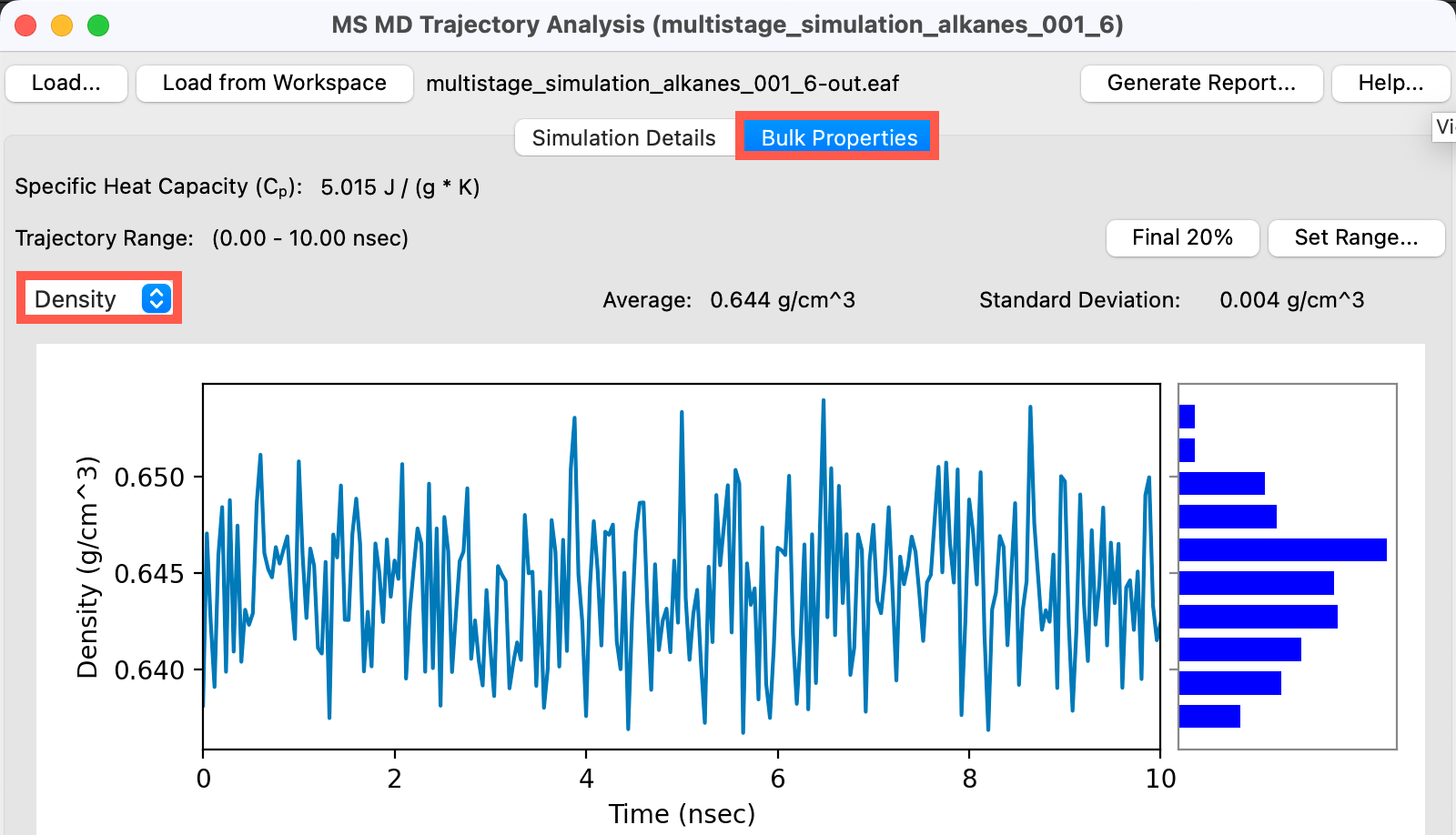
- Go to the Bulk Properties tab
- Change the first property to Density from the dropdown
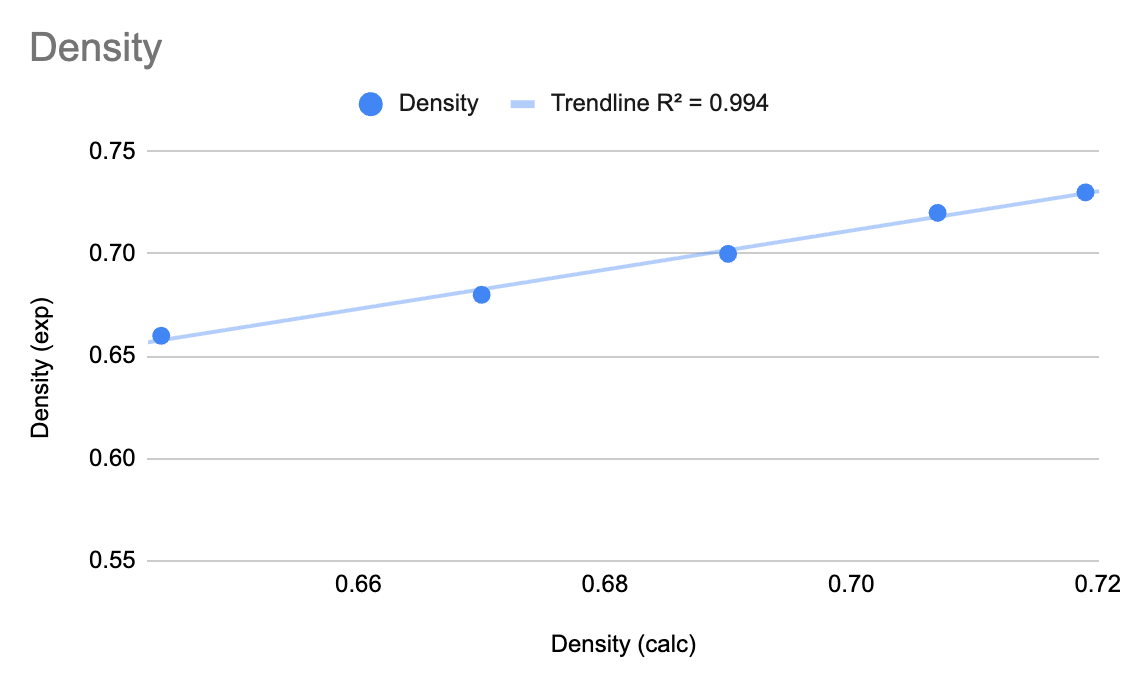
We can see that for hexane, the system has equilibrated to an average density of 0.644 which is in quite good agreement with the experimental density of 0.66. This system is ready for viscosity calculations. Feel free to check each of the alkanes.
- Afterwards, close the MS MD Trajectory Analysis panel
6. Running and Analyzing the Viscosity Calculation
We will now proceed to run the viscosity calculation on each of our five equilibrated systems. We will analyze the outputs and compare them to experimentally known values.
- Select(1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries all five new equilibrated entries from the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion (Shift + Click on all 5 rows)
- Go to Tasks > Materials > Classical Mechanics > Viscosity > Viscosity Calculations
- The Viscosity panel opens
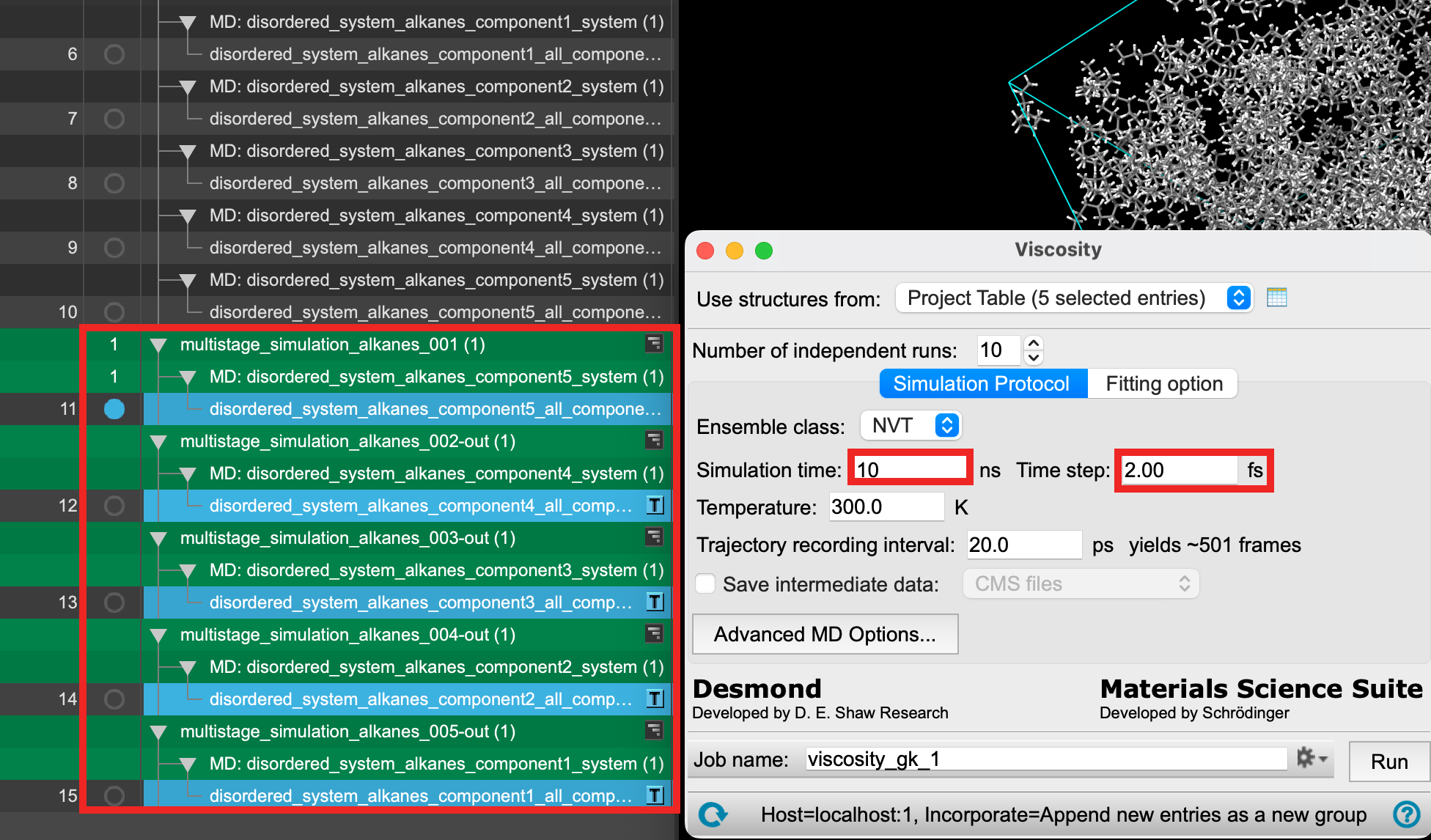
- Change the Simulation time to 10 ns
- Change the Time step to 2.0 fs
- Ensure that Use structures from directs to the 5 entries from Section 5
- Open Advanced MD Options
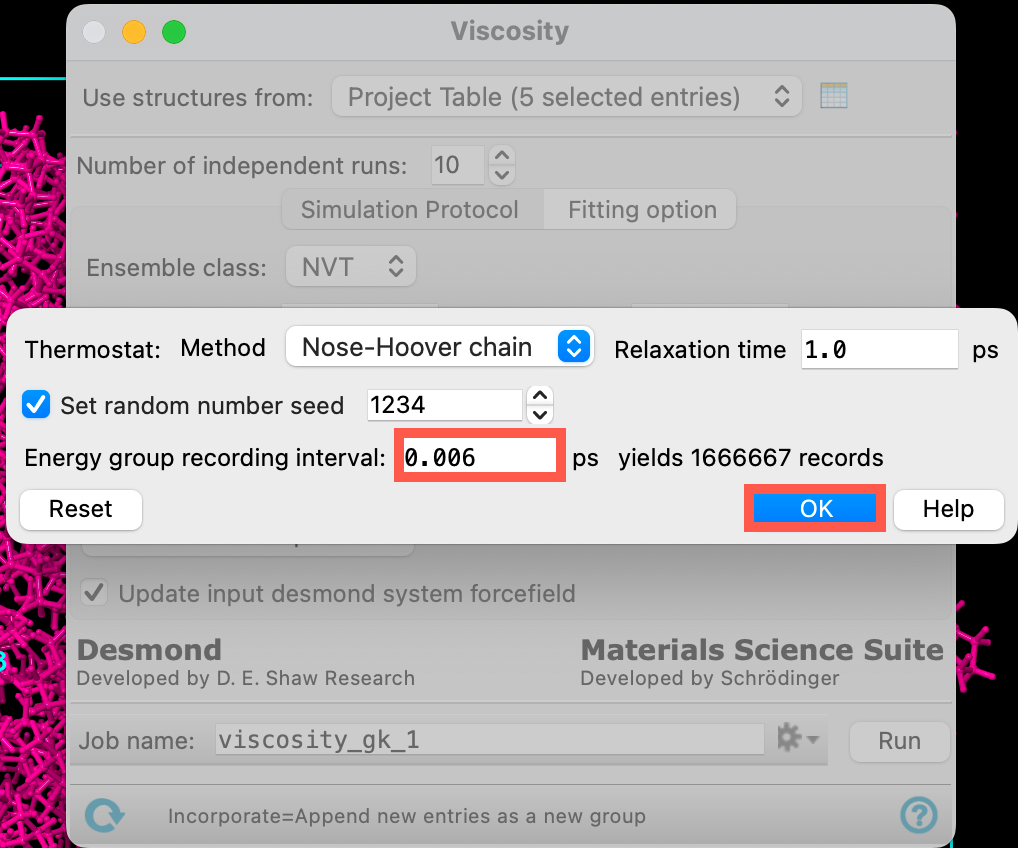
- Check Set random number seed
- Change the Energy group recording interval to 0.006 ps. Make sure to press Enter on your keyboard after updating the input to ensure that the panel updates to read yields 1666667 records
- In the cells below, we will discuss this parameter and some other best practices for using the Viscosity panel
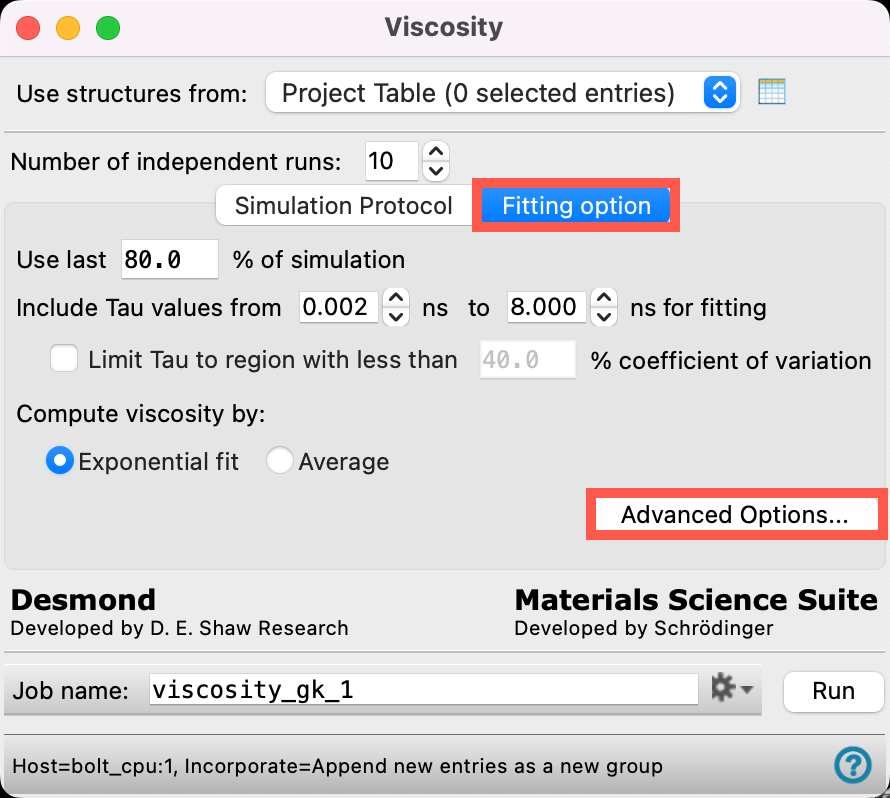
- Switch to the Fitting option tab
- Click Advanced Options
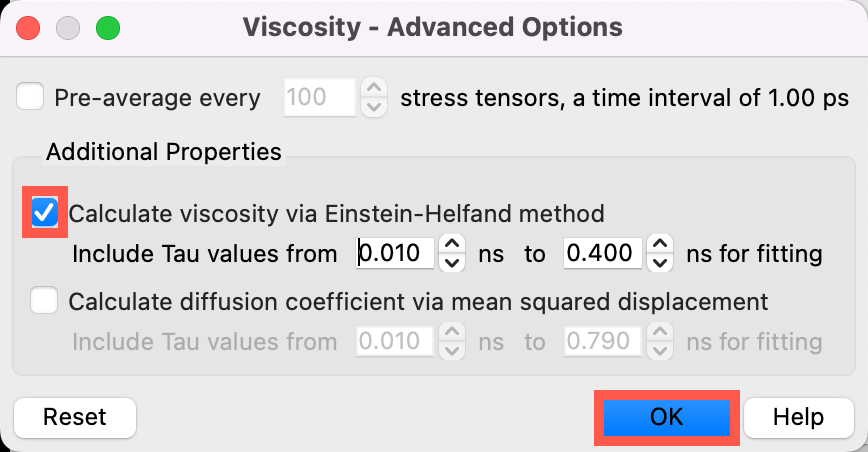
- Check the Calculate viscosity via Einstein-Helfand method box
- In the Einstein-Helfand method, the viscosity is determined from the ensemble average of the mean square displacement (MSD) of the time integral of the pressure tensors at long tau values
- Calculating the viscosity two different ways provides extra validation for the computed values
- Click OK to close the Viscosity - Advanced Options window
- Change the Job name to viscosity_alkanes
- This job takes several hours on a CPU (for the master job) and GPU Host (for the simulations), highly depending on your GPU resources. If you would like to run the job yourself, adjust your Job settings as needed and click Run
If you would prefer to proceed with pre-generated results, you can import the viscosity jobs via File > Import Structures. Navigate to where you downloaded the tutorial files and choose any or all of the Section_06 > viscosity_alkanes-00N > viscosity_alkanes-00N-out.cms files
- Close the Viscosity panel
Let’s take a moment to discuss the parameterization of the Viscosity panel and some best practices. For a complete description, refer to the help documentation
- As mentioned previously, the viscosity workflow does not include any equilibration steps, thus it is essential the system is first equilibrated (Section 5 in this tutorial).
- The Number of independent runs specifies the amount of independent MD simulations with different initial velocities to be run and analyzed when extracting shear viscosity information.
- Sampling is critical for accurate viscosity calculations. For <2 cP systems like these, 10 independent replicas each run for 10 ns of simulation time is validated as sufficient; for calculations of fluids with higher viscosities, more independent runs might be beneficial.
- The default ensemble class is NVT, but NVE is also available and is likely to produce similar results. Although NVE is in general more reliable, it requires a very well equilibrated system to ensure that the desired temperature is obtained and to avoid drift in the simulations. The NVT method is less dependent on the input structure equilibration, and has been found to produce reliable results (see References).
- In the tutorial, we use 10 ns simulations. For these <2 cP liquids, the results are very good. However, simulations of tens of nanoseconds or more are recommended for liquids with higher viscosities or for in-practice production runs; the time must be long enough to reach a plateau in the pressure tensor integral.
- The Energy group recording interval is the recording interval for saving pressure tensor information (in ps). The final result is sensitive to this value, thus we decreased the recording interval to generate 166,666 records. The interval can be lowered even more in some cases but lowering the value too much will record more frequently and can waste memory. On the other hand, if the energy group recording interval is too high (a little higher than default) an overestimation of the viscosity could occur. A lower energy group recording interval is best for higher viscosity systems.
- In the Advanced Options, you can pre-average stress tensors before the viscosity calculation. This reduces data and improves the response time of the Viscosity Viewer
- Finally, the fitting options defaults are well-parameterized for obtaining the shear viscosity, which can be calculated either by exponentially fitting the data (a double-exponential fit) or averaging the shear viscosity over the specified Tau range across the independent simulations. The Limit Tau to region parameter can be used to quantitatively decide where to stop fitting the viscosity data. Once the size of the error bars exceeds the average value of the autocorrelation function by the designated percentage, the fitting is stopped.
- Use the WAM button (
 ) to open the Viscosity Results panel
) to open the Viscosity Results panel
- Alternatively, access the panel via Tasks > Materials > Classical Mechanics > Viscosity > Viscosity Results
- The Viscosity Viewer panel opens
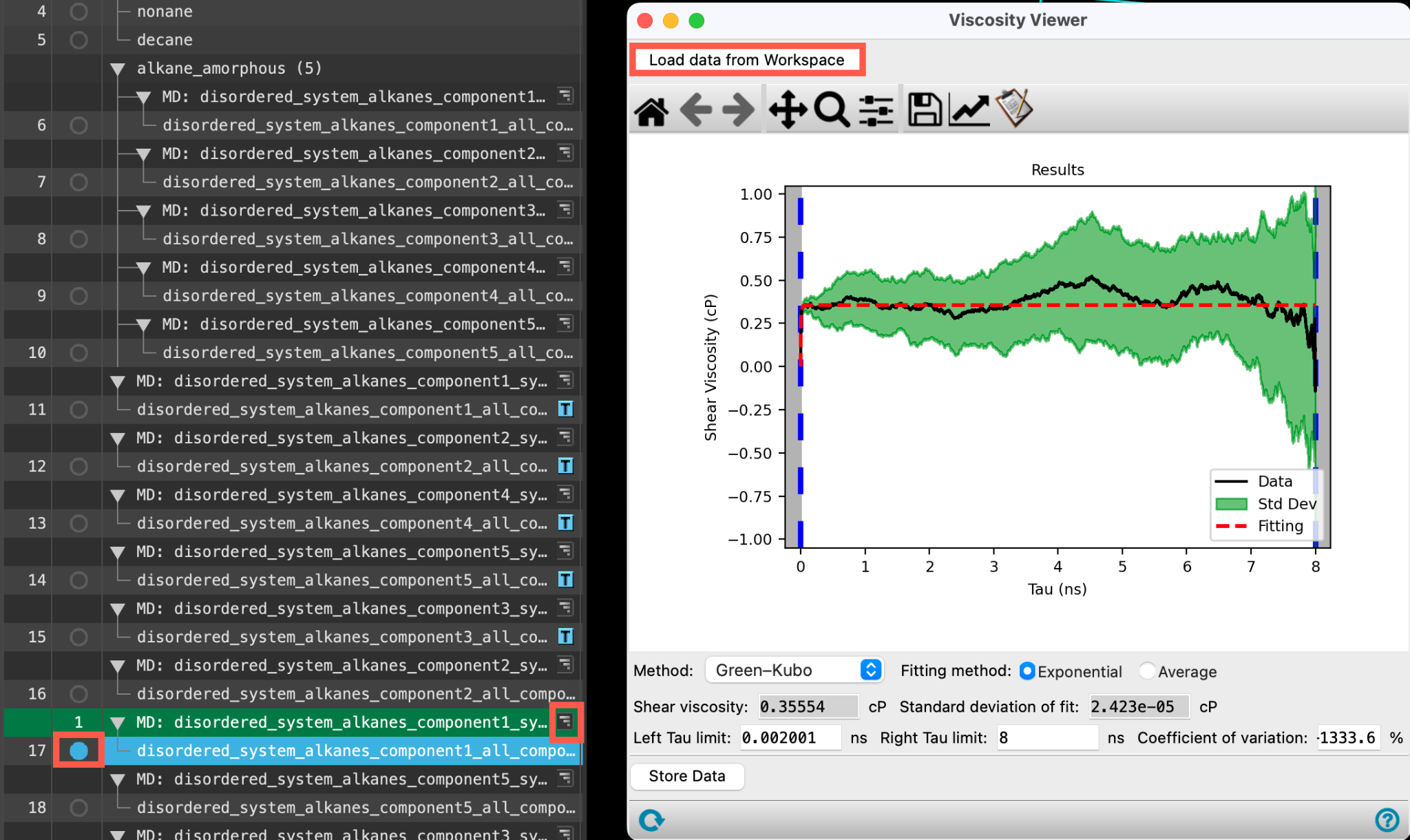
- To view the output for any of the alkanes (for example, hexane), include the relevant output from the entry lista simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion and click Load data from Workspace
- The Viscosity Viewer Panel updates to show the output
- The Shear Viscosity (cP) and Standard deviation of fit (cP) are shown below the results figure
- The Einstein-Helfand Method can be shown by toggling the Method drop-down menu, the Einstein-Helfand Method will produce a Linear fit
- Both the Green-Kubo and Einstein-Helfand Method should give a similar result
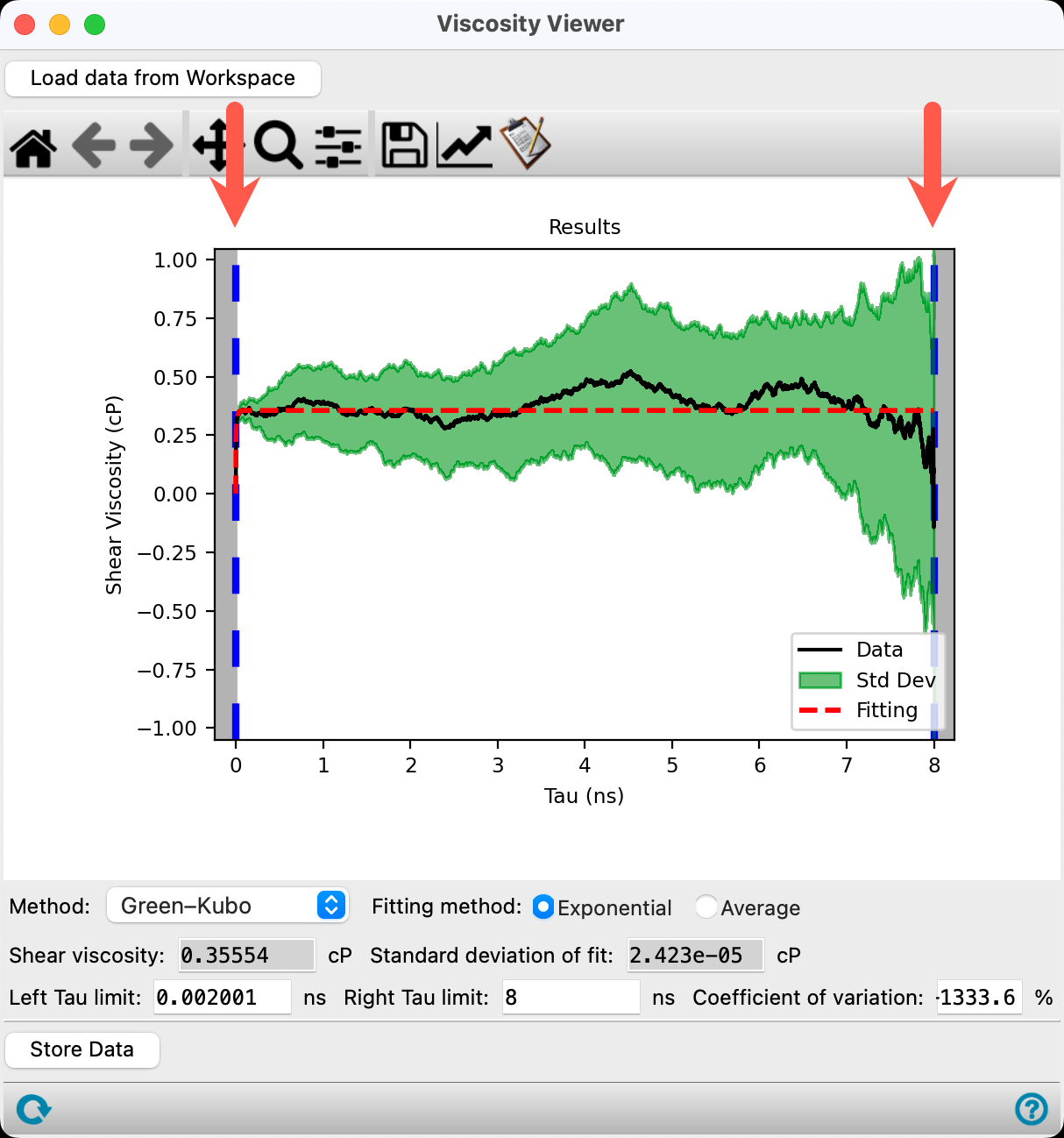
- The plot shows the last 80% of the simulation time by default
By dragging the blue dashed vertical lines, the fitting region can be manually adjusted, however, finding the best fit region for viscosity is not an exact science. The panel usually makes a pretty good guess at where to fit things but it is not perfect.
The coefficient of variation is one way to get the right tau limit. If the coefficient is set, then it will choose the right limit such that the standard deviation is x% of the viscosity at time t.
The Green-Kubo Method has two different Fitting methods. The exponential fit is empirical, but has a little bit more of a physical basis to it, while the average fitting method is good for eye-balling noisy data.
7. Conclusion and References
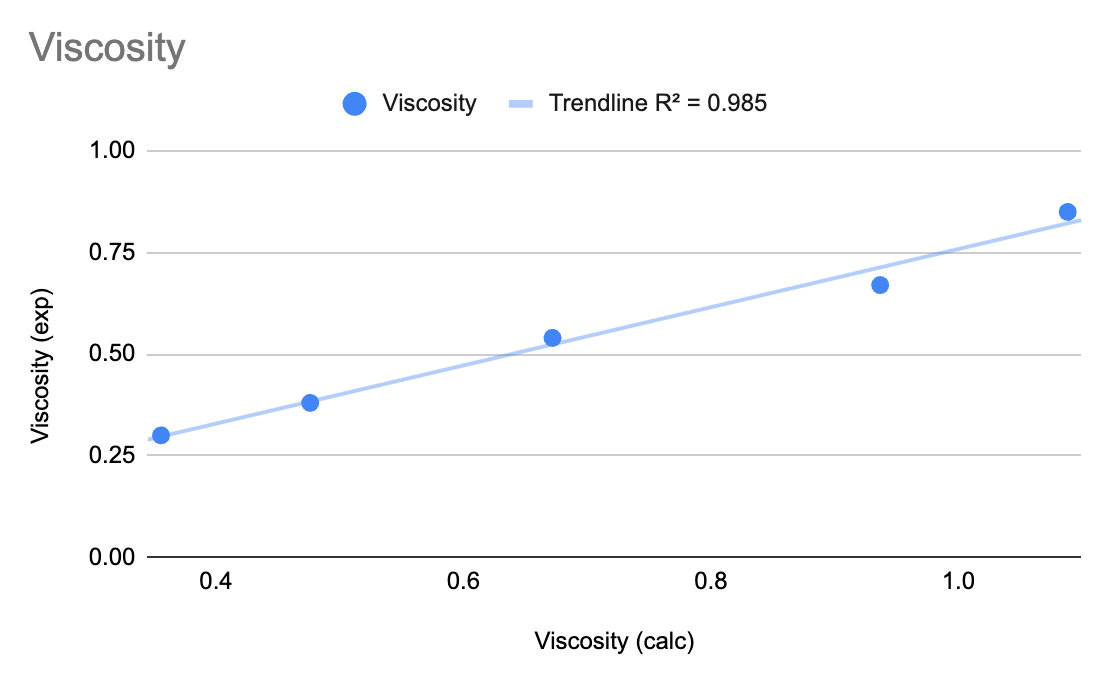
In this tutorial, we learned to calculate the viscosities of a series of alkanes using Materials Science Maestro. The viscosities and densities from the workflow were compared to known experimental values, showing quite good agreement.
For further learning:
For introductory content, focused on navigating the Schrödinger Materials Science interface, an Introduction to Materials Science Maestro tutorial is available. Please visit the materials science training website for access to 100+ tutorials. For scientific inquiries or technical troubleshooting, submit a ticket to our Technical Support Scientists at help@schrodinger.com.
For self-paced, asynchronous, online courses in Materials Science modeling, including access to Schrödinger software, please visit the Schrödinger Online Learning portal on our website.
For some related practice, proceed to explore other relevant tutorials:
For further reading:
- Reliable Viscosity Calculation from Equilibrium Molecular Dynamics Simulations: A Time Decomposition Method. DOI:10.1021/acs.jctc.5b00351
- Best Practices for Computing Transport Properties 1. Self-Diffusivity and Viscosity from Equilibrium Molecular Dynamics. DOI:10.33011/livecoms.1.1.6324
- Effects of Temperature Control Algorithms on Transport Properties and Kinetics in Molecular Dynamics SImulations. DOI:10.1021/ct400109a
- See the help documentation
8. Glossary of Terms
Entry List - a simplified view of the Project Table that allows you to perform basic operations such as selection and inclusion
Included - the entry is represented in the Workspace, the circle in the In column is blue
Project Table - displays the contents of a project and is also an interface for performing operations on selected entries, viewing properties, and organizing structures and data
Recent actions - This is a list of your recent actions, which you can use to reopen a panel, displayed below the Browse row. (Right-click to delete.)
Scratch Project - a temporary project in which work is not saved, closing a scratch project removes all current work and begins a new scratch project
Selected - (1) the atoms are chosen in the Workspace. These atoms are referred to as "the selection" or "the atom selection". Workspace operations are performed on the selected atoms. (2) The entry is chosen in the Entry List (and Project Table) and the row for the entry is highlighted. Project operations are performed on all selected entries
Working Directory - the location where files are saved
Workspace - the 3D display area in the center of the main window, where molecular structures are displayed